Abstract
Background.—
The role of genetics in determining an individual's susceptibility to high altitude pulmonary edema (HAPE) is unclear. However a number of genetic polymorphisms have recently been found to be overrepresented in patients with HAPE. Changes at the mitochondrial level may play an important role in the human body's adaptation to hypoxia. Polymorphisms of mitochondrial DNA (mtDNA) have been shown to be responsible for differences in organelle function. Therefore, the frequency of mtDNA 3397A/G and 3552T/A polymorphisms were studied to determine their potential role in HAPE.
Objectives.—
To further study the role of mtDNA 3397A/G and 3552T/A variations of reduced nicotinamide adenosine dinucleotide dehydrogenase 1 in HAPE susceptibility.
Methods.—
The single-nucleotide polymorphisms of mtDNA 3397 and 3552 in patients with HAPE (n = 132) and their matched control subjects (n = 233) were studied using polymerase chain reaction sequencing.
Results.—
The frequency of mtDNA 3397G in the HAPE group (2.3%) was significantly higher than that of the control group (0%; P = .021; odds ratio, 2.806; 95% confidence interval, 2.443 to 3.223). The frequency of mtDNA 3552A in the HAPE group (6.8%) also was significantly higher than in the control group (1.7%; P = .012; odds ratio, 4.198; 95% confidence interval, 1.264 to 13.880).
Conclusions.—
In this study, we present the first evidence of differences in mtDNA polymorphism frequencies between HAPE victims and healthy Han Chinese. Genotypes of mtDNA 3397G and 3552A were correlated with HAPE susceptibility. This result could contribute to defining the role of the mitochondrial genome in the pathogenesis of HAPE.
Keywords
Introduction
High altitude pulmonary edema (HAPE) results from a breakdown of the alveolar capillary membrane and the accumulation of a protein-rich fluid in the respiratory tract. This condition normally occurs at altitudes above 3000 m and affects many people including previously fit individuals. 1 –4 Advances in the understanding of the biochemical pathways involved in the physiologic response to hypoxia have led to greater insights into the pathogenesis of HAPE. Polymorphisms within certain genes may explain the individual variation in hypoxic responses and suggest the susceptibility to HAPE. These genes include nitric oxide synthase 3, cytochrome b-245, angiotensin-converting enzyme, surfactants A1 and A2, hypoxia-inducible factor-1 (HIF-1), major histocompatibility complex A, and the aldosterone synthase CYP11B2. 5 –10 However, the current crop of polymorphisms is not enough to provide a full genetic explanation for HAPE. We suspect that polymorphisms of other genes, such as genes encoded by mitochondria, 11 may play an important role in the HAPE susceptibility.
Mitochondria and mitochondrial DNA (mtDNA) play a crucial role in the response to hypoxia. In our previous study, we identified 15 novel mtDNA-encoded amino acid changes that had potential function in modulating oxidative phosphorylation (OXPHOS) function during cold and hypoxia adaptation in the plateau pika (Ochotona curzoniae). 11 Human mtDNA is a covalently closed, circular double-strand genome of 16 569 bp. 12 It is different from nuclear DNA (nDNA) because mtDNA is not protected by histones and locates very close to the inner membrane of the mitochondria. Therefore, mutation rates are much higher in mtDNA than in nDNA. 13 Some mutations in mtDNA may result in a reduction of mitochondrial ATP production capacity, whereas other variations may have subtle effects on oxygen-sensitive K+ and Ca2+ channels. 14
Previous studies have suggested that mtDNA mutations are associated with a variety of disorders, including diabetes 15 –17 and other metabolic defects. 18 One of the well-studied mutated genes is the mitochondria-encoded reduced nicotinamide adenosine dinucleotide (NADH) dehydrogenase 1 (ND1). ND1 mutation increases the reactive oxygen species (ROS) level,19,20 which, in turn, induces HIF-1 destabilization. 21 As a result, HIF-1α is unable to bind to HIF-1β during hypoxia, failing to form HIF and preventing adaptation to the hypoxic environment. We suspect that HAPE pathogenesis could be related to a mutation in ND1. 22 Mutations of ND1 are reported to be associated with a number of different diseases, including Alzheimer disease, Parkinson disease, mitochondrial myopathy, Wolfram syndrome, and maternal inherited diabetes. 23 –27 A major mutation of ND1 that is related to these diseases is mtDNA 3397A→G, which replaces methionine with valine at amino acid 31 in a highly conserved region of ND1. The mutation mtDNA 3552T→A, which is a synonymous homoplasmic mutation in the ND1 codon 82 position 3, was found in some patients with Leber hereditary optic neuropathy 28 and type 2 diabetes. 29 We speculated that those mutations may also be associated with HAPE. To test this hypothesis, polymerase chain reaction (PCR)-sequencing was used to compare the mtDNA sequences at positions 3397 and 3552 in HAPE patients and healthy control subjects.
Materials and methods
Patients and Control Subjects
HAPE patients and healthy individuals (control subjects) were recruited from the same ethnic group of Han in China in Lhasa, Tibet (3658 m). The HAPE patients consisted of 132 individuals (25.01 ± 10.70 years old) who had HAPE, as determined by the standard diagnostic criteria. 30 This included cough and dyspnea at rest, with pulmonary rales, cyanosis, and patchy shadows detected using chest x-ray. All patients with HAPE met the criteria at the onset of the disorder and recovered promptly after hospitalization. All the HAPE patients and control subjects were from southwest China. HAPE patients experienced the disease after traveling from low attitude (around 300 m) to high altitude (Lhasa, 3658 m) within 7 days. We also randomly selected 233 healthy individuals (21.80 ± 11.08 years old), matching the HAPE patients in age, gender, and ethnic group. These individuals were healthy after moving to Lhasa for at least 1 week, after which a venous blood sample was collected. Individuals in both groups were unrelated to each other (all individuals had no blood relation for at least 3 generations in this study) and had no known mtDNA-related diseases (such as diabetes, Parkinson disease, or Alzheimer disease) in their medical history. This study was approved by the ethical committee of the Third Military Medical University in China.
Isolation of DNA
The HAPE patients' blood was collected before any medical treatment. Venous blood (2 mL) was collected from both HAPE patients and healthy control subjects and placed in EDTA-anticoagulation tubes, then stored at −80 °C before the experiments. Genomic DNA was extracted from peripheral blood leukocytes from 1 mL of whole blood using Omega (Omega Bio-Tek, Inc, Norcross, GA) DNA extraction kits. Genomic DNA was tested using gel electrophoresis on 0.8% agarose gel stained with ethidium bromide.
Genotyping of mtDNA 3397 and 3552 Single-nucleotide Polymorphisms By PCR Sequencing
Single-nucleotide polymorphisms (SNPs) of mtDNA 3397 and 3552 were genotyped using PCR-sequencing. PCR primers were forward: 5′-GTCCTAAACTACCAAACCTGC-3′ and reverse: 5′-GTGTTAGTCATGTTAGCTTG-3′. Reactions were performed in a 50-μL volume using 50 ng of template DNA, 5 μM deoxyribonucleotide triphosphate (Takara Bio Inc, Shiga, Japan), and 20 pmol of the forward and reverse primers in total, together with 2.5 U of Taq DNA polymerase enzyme (Takara Bio) in its supplied Taq buffer supplemented with 75 μM MgCl2. PCR was performed using the PTC-100 thermal cycler system (MJ Research Inc, Waltham, MA). Program parameters were 1 cycle of 94°C for 5 minutes followed by 35 cycles of 94°C for 45 seconds, 56°C for 30 seconds, and 72°C for 3 minutes. After a final incubation at 72°C for 4 minutes, the products were cooled to 4°C. PCR products were tested using gel electrophoresis on 1% agarose gel stained with ethidium bromide. PCR fragments were purified with the QIAquick PCR purification kit (Qiagen, Inc, Valencia, CA) according to the manufacturer's protocol and assayed using a Nanodrop spectrophotometer (NanoDrop Technologies, Inc, Wilmington, DE). Five picomoles of the forward or reverse primer and 200 ng of purified PCR product were combined in a final volume of 16 μL. Sequencing reactions were performed using the Applied Biosystems Dye-Terminator Kit and analyzed on an ABI Prism 3730 DNA sequencer (Applied Biosystems, Inc, Foster City, CA). All PCR products were sequenced at Shanghai Sangon Biological Engineering Technology & Services Co (Shanghai, China). Primers used to sequence the mtDNA 3397 and 3552 were 5′-GTCCTAAACTACCAAACCTGC-3′.
Statistical Analyses
Statistical analyses were performed using the SPSS software (version 12.0, SPSS, Inc, Chicago, IL). The allele frequencies in the patients and the controls were compared using the Pearson χ2 test. We also used the Bonferroni correction to correct for multiple testing. Difference was considered significant if P < .025. 31 An odds ratio (OR) for each genotype, with its 95% confidence interval (CI), was calculated with the χ2 test.
Results
In this study, we compared the genotypes of mtDNA 3397 and 3552 from 132 HAPE patients and 233 healthy individuals. As shown in the Table and the Figure, all of the 233 healthy individuals had a genotype 3397A, whereas 3 of 132 (2.3%) of the HAPE patients had 3397G. The percentage appeared low, but Pearson χ2 test and Bonferroni correction showed that the difference was statistically significant (P = .021; OR, 2.806; 95% CI, 2.443 to 3.223). Another mutation we found that appeared to correlate with HAPE was mtDNA 3552T→A (see the Figure). Our results showed that the frequency of 3552A in HAPE (6.8%) was higher than that of the control group (1.7%; P = .012; OR, 4.198; 95% CI, 1.264 to 13.880).
The association between mt3397G, mt3552A genotypes and high altitude pulmonary edema susceptibility
CI, confidence interval; HAPE, high altitude pulmonary edema; OR, odds ratio.
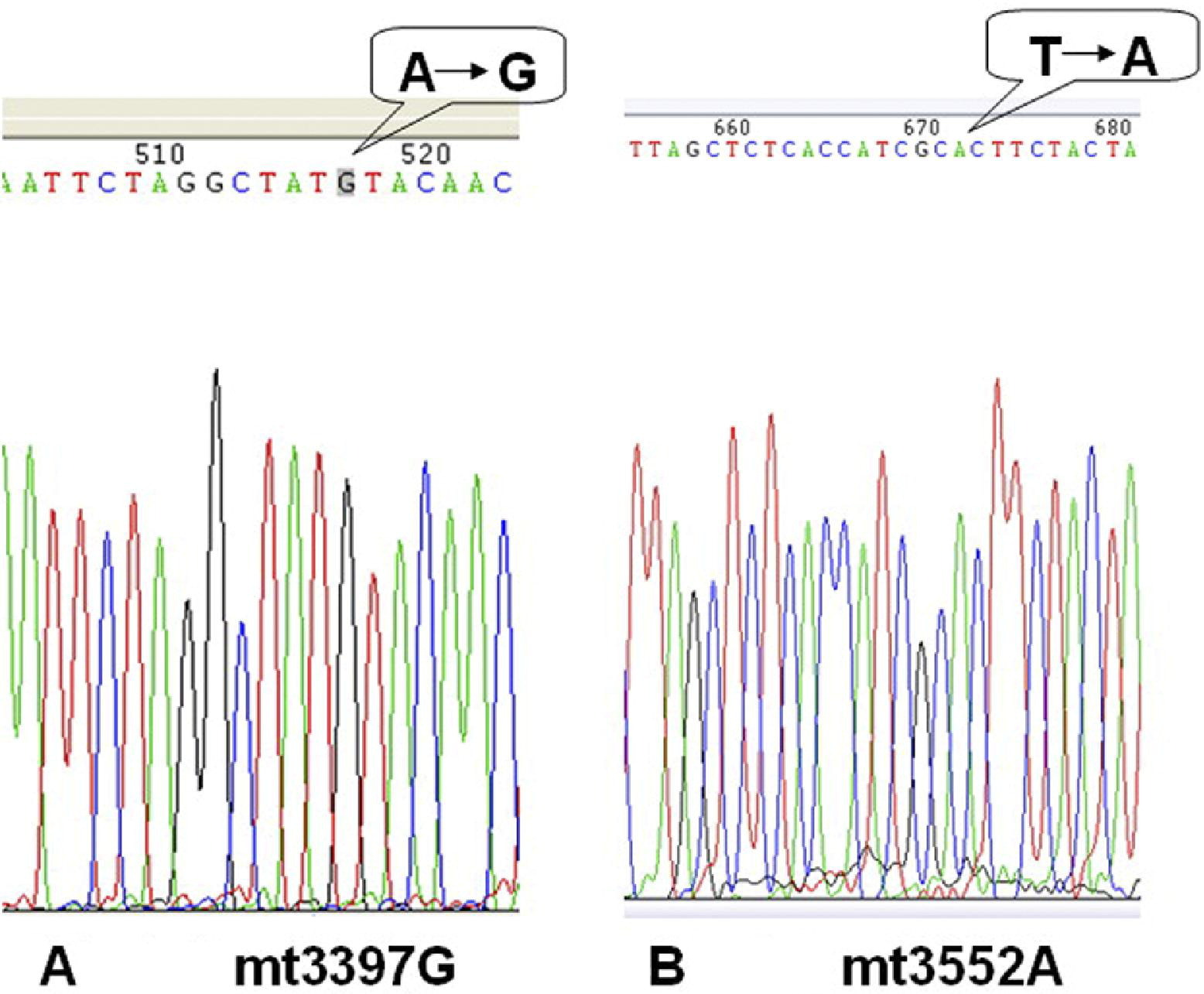
Single-nucleotide polymorphism sequences, showing the substitutions of 3397A→G and 3552T→A. At the mt3397 site, the A was replaced by G; at the 3552 site, the T was replaced by A.
Discussion
In this study we present for the first time evidence of differences in mtDNA polymorphism frequencies between HAPE patients and healthy Han Chinese. Mitochondria are the main site of OPHOS in cells. Previous studies showed that mtDNA-encoded cytochrome c oxidase and NADH dehydrogenase subunits 4 and 5 play an important role in hypoxic adaptation in plateau pika or Tibetan wild ass.11,32 Tibetans are well adapted to living and thriving in high altitude environments. Luo et al 33 found mitochondrial nt3010G–nt3970C haplotype is also associated with high altitude adaptation in the Tibetan population. It has also been reported that certain mtDNA mutations are related to individual tolerance to acute mountain sickness (AMS), or are risk factors for AMS among Han Chinese. 34 HAPE is a disease associated with poor acclimatization. Taken together, we believe that mtDNA polymorphism may play a role in the diseases related to high altitude.
mtDNA mutations have been shown to be associated with a variety of diseases. Little is known, however, about the relationship between the mtDNA variation and HAPE. The Human Mitochondrial Genome Database (
There were some limitations to this study. The sample size was small and drawn exclusively from a population that originated from southwest China. Larger samples from other populations would be useful to identify whether the association between mtDNA SNPs and HAPE is more widely found. The rarity of the SNP findings in HAPE patients underlines the complexity of HAPE. HAPE is likely to be caused by a much wider range of genetic and environmental factors than was studied here.37,38 Nevertheless, the study of SNPs in HAPE patients may help us elucidate the potential pathways involved in the pathogenesis of this life-threatening disease. In summary, using a strict case-control matching design, we found genotypes of mtDNA 3397G and 3552A were correlated with HAPE susceptibility.
Footnotes
Funding
This research was supported by the National Natural Science Foundation of China, National Science and Technology Ministry of China, and the 973 Project of China.
Acknowledgments
We are grateful to all the people who participated in this study.
Electronic-Database Information
Accession numbers and URLs for data presented herein are as follows: GenBank,
