Abstract
Objective
The purpose of this study was to test the hypothesis that polymorphisms in the endothelial PAS domain protein 1 (EPAS1) gene are associated with the susceptibility to high altitude pulmonary edema (HAPE) in Han Chinese.
Methods
This study enrolled 153 HAPE patients (HAPE-p), matched with Han Chinese resistant to HAPE (HAPE-r) and local highland Tibetans from Yushu earthquake construction population in Qinghai where the altitude is more than 3500 m above sea level. The polymorphism of EPAS1 chr2:46441523(hg18) was genotyped by polymerase chain reaction restriction fragment length polymorphism and confirmed by DNA sequencing.
Results
The frequencies of EPAS1 chr2:46441523(hg18) polymorphism C allele were significantly higher in the HAPE-p group than in the HAPE-r group (P < .001), but the frequencies of heterozygous C/G were significantly higher in the HAPE-r group than in the HAPE-p group (P < .001). Moreover, the frequencies of the EPAS1 chr2:46441523(hg18) polymorphism G allele were significantly higher in the highland Tibetan group than in the HAPE-p and HAPE-r groups.
Conclusions
The EPAS1 chr2:46441523(hg18) polymorphism C is strongly associated with susceptibility to HAPE in Han Chinese, and the EPAS1 chr2:46441523(hg18) polymorphism G is present at high frequency and may be associated with high altitude adaptation in the Tibetans.
Introduction
High-altitude pulmonary edema (HAPE) is noncardiogenic pulmonary edema that usually occurs at altitudes above 3,000 m in rapidly ascending, nonacclimatized persons within the first 2 to 5 days after arrival.1,2 HAPE is characterized by high pressure in pulmonary arteries, with edema in pulmonary interstitial tissue and alveoli, leading to pulmonary capillary stress failure and a high permeability type of edema. 3 Although hypoxia is a major trigger factor, the pathogenesis of HAPE remains unclear because some persons are more susceptible to HAPE than others when exposed to the same hypoxia conditions. Growing evidence suggests that genetics plays an important role in the risk of HAPE.4,5 Recently, several candidates have been proposed as HAPE susceptibility genes. 6 –8 However, the exact genetic mechanism responsible for the development of HAPE is largely unknown.
The endothelial PAS domain protein 1 (EPAS1) gene is one of the most notable candidate genes that have been identified as contributing to evolutionary adaptation to high altitude in Tibetans, probably by some unknown molecular pathways that delicately control erythropoietic response to hypoxia. 9 –11 The EPAS1 gene is located on the short (p) arm of chromosome 2 between positions 21 and 16, from base pair 46,524,540 to base pair 46,613,841 on chromosome 2. Human EPAS1 encodes the oxygen-sensitive alpha subunit of hypoxia-inducible factor-2 (HIF-2) transcription factor, which is a key regulator of chronic hypoxia by regulating a large number of genes involved in the cellular and systemic responses to hypoxia including erythropoiesis, angiogenesis, vascular regulation, and anaerobic metabolism. 10 The expression of HIF-2 is induced by hypoxia, and it demonstrates a selective pattern of expression in microvascular endothelial cells, lung epithelial cells, cardiac myocytes, and the brain; and an EPAS1 gene knockout mouse model revealed a significant reduction in hematocrit levels and peripheral blood counts. 10 Recent genetic studies have provided evidence for natural selection on EPAS1 gene in the Tibetan population during their adaptation to living on the Tibetan Plateau. 11 –14
Notably, the EPAS1 gene has a single nucleotide polymorphism (SNP), chr2:46441523 (hg18), 5 base pairs from the beginning of exon 6, and this SNP has a 78% frequency difference between highland Tibetan and lowland Han Chinese. 13 Furthermore, our preliminary experiments on the sequencing of EPAS1 gene in HAPE patients and healthy controls showed that there were no changes in the sequences of all exons, but the frequency of SNP chr2:46441523 (hg18) was different between HAPE patients and healthy controls. Therefore, we hypothesized that SNP chr2:46441523 (hg18) of EPAS1 is associated with the susceptibility to HAPE in Han Chinese. To test our hypothesis, in this study we detected the EPAS1 chr2:46441523 (hg18) polymorphism in HAPE patients, matched Han Chinese and local Tibetans from Yushu earthquake construction population in Qinghai where the altitude is more than 3,500 m above sea level.
Methods
Subjects
A total of 153 HAPE patients (HAPE-p) had been hospitalized in Yushu People’s Hospital between March 2010 and June 2012 owing to the onset of HAPE after arriving 1 to 2 days at Yushu (3760 m). The diagnosis of HAPE was based on chest radiographs and standard diagnostic criteria. 15 A total of 298 healthy controls resistant to HAPE (HAPE-r) were randomly selected from among the coworkers of HAPE-p, matching the patients in age, sex, ethnicity, and working conditions. These subjects remained healthy after working at Yushu for at least 3 months, without having HAPE or high altitude cerebral edema. As controls, 245 healthy highland Tibetans (HLT) from the Yushu area were randomly selected. The study protocols were approved by the Ethics Committee of Qinghai University (Xining, China). All participants in this study signed informed consent.
Sample Collection
Venous blood samples, 5 mL, were collected from each participant. The whole blood was separated into blood cells and plasma by centrifuging at 1000g for 10 minutes, and then the blood cells and plasma were transported to Xining for biochemical assays and genetic polymorphism analysis.
DNA Extraction and GENOTYPING Assays
Genomic DNA was extracted from venous blood by Gentra Puregene Blood Kit (158389; Qiagen GmbH, Hilden, Germany) according to standard procedures. The EPAS1-[ch2:46441523(hg18)] polymorphism was examined by polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP) with the following primers: 5’-AGGCTCTGGTTTTGGGAA-3’ (forward) and 5’-TGGGATGGGTGCTGGATT-3’ (reverse). The PCR mixture contained 12.5 µL Phusion High-Fidelity PCR Master Mix (F-532L; Thermo Fisher Scientific, Waltham, MA), 1 µL each primer (final concentration 10 µM), and 50 ng genomic DNA in a final volume of 25 µL. The PCR cycles consisted of denaturation at 96°C for 1 minute, 35 cycles of 96°C for 10 s, 60.5°C for 30 s, and 72°C for 20 s, and a final elongation at 72°C for 10 minutes. Amplified fragments were digested by BsmAI (New England Biolabs, Ipswich, MA), and the digested PCR products were resolved on 1.8% agarose gels stained with ethidium bromide. The fragments run as 247 and 130 bp fragments in the presence of the C nucleotide or as uncut 377 bp fragment in the presence of the G nucleotide. The polymorphism was confirmed by DNA sequencing with ABI 3730 DNA analyzer.
Statistical Analysis
Statistical analyses were performed using the SPSS software (version 17.0; SPSS Inc, Chicago, IL). Continuous values were expressed as means ± SD. Deviation of genotype frequency from Hardy-Weinberg equilibrium was assessed by χ2 test with 1 degree of freedom. Allele frequencies were calculated based on genotype frequencies in the HAPE and control groups, and the intergroup difference was estimated with the χ2 test.
Results
Clinical Characteristics
The average age, heart rate, arterial oxygen saturation, and concentration of hemoglobin plus hematocrit for the HAPE-p, HAPE-r, and HLT groups are listed in Table 1. The incidence of HAPE was much higher among men than among women, consistent with the predisposition of men to HAPE. 1 We found that arterial oxygen saturation was significantly lower whereas heart rate was significantly higher in the HAPE-p group compared to the HAPE-r and HLT groups. Compared to the HLT group, hematocrit was at a similar level in the HAPE-p group but was significantly higher in the HAPE-r group.
High altitude exposures and physiological phenotypes for the study populations
Values are means ± SD.
HGB, hemoglobin; Hct, Hematocrit; SaO2, oxyhemoglobin saturation; HR, heart rate.
P<.05 vs HAPE-r
P<.05 vs HLT
P<.05 vs HAPE-p.
Genotype and Allele Distribution
The EPAS1 chr2:46441523(hg18) polymorphism was examined by PCR-RFLP and DNA sequencing. As shown in the Figure, the results of PCR-RFLP matched to the corresponding results of DNA sequencing, confirming the validity of our experimental approach.
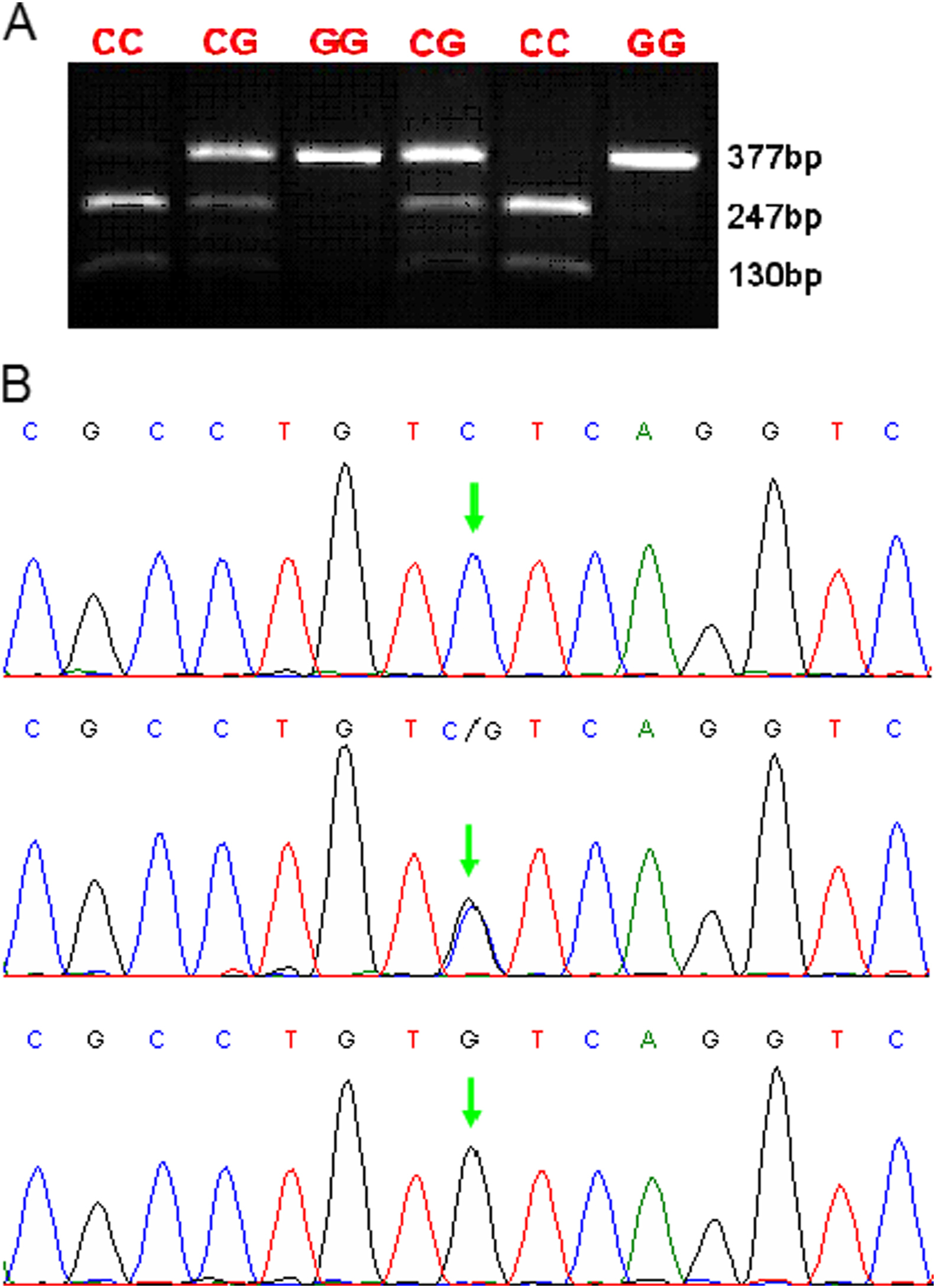
Endothelial PAS domain protein 1 (EPAS1) chr2:46441523(hg18) genotypes analyzed by polymerase chain reaction restriction fragment length polymorphism (PCR-RFLP) and DNA sequencing. (A) The PCR products were digested with BsmAI and separated by gel electrophoresis. The length of PCR products was 377 bp. The RFLP patterns were as follows: C/C genotype (247 bp and 130 bp); C/G genotype (377 bp, 247 bp, and 130 bp); G/G genotype (377 bp). (B) Sequencing results of PCR product containing EPAS1 chr2:46441523(hg18) polymorphism site. The single nucleotide polymorphism site is indicated by the red arrowhead. The results were completely matched to the corresponding results derived from PCR-RFLP.
The genotypic distributions and the allelic frequencies of the EPAS1 chr2:46441523(hg18) polymorphism in each group are summarized in Tables 2 through 4. The polymorphism was found to be in Hardy-Weinberg equilibrium (HWE) only for HLT group (0.337), whereas deviations from HWE were found for this SNP in the HAPE-p group (0.013) and HAPE-r group (0.00003308). There were significant differences in the allele frequency among the 3 groups (P<.001). The C allele was significantly more prevalent among the HAPE-p group (92.5%) than the HAPE-r group (75.2%, P < .001) with an odds ratio of 0.246 (95% CI: 0.155 to 0.391) and the HLT group (24.9%, P < .001) with an odds ratio of 0.025 (95% CI: 0.015 to 0.040), respectively. The C allele was also significantly more prevalent among the HAPE-r group (75.2%) than the HLT group (24.9%, P < .001) with an odds ratio of 0.101 (95% CI: 0.076 to 0.133). There was no significant difference in the genotypic distribution of CC between the HAPE-p group and the HAPE-r group (P = 0.627) with an odds ratio of 0. 699 (95% CI: 0.164 to 2.981), but there was significant difference in the genotypic distribution of CC between the HAPE-p group and the HLT group (P < .001) with an odds ratio of 0. 003 (95% CI: 0.001 to 0.010), as well as between the HAPE-r and the HLT group.
Genotypic distributions and allelic frequencies of the EPAS1 chr2:46441523(hg18) polymorphism in 3 groups
HAPE-p, high altitude pulmonary edema patients; HAPE-r, high altitude pulmonary edema resistant; HLT, highland Tibetans; OR, odds ratio.
The χ2 and P values for genotypes in the 3 groups
HWE, Hardy-Weinberg equilibrium; HAPE-p, high altitude pulmonary edema patients; HLT, highland Tibetans; HAPE-r, high altitude pulmonary edema resistant.
The χ2 and P values for allele frequencies in the 3 groups
HAPE-p, high altitude pulmonary edema patients; HAPE-r, high altitude pulmonary edema resistant; HLT, highland Tibetans.
The genotype distribution of CG was significantly more prevalent among the HAPE-p group than the HAPE-r group (P = 0.026) with an odds ratio of 4.871 (95% CI: 1.068 to 22.212) and the HLT group (P<.001) with an odds ratio of 0.108 (95% CI: 0.031 to 0.378). The genotype distribution of CG was also significantly more prevalent among the HAPE-r group than the HLT group (P < .001) with an odds ratio of 0.108 (95% CI: 0.031 to 0.378) and 0.004 (95% CI: 0.001 to 0.010), respectively. In contrast, the incidence of the G allele and the genotype distribution of GG in the HLT group were significantly more prevalent among the HLT group, suggesting a dominant effect of the G allele and the genotypic distribution of CC in the Tibetans.
Discussion
In this study we investigated the EPAS1 chr2:46441523(hg18) polymorphism in HAPE patients, matched Han Chinese, and local Tibetans from the Yushu earthquake construction population in a high altitude area. We found that the frequencies of EPAS1 chr2:46441523(hg18) polymorphism C allele were significantly higher in the HAPE-p group than in the HAPE-r group, but the frequencies of heterozygous C/G were significantly higher in the HAPE-r group than in the HAPE-p group. Moreover, the frequencies of the EPAS1 chr2:46441523(hg18) polymorphism G allele were significantly higher in the HLT group than in the HAPE-p and HAPE-r groups.
Recently, different research groups have reported the genome-wide studies of Tibetans in extreme hypoxia environments, and candidate genes in the HIF oxygen-signaling pathway have been suggested to be responsible for the genetic adaptation in Tibetans living at high altitude.11,13 Further association studies found that the SNPs located in and around EPAS1, EGLN1, and PPARA genes are correlated with not only the concentration of hemoglobin but also energy utilization. 16
Based on the Affymetrix v5.0 SNP arrays (Affymetric, Santa Clara, CA) of a Han Chinese family with multiple members affected with HAPE, we investigated the association of EPAS1 haplotype with susceptibility to HAPE in Han Chinese. 9 The noncoding nucleotide variants in EPAS1 gene are strongly associated with reduced blood concentration of hemoglobin in Tibetans. 6 ,11,13 The EPAS1 chr2:46441523(hg18) SNP is located between exon 5 and exon 6 and may be involved in alternate splicing of mRNA owing to its close proximately to the exons. 17
In this study, the EPAS1 SNP was found to be in HWE only for the HLT group, and deviations from HWE were found for the EPAS1 SNP in the HAPE-p and HAPE-r groups. We found a significantly higher incidence of the EPAS1-C allele in the HAPE-p group compared to HAPE-r group (P < .001), and a significantly lower incidence of the EPAS1-C allele in the HLT group compared to the HAPE-p group and HAPE-r group (P < .001). Interestingly, we found that the incidence of the EPAS1-G allele in the HLT group was much higher than in the HAPE-p group and HAPE-r group. These results are consistent with the significant divergence of alleles for the EPAS1 SNP (C/G) between Han and Tibetan Chinese reported previously.13,17 For the EPAS1 SNP, the significant departure from HWE in the HAPE-p group was the result of a lack of EPAS1-GG genotypes compared to HAPE-r and HLT groups. This overdominance effect may be due to the balancing selection for the EPAS1 SNP in the highland Tibetan who have adapted to high altitude for generations.
Several limitations of this study should be pointed out. First, owing to the limitations of the equipment, we did not measure other useful clinical parameters in the participants, such as body mass index, systolic blood pressure, diastolic blood pressure, mean arterial pressure, and pulmonary artery systolic pressure, which would provide more insight into the implication of EPAS1 variants in HAPE. Second, we selected only 1 SNP and did not screen other SNPs. Third, a moderate number of participants was enrolled in the study. Larger samples of GG persons would be included in the HAPE group to increase the power to detect differences between the genotypes. We will enroll more samples and screen more EPAS1 variants to investigate their association with HAPE. In addition, with the advances of new techniques, it is necessary to employ genome-wide association studies, whole exome sequencing, and whole genome resequencing to understand molecular mechanisms of HAPE diseases, which will help develop new approaches to the prevention and treatment of HAPE.
Conclusions
In summary, our data suggest that the EPAS1 chr2:46441523(hg18) polymorphism is strongly associated with the susceptibility to HAPE in Han Chinese, and the EPAS1 chr2:46441523(hg18) polymorphism G is present at high frequency and may be associated with high altitude adaptation in Tibetans.
Footnotes
Acknowledgments
This work was supported by grants from National Basic Research Program of China (No. 2012CB518200), Program of International S&T Cooperation of China (No. 2011DFA32720), Natural Science Foundation of China (No. 31160219 & 31160232), and Natural Science Foundation of Qinghai (No. 2011-Z-919Q). The authors declare that they have no competing interests.
