Abstract
Objective
High altitude pulmonary edema (HAPE) is a potentially fatal high altitude illness occurring as a result of hypobaric hypoxia with an unknown underlying genetic mechanism. Recent studies have shown a possible association between HAPE and polymorphisms in genes of the renin-angiotensin-aldosterone system (RAAS), which play a key role in sensitivity of an individual toward HAPE.
Methods
For the present investigation, study groups consisted of HAPE patients (HAPE) and acclimatized control subjects (rCON). Four single-nucleotide polymorphisms (SNPs) were genotyped using restriction fragment length polymorphism (RFLP) analysis in genes of the RAAS pathway, specifically, renin (REN) C(–4063)T (rs41317140) and RENi8–83 (rs2368564), angiotensin (AGT) M(235)T (rs699), and angiotensin-converting enzyme (ACE) insertion/deletion (I/D) (rs1799752).
Results
Only the I/D polymorphism of the ACE gene showed a significant difference between the HAPE and rCON groups. The frequency of the D allele was found to be significantly higher in the HAPE group. Arterial oxygen saturation levels were significantly lower in the HAPE group compared with the rCON group and also decreased in the I/D and D/D genotypes compared with the I/I genotype in these groups. The other polymorphisms occurring in the REN and AGT genes were not significantly different between the 2 groups.
Conclusions
These findings demonstrate a possible association of the I/D polymorphism of the ACE gene with the development of HAPE, with D/D being the at-risk genotype.
Introduction
High altitude illness may be divided into different syndromes that affect lowland or highland residents ascending to altitudes greater than those to which they are accustomed. Susceptibility to high altitude maladies is considered as the major factor limiting the individual’s performance at high altitude (HA). The processes of acclimatization help in maintenance of physical performance in tolerant individuals, whereas the nonacclimatized maladapted individuals succumb to life-threatening complications. These HA syndromes most commonly include acute mountain sickness, high altitude pulmonary edema (HAPE), and high altitude cerebral edema. Development of HAPE is a potentially fatal condition affecting healthy lowlanders or nonacclimatized individuals. This disease is a form of noncardiogenic pulmonary edema that originates with accelerated permeability and usually occurs within 2 to 4 days of ascent above 2500 to 3000 m. 1 –3 Its initial pathogenesis is owing to nonuniform hypoxic pulmonary vasoconstriction leading to pulmonary capillary stress failure and a high-permeability type of edema of the lungs in the absence of infection with normal left atrial pressure. 4 Acclimatization at HA is mediated by a combination of mechanisms involved in sustained oxygen delivery, involving increase in ventilation, cardiac output, and hemoglobin concentration, as well as metabolic changes. 5 Genetic polymorphisms in many genes play a pivotal role in determining susceptibility of an individual to HAPE. 6
It is well known that the renin-angiotensin-aldosterone system (RAAS) acts as a circulating hormonal system that regulates blood pressure, electrolyte balance, and fluid homeostasis.
7
Genetic variations occurring in the genes of the RAAS pathway have been studied in various populations with conflicting results in relation to hypertension,
8
diabetic complications,
9
coronary heart disease,
10
and renal disease.
11
Many studies have suggested the role of human RAAS in HA hypoxic adaptation also.
12
RAAS is an enzymatic cascade that is composed of renin (REN), which converts angiotensinogen (AGT) to angiotensin, which is in turn cleaved by angiotensin-converting enzyme (ACE) to produce angiotensin II (AngII), the final product of this pathway. AngII in turn interacts with its receptors to release aldosterone and bring about its other biological actions. Renin plays a crucial role in regulation of blood pressure, and its polymorphisms have been shown to have both positive
13
and negative
14
associations with essential hypertension. The AGT gene variants have also been studied in relation to hypertension
15
and diabetes
16
with conflicting results. Woods et al
17
suggested the preferential association of the I allele of ACE in the maintenance of arterial oxygen saturation (Sa
In view of the above literature, the present study was undertaken to determine the possible association of 4 genetic variants (single-nucleotide polymorphisms [SNPs] and insertion/deletion [I/D] polymorphism) of 3 genes of the RAAS pathway, ie, REN C(-4063)T (rs 41317140) ) and A/Gi8-83 (rs 2368564), AGT M(235)T (rs 699), and ACE (I/D) (rs 1799752) with HAPE in the Indian population and also to examine the association of these polymorphisms with Sa
Methods
Subjects
A total of 154 Indian lowlanders, unrelated to each other to the best of our knowledge, belonging to the Indian Army were studied. These subjects were serving Army personnel and had no familial relationships among them. They were acutely introduced to HA (≥3500 m) in Leh, in the Ladakh region of India, at different times (over the course of approximately 2 years) and were heterogeneous without predominance of any particular ethnic group. Because these subjects were all healthy Army personnel, they did not have any medical history of other pulmonary disorders. They were born and recruited at sea level and introduced to HA on posting; of these, 75 of the volunteers were acclimatized controls (rCON) and 79 individuals were those with high altitude pulmonary edema (HAPE), which had developed within 48 to 72 hours after ascent to HA. HAPE was diagnosed by physicians through chest radiography and other clinical parameters. The study was approved by the Institutional Ethics Committee of the Defence Institute of Physiology and Allied Sciences (DIPAS), and written informed consent was obtained from the volunteers before recruiting them for study. Approximately 5 mL of venous blood was collected in K3 EDTA vacutainers (Greiner GmbH, Pleidelsheim, Germany), stored at 4°C, dispatched on ice to the Delhi laboratory, and stored at –20°C until further processing.
Physiological Measurements
Physiological recordings and sample collection for 154 subjects (79 HAPE and 75 rCON), such as age, body weight, heart rate (HR), blood pressure, and Sa
Physiological characteristics of the 2 groups at high altitude: acclimatized lowlanders (controls) and individuals with high altitude pulmonary edema
BP, blood pressure; HAPE, high altitude pulmonary edema; Sa
P value is obtained using unpaired Student’s t test.
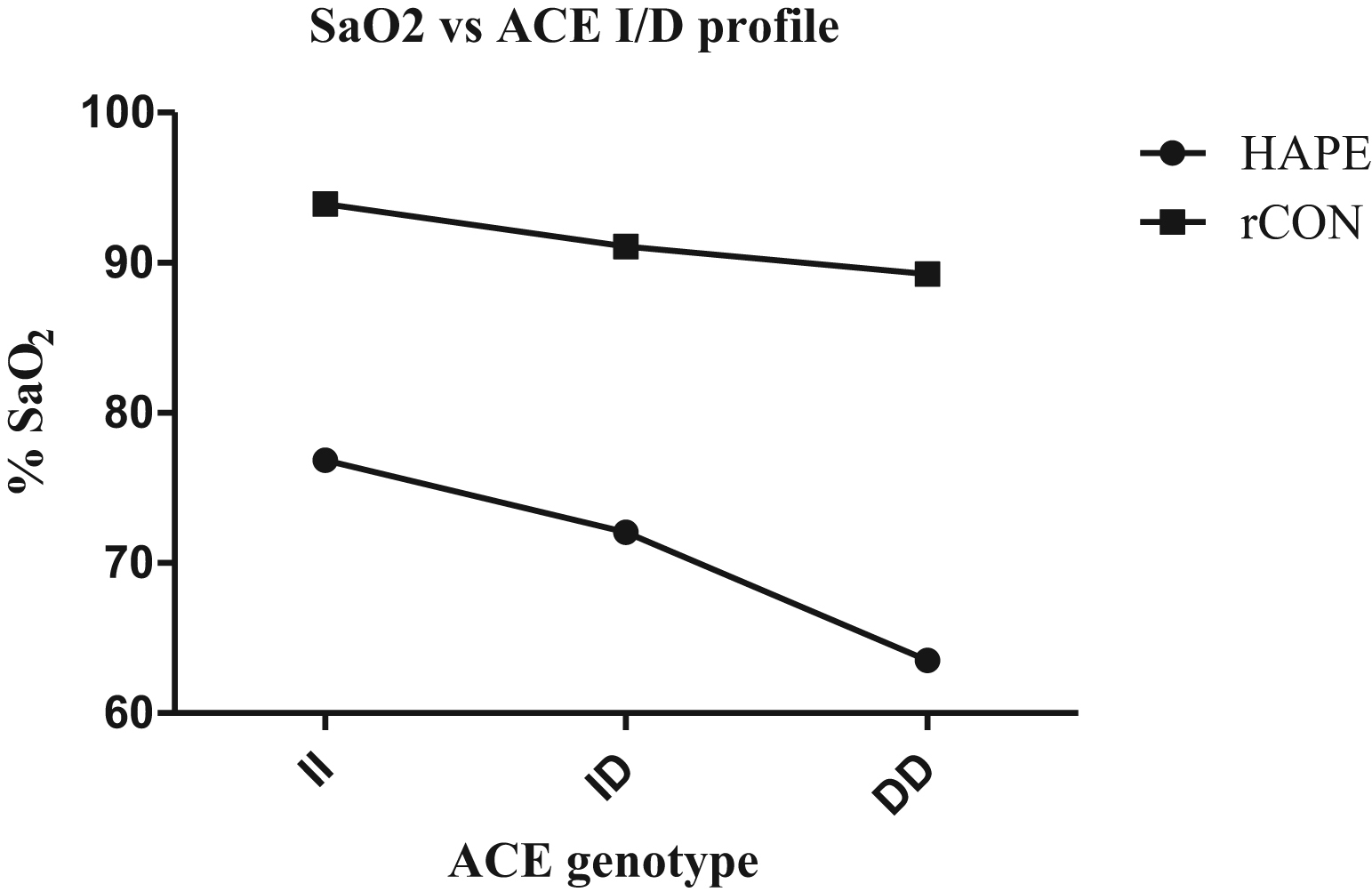
Arterial oxygen saturation compared with genotypic profile of the angiotensin-converting enzyme gene (ACE) I/D polymorphism in high altitude pulmonary edema (HAPE) and acclimatized control subjects (rCON) groups. A steady decline in the arterial oxygen saturation (Sa
Genomic Dna Isolation And Genotypic Analysis
Blood samples were taken from –20°C storage and thawed on ice. Extraction of genomic DNA from peripheral leukocytes was performed using phenol extraction from sodium dodecyl sulfate–lysed, proteinase K–treated cells by overnight incubation at 37°C. A quantitative check of DNA was done using DNA/RNA Quant calculator (Amersham Pharmacia, Amersham, UK). For qualitative check, DNA samples were diluted to concentration of 20 ng/µL and loaded on 0.7% agarose gel containing ethidium bromide (EtBr) and run in 0.5× Tris/borate/EDTA buffer for approximately 15 minutes. Gel images were visualized under UV light in a Fluor-S Multi-Imager with Quantity One software (BioRad, Hercules, CA). The present study evaluated 4 genotypic polymorphisms in genes of the RAAS pathway selected in view of their functional relevance in the published literature: REN, AGT, and ACE. In the REN gene, 2 polymorphisms, C(–4063)T occurring in the promoter region and A/G occurring in intron 8, were studied. In the AGT gene, a polymorphism occurring in the region of exon 2, M(235)T was studied, and in the ACE gene, the I/D polymorphism in intron 16 was evaluated. These specific regions of genes were amplified using 5 to 10 pmol of specific primer sequences 24 along with 1× buffer containing 1.5 mM MgCl2, 0.6 U Taq DNA polymerase, 2.2 mM/L deoxyribonucleoside triphosphate in a final volume of 25 µL. A negative control containing no genomic DNA and a positive control with a known genotype were always included in all experiments. Amplified gene sequences were digested by specific restriction enzymes with respective buffers in a final reaction volume of 35 µL. The digested polymerase chain reaction (PCR) products were resolved on 2% to 3% agarose gels stained with EtBr. Gel images were visualized under UV (Fusion Fx5). For the ACE I/D polymorphism, as there is a tendency of the D allele in heterozygous samples to display a preferential amplification, 25 samples found to have the D/D and I/D genotype were subjected to a second independent PCR amplification using an insertion-specific primer sequence 26 for confirmation of the I allele. Table 2 shows the PCR primers and cycling conditions, together with restriction enzymes for RFLP and their respective resulting fragments. An independent observer read and confirmed all the genotypes; discrepancies, if any, were resolved by repeated RFLP.
Details of single nucleotide polymorphisms studied in the rennin-angiotensin-aldosterone system pathway showing the primer sequences, polymerase chain reaction product size, annealing temperature, restriction enzyme, and the fragment sizes
ACE, angiotensin-converting enzyme gene; AGT, angiotensinogen gene; PCR, polymerase chain reaction; REN, renin gene.
Statistical Analysis
Allele and genotype frequencies were determined by gene counting and compared by 2 × 2 and 3 × 2 contingency table, respectively. Pearson χ2 test (3 × 2 contingency table) was used to assess the association of genotype frequencies. For Hardy-Weinberg’s equilibrium (HWE) calculations, a 2 × 2 statistic 1 degree of freedom) was computed by an online tool, the Hardy-Weinberg calculator.
Differences in Sa
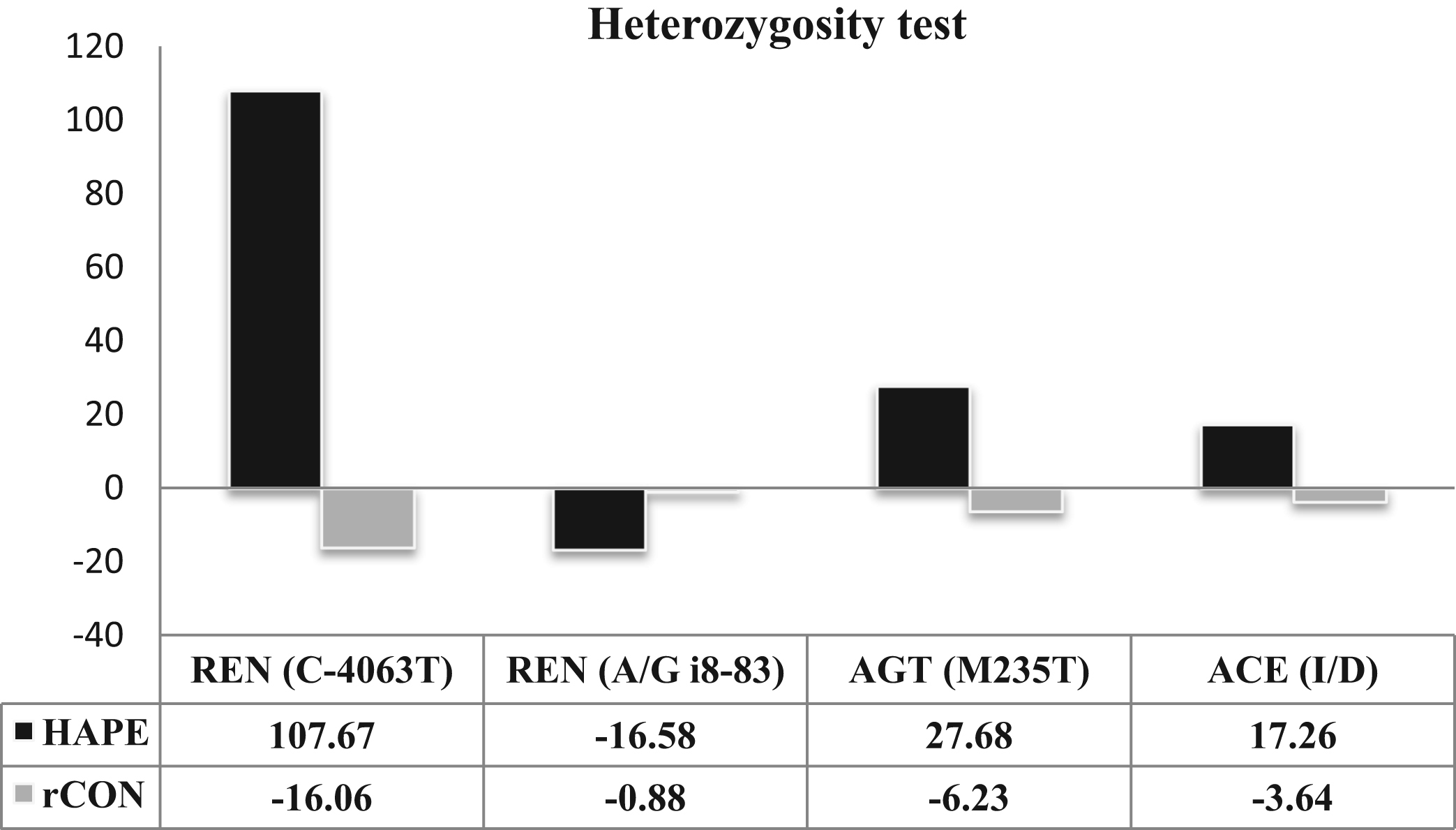
The graph shows the percent heterozygosity (expected heterozygosity – observed heterozygosity) for the high altitude pulmonary edema (HAPE) and acclimatized control subjects (rCON) groups in all 4 single-nucleotide polymorphisms (SNPs). The renin gene (REN) (C–4063T) SNP shows the highest difference in heterozygosity for HAPE and rCON samples: HAPE samples for this SNP had a much lower observed heterozygosity than expected, as evident from the graph. Also, in the HAPE group the angiotensinogen gene (AGT) (M235T) and the angiotensin-converting enzyme gene (ACE) (I/D) SNPs showed a lower observed heterozygosity than expected; however, the difference is not very prominent.
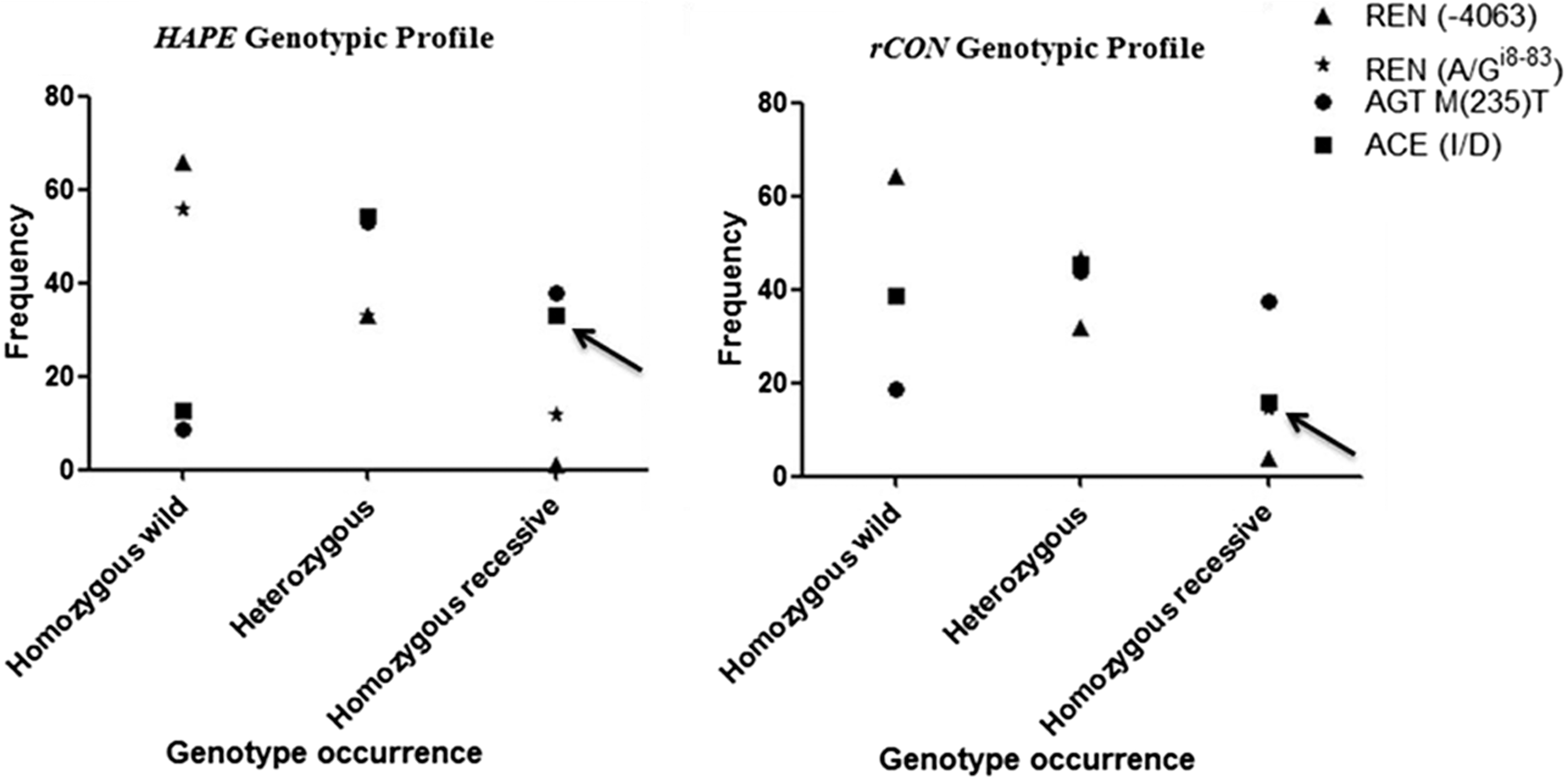
The observed genotypic diversity for high altitude pulmonary edema (HAPE; A) and acclimatized control subjects (rCON; B) is shown for the 4 single-nucleotide polymorphisms (SNPs) under study. A noticeable difference occurs for the angiotensin-converting enzyme gene (ACE) (I/D) SNP in homozygous recessives between HAPE patients and control subjects. AGT, angiotensinogen gene; REN, renin gene.
Results
Baseline Characteristics
The physiological characteristics of all 154 volunteers (including both HAPE patients and acclimatized control subjects) were recorded at HA. These readings were recorded before collection of blood samples and are presented in Table 1. Significant differences were found between the HAPE and rCON groups for HR and Sa
Genotyping
Renin
In the present study, genotyping was carried out for 2 previously reported SNPs of the renin gene ie, C(–4063)T (rs41317140) and A/Gi8-83 (rs2368564). The RFLP analysis demonstrated no significant difference between the HAPE and rCON groups for both C(–4063)T and A/Gi8-83 polymorphisms at the genotypic as well as the allelic level (Table 3). The distribution of genotypes for both SNPs was in accordance with HWE for both patient and control groups (Table 3).
Observed genotypic and allelic frequencies of polymorphisms in AGT, ACE, and REN genes, with statistical analysis for the single-nucleotide polymorphisms under study
ACE, angiotensin-converting enzyme gene; AGT, angiotensinogen gene; HWE, Hardy-Weinberg’s equilibrium; OR, odds ratio; REN, renin gene.
Angiotensinogen
The M(235)T polymorphism of the AGT (rs699) gene also showed a statistically insignificant difference between the HAPE and rCON groups both at the genotypic and allelic levels (Table 3). However, the percentage distribution of genotypes was in accordance with HWE.
Angiotensin-converting enzyme
The ACE I/D polymorphism (rs1799752) showed a significant difference at the genotypic (χ2 = 15.3; P = .0005) as well as the allelic level (OR, 0.41; 95% CI, 0.26–0.66; P = .0002; Table 3). The percentage frequency of the D/D genotype was higher in the HAPE group (32.91%) compared with the rCON group (16.0%). Although the D allele had significantly higher prevalence in the HAPE group, it was in accordance with HWE (χ2 = 0.89; P = .34) for this SNP (Table 3). Both dominant and recessive models for the I/D polymorphism showed significant association with the phenotype of HAPE (Table 4).
Effect of D allele assuming the dominant and recessive modes on patients with high altitude pulmonary edema
P value was calculated by Fisher’s exact test (2 × 2 contingency table; 1 degree of freedom).
Analysis of Heterozygosity
The observed heterozygosity was lower than expected for SNPs REN C(–4063)T, AGT M(235)T, and ACE I/D in the HAPE group (Figure 2). There was a striking difference between the observed and expected heterozygosity of the HAPE group in REN C(–4063)T polymorphism, specifically, the observed heterozygosity was much lower than expected. These differences could be caused by ascertainment bias in the biological sample, which may in turn lead to gains or losses in observed counts.
Figure 3 depicts the distribution of observed genotypes for all 4 SNPs under study in both the HAPE (A) and rCON (B) groups on the basis of percentage of occurrence of homozygous dominants, heterozygotes, and homozygous recessives. The graph indicates a similar distribution of genotypes in both groups for SNPs REN C(–4063), REN A/Gi8-83, and AGT M(235)T. The D/D genotype distribution of the ACE gene visibly differs between the 2 groups.
Discussion
The present study examines the association of gene polymorphisms of the RAAS pathway with an individual’s susceptibility to HAPE and found positive association of the ACE I/D polymorphism with HAPE susceptibility. HAPE is the cause of the majority of deaths at HA 2 and is typically associated with pulmonary hypertension and elevated capillary pressure. 27 The previous reports have shown that genes of the RAAS pathway play an important role in regulating blood pressure by influencing vascular tone, electrolyte balance, and salt and water homeostasis 7 and thus could be a logical candidate for evaluation of sensitivity to HAPE. A recent study has shown a possible association between the polymorphisms existing in the genes of the RAAS pathway and HAPE susceptibility. 28
In the search for the inheritable genetic biomarkers of HAPE, the gene encoding the ACE protein (chromosome 17q23) ranks high. ACE catalyzes generation of angiotensin II and has the capability to modulate the pulmonary vasoconstrictor response to hypoxia via interaction with its receptors.
29
However, there is wide interindividual variability of plasma ACE levels, which might be explained by the existence of the I/D polymorphism. Qi et al
30
conducted a meta-analysis of consolidated data to investigate the association between the ACE I/D polymorphism and HAPE. Their results supported the notion that ACE D allele carriers were at significantly increased risk of developing HAPE in a Chinese population. The result of the present study concurs with those of Qi et al.
30
We report a significant genetic difference in genotypic (χ2 = 15.3, P = .0005) and allelic frequencies (P = .0002) of the ACE I/D polymorphism between individuals with HAPE in comparison to acclimatized control subjects; the frequency of the D/D genotype was much higher in the HAPE group (32.91%) compared with that of the rCON group (16%; Table 3). The I allele has been associated with lowered ACE protein levels compared with the D allele.20,29 Moreover, Woods et al
17
showed that individuals with the I/I genotype maintain a higher Sa
For the 2 polymorphisms studied in the REN gene, C(–4063)T and A/Gi8-83, although data are available for their association with essential hypertension, 13 we could not establish their contribution to susceptibility to HAPE in the Indian population. One of our previous studies had shown an association of the T(174)M polymorphism of the AGT gene with HAPE, 35 which prompted us to examine the association of another SNP of this gene, M(235)T, with HAPE. This SNP was previously shown to be associated with both hypertension 36 and HAPE 28 ; however, the present investigation did not show any association of AGT M(235)T with HAPE. Besides polymorphisms in the RAAS pathway, recent findings of Aggarwal et al 37 showed that polymorphisms in the EGLN1 gene are associated with HA adaptations and certain genotype combinations are overrepresented in lowlanders, which make them more susceptible to the risk of HAPE.
Limitations
An association study between genetic polymorphism and genetic predisposition remains controversial. Although it is an efficient method to evaluate the association of genotype or allele frequencies of candidate genes with a disease, it is not able to shed light on the pathological process. One limitation of the present study lies in its sample size and homogeneous ethnic background; a larger and more ethnically diverse sample size could have increased the reliability of our observations and its application to other ethnic groups.
Conclusions
In summary, among the genes of the RAAS pathway, REN C(–4063)T and A/Gi8-83, AGT M(235)T, and ACE (I/D), the D/D genotype of the ACE gene appears to be associated with an increased risk of HAPE in the Indian population. In the future, other SNPs in the ACE gene could be evaluated for an association with HAPE and lead to the determination of the pathway that connects ACE gene polymorphisms with this condition.
Footnotes
Acknowledgments
The authors sincerely thank officers in charge: Col G Himashree, Col RKS Tomal, Col Rajiv Bajaj, and other supporting staff of HAMRC, Leh, for their cooperation and logistic support in collection of blood samples and to all the volunteers involved in the study. We are also highly grateful to Mr Khem Chandra and Ms Babita Kumari for excellent technical assistance.
Funding
The funding for the experiments was provided by the Defence Research and Development Organization, Ministry of Defence, India.
☆
Conflict of interest: The authors declare that they have no conflict of interest.
