Abstract
Despite great strides made in the past decades, the detection of microbial pathogens and their toxins in foods remains a challenging task. This is due primarily to several inherent difficulties associated with food analysis, that is, the complexities of food matrices (inhibitors and normal flora), the attributes of target analytes in foods (low level, heterogeneous distribution, and cell injury during processing), and the ratio between the amount of food samples and the detection assay volume. This review aims to provide an overview and a better understanding of the limitations, current applications, and future perspectives in terms of pathogen and toxin detection in foods.
Introduction
Microorganisms are the primary cause of food spoilage and foodborne illness. The ability to detect pathogenic organisms and their toxins serves as a cornerstone to ensure the safety and quality of our food supplies. Although not a solution, improved detection methods with better sensitivity and speed will be a valuable tool in defining the problems and outlining solutions. Numerous technologies have been developed to enumerate the total and groups of microorganisms and to detect and identify specific pathogens and toxins present in foods. 1 Traditional methods, to a large extent, depend on using suitable culture media. To detect specific microorganisms, which often are a small proportion of the total microorganisms present in food, selective media are used that enhance the growth of the target organism(s) and suppress the growth of the rest. Although progress has been made in the formulation of improved media for selective enrichment and identification, the process remains lengthy and labor intensive. Advanced technologies, including convenience-based, antibody-based, and nucleic acid-based assays, have revolutionized the detection methodology for microbial pathogens and toxins in foods. Often times, such assays not only identify the specific microorganism, but also provide a rather comprehensive genetic or metabolic profile of the organism. 2 In addition, the rapid evolving field of nanotechnology has provided its unique approach to pathogen and toxin detection in foods. Although significant challenges do exist, collaborative efforts among scientists with expertise in multiple disciplines should lead to major technology advancements and result in future generations of pathogen and toxin detection methodologies in foods.
Challenges
Detecting microbial pathogens and toxins in foods faces many inherent challenges associated with food analysis. There are a wide variety of food products, liquid versus solid, meat versus produce (fresh fruits and vegetables), animal carcass versus ground meat, and raw versus ready-to-eat foods. The complexity of food matrices and compositions presents a major obstacle to developing effective sampling, sample preparation, and testing strategies. 3 For example, efficient separation of target analytes (pathogens or toxins) from food matrices is rather difficult in meat, poultry, and egg products. In addition, food matrices frequently harbor inhibitors for key reagents used in detection assays, such as PCR enzymes. Furthermore, raw foods naturally contain high levels of normal flora, which may interfere with the performance of a wide array of pathogen detection technologies.
On the other hand, the attributes of target analytes in foods also significantly impact the detection efficiency. For example, unlike clinical samples, pathogens in foods are generally present at a much lower level and are frequently injured during food processing, which demands improved detection sensitivity and pre-enrichment to resuscitate the injured cells. Additionally, the heterogeneous distribution of pathogens and toxins in foods presents a challenge for effective and statistically sound sampling procedures, particularly when detecting pathogens that cause severe illness in humans with very low infectious doses such as Escherichia coli O157:H7. 4 One additional consideration for sampling is the type of sampling procedures to be used, destructive versus nondestruction, which can dramatically affect microbial recovery rates, hence the detection sensitivity and overall performance.
Another important challenge for detecting pathogens and toxins in foods is related to the ratio between the amount of food samples and the detection assay volume, which is particularly noteworthy in advanced technologies that only a tiny fraction of the original food samples is subjected to the actual detection assay after sample preparations. As most sampling and sample preparation procedures tend to accommodate liquids, dilution is generally needed for testing solid samples (e.g., 1:10, 1:100), which further reduces bacterial concentrations in foods. Table 1 shows an example of how this sample volume issue affects the ultimate sensitivity of PCR assays in comparison with culture-based assays. Given a one cell per reaction sensitivity for PCR, the deduced detection limit in that scenario is 1000 CFU/g, whereas the limit can be as few as 0.003 CFU/g when 325 g ground beef samples are collected and subjected to culture enrichment procedures.
Detection limits of assays as affected by sample size and assay volume

The detection sensitivity for PCR is presumed to be one cell per assay reaction, whereas it is also assumed that culture-based method can detect 1 CFU per plating event.
Because of these challenges, it is crucial that effective sample preparation strategies are implemented before they are actually subjected to the detection assay. Enrichment has been commonly used to overcome the problems associated with pathogen detection in foods. In addition to recovering injured cells, enrichment serves as a major strategy to increase the concentration of target pathogens in foods and thereby diluting the effects of inhibitors and normal flora on the testing. In DNA-based assays, enrichment has been applied to indicate the detection of viable rather than nonviable cells. 3 However, enrichment is labor intensive, and inevitably increases the total analysis time, stretching it to days rather than hours or even minutes. In addition to enrichment, concentration and extraction techniques are commonly applied to facilitate both pathogen and toxin detection in foods. 5 However, when food samples are homogenized and concentrated, components that are inhibitory to certain detection assays are often co-concentrated. 2 Furthermore, many techniques used to extract and purify pathogens and toxins in foods suffer poor recovery rates, resulting in reduced assay sensitivity and efficiency.
Current Procedures
Overview
According to a recent report “Food Micro—2008 to 2013” by the Strategic Consulting, Inc. (www.strategic-consulting.com), worldwide food industry microbiology testing includes 738.3 million tests in 2008, with a market value exceeding $2 billion. 6 This represents an increase of testing volume by 17.8% over the past 3-year period. Routine tests account for 81.3% of all food microbiology testing at 600.2 million, a growth of 16.1% since 2005. Testing for foodborne pathogens has grown even faster at 25.6% and now represents 138.1 million tests. 6 Conventional methods still account for approximately 60% of the microbiological food tests in 2008, although down from the 65% level in 2005. In contrast, rapid methods have increased by 36.8% from 2005, now accounting for 40% of all food microbial tests performed worldwide. 6
In the United States, food safety regulatory agencies including the U.S. Food and Drug Administration and the U.S. Department of Agriculture (USDA) have each developed and validated detection methods for their internal use. These methods combine several techniques in concentration, screening, and identification and are available online at Bacteriological Analytical Manual (www.cfsan.fda.gov/%7Eebam/bam-toc.html; 2001 Edition) and Microbiology Laboratory Guidebook (www.fsis.usda.gov/Science/Microbiological_Lab_Guidebook/; last updated February 2008). Validations for detection methods for foodborne pathogens and toxins are conducted by independent organizations such as the Association of Official Analytical Chemists International (AOAC; www.aoac.org). The AOAC method validation programs include Official Methods of Analysis Program and Performance-Tested Methods Program.
Sample Preparation
A complete analysis for pathogens and toxins in foods encompasses three components: sampling, sample preparation, and detection assay. Even though running the actual assay may take minutes to hours, the complete analysis can sometimes take days. As pathogens are usually represented in low numbers in food products, the processing of large sample volumes is often required for effective detection. Several methods of concentrating and separating bacteria from food matrices have been used to address the problem, including physical separation such as centrifugation and filtration; chemical separation such as metal hydroxides, resins, and lectins; and immunomagnetic separation (IMS) using magnetic beads coated with antibodies to specific pathogens as reviewed by Stevens and Jaykus. 5 In addition, pathogen-testing methods currently in use typically use a one- or two-stage broth enrichment step to increase the target pathogen to levels that can be detected by subsequent steps in the method. After enrichment, a rapid screening test such as enzyme-linked immunosorbent assay (ELISA) and PCR may be used to expedite determination of samples that are negative. Use of combinations of methods has many advantages to achieve the required test sensitivity and specificity and some degree of rapid detection. 3
Conventional Culture Methods
Generally regarded as the “gold standard” for food detection, traditional methods for microbial analysis of foods require multiple steps of incubations (pre-enrichment, selective enrichment, and plating on differential and selective agars) followed by biochemical, serological, or molecular tests for confirmation and further characterization. 3 The hallmark of such conventional pathogen detection methods is the use of suitable selective and differential culture media. Selective media are primarily used to enhance the growth of the target pathogens to detectable levels and simultaneously suppress the growth of the rest, which may interfere with the target detection. Differential media, on the other hand, rely on specific biochemical traits possessed by the target organisms, which are usually reflected on the color change of the growth media. Progress has been made in the formulation of improved media for selective enrichment and identification, for example, USDA recently revised the procedures used for the identification of E. coli O157:H7 by using an improved enrichment broth, mTSB + n (modified tryptone soy broth with novobiocin and casamino acids) to replace the mEC + n broth. 7
Rapid Methods
Rapid methods, including convenience-based, antibody-based, and nucleic acid-based assays, have revolutionized the detection methodology for microbial pathogens and their toxins in foods. Examples of convenience-based assays include hydratable media gel cards such as Petrifilm (3M Microbiology, St. Paul, MN) and portable device for testing ATP activities used in food industry for sanitation control. It is noteworthy that while not a direct indicator of the microbial contamination of food products, ATP testing does provide immediate feedback on the sanitation conditions of food contact surfaces and the environment. As a result, currently over 37% of all food plants worldwide have ATP systems and over 30 million tests are performed annually. 6 Additionally, specialty substrate media, such as chromogenic agar, have been developed and used to test groups of microorganisms such as coliforms and E. coli, as well as presumptive identification of specific foodborne pathogens such as E. coli O157:H7, 8 Listeria monocytogenes, 9 and Vibrio vulnificus. 10 The use of miniaturized biochemical assays such as API (bioMérieux, France) is another widely used convenient technique for pathogen identification. Table 2 lists several convenience-based detection assays as examples. Additional information is available in a book chapter by Feng for miniaturized and automated identification kits for bacteria and commercial available assays and specialty substrate media for detection of foodborne pathogens with their AOAC status. 3
Examples of rapid methods for detecting bacterial pathogens and toxins in foods (based on information from reference 3)
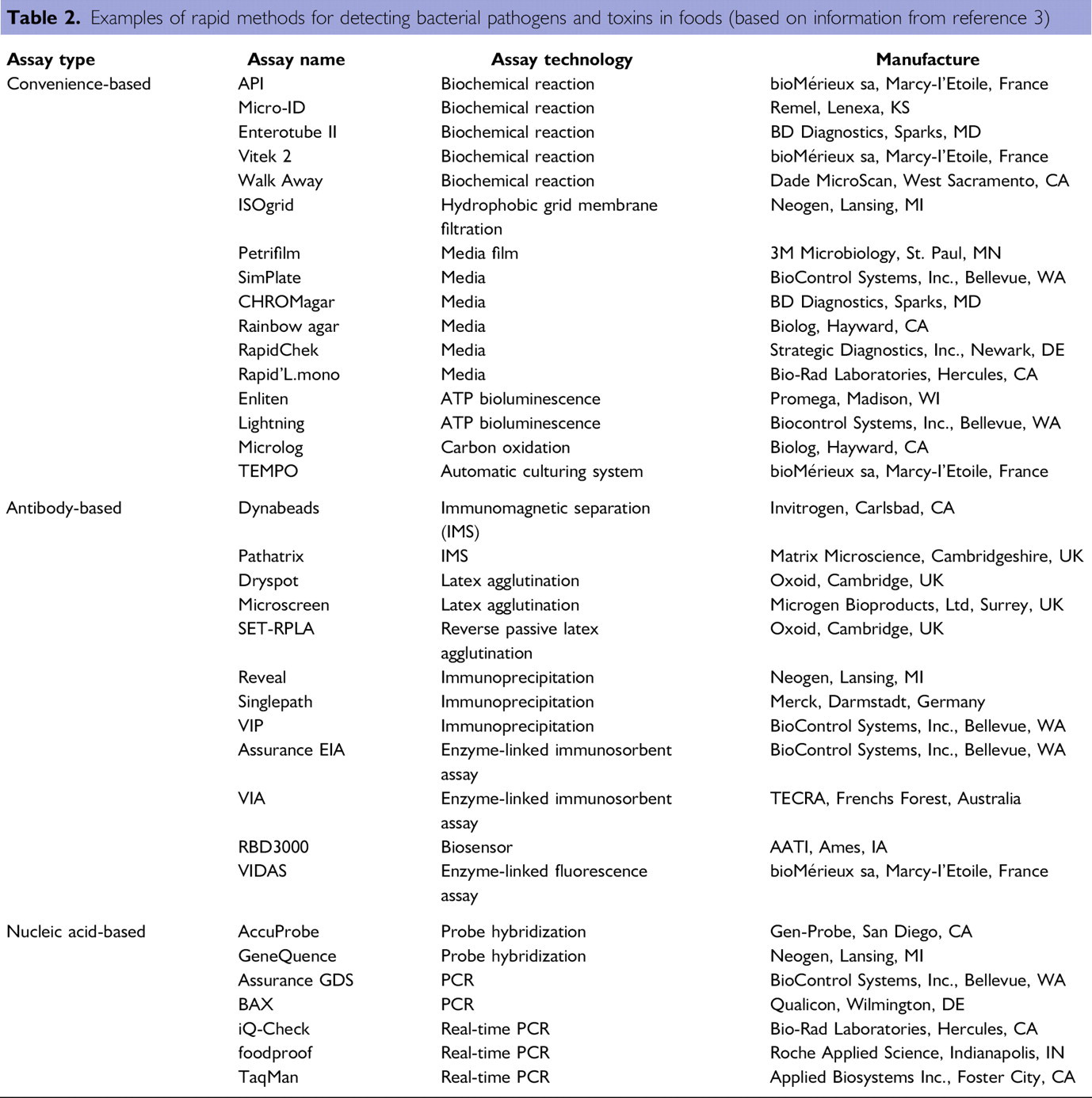
Antibodies that recognize bacterial pathogens have been used in the direct identification of foodborne pathogens or to aid in detection by separating microbial cells from the food matrix, that is, IMS. Bacterial pathogens can be concentrated before culture using IMS. The captured pathogens may then be detected by PCR and other technologies. Because much of the matrix is removed during the IMS, the resulting concentrate has lower levels of inhibitors and is also of smaller volume, both factors serving to improve the sensitivity of PCR-based tests. Additional platforms of antibody-based assays for detecting pathogens and toxins include latex agglutination, ELISA, and biosensor (Table 2).
Nucleic acid-based assays consist of two main types, hybridization using probes and amplification by PCR and related techniques. The recent development of microarray technology offers a powerful platform for probe hybridization where millions of probes can be deposited and analyzed simultaneously. Methods linking PCR detection to samples enriched for pathogen proliferation (usually overnight) are available for the majority of foodborne pathogens. For example, the USDA method for detecting Salmonella combines enrichment and PCR screening, resulting in reduced numbers of samples for further testing and increased testing sensitivity. 11 Design of improved media formulations that would facilitate “flash” enrichment might further decrease the time-to-test results. However, PCR is adversely affected by potential inhibitors in the food matrices, therefore it is critical that when designing PCR assays for food testing, internal amplification controls are included. 12 Real-time PCR provides near instantaneous amplification and detection at the same time. Real-time PCR is also meritorious for the capability to quantitatively identify foodborne pathogens in foods when no enrichment is required. Table 2 includes a selected list of probe, PCR, and real-time PCR-based detection assays. In addition to PCR assays used for the amplification of nucleic acids, some isothermal amplification techniques have been developed and applied in bacterial and viral detections in foods, which include strand displacement amplification, 13 nucleic acid sequence-based amplification, 14 loop-mediated isothermal amplification, 15,16 and ramification amplification. 17
One major limitation of DNA-based molecular detection assays is the inability to differentiate live versus dead cells. Recently, a technique incorporating either propidium mono-azide or ethidium bromide monoazide during the sample preparation steps was reported, 18 20 in which chemicals selectively intercalate into free DNA (in the case of dead cells) but not DNA in intact live cells, therefore preventing the amplification of dead cell DNAs. Such techniques offer great promise in inhibiting the amplification of dead cell DNAs by PCR or real-time PCR in complex samples. 18 20
It is noteworthy that when used for detecting pathogens and toxins in foods, the rapid methods described above are used primarily as a screening tool, and negative results are regarded as definitive but positive results are regarded only as presumptive, which requires further confirmation by traditional culture-based methods for pathogens, and the validation of toxin productions rather than merely the presence of toxin genes. 3 Additionally, for the majority of antibody-based and nucleic acid-based assays, enrichment is almost always needed to achieve the necessary sensitivity required of food testing and simultaneously diluting inhibitors in foods, which results in a total analysis time of several days rather than hours or minutes.
Nanotechnology
The recent advent and rapid evolving field of nanotechnology has provided a unique approach to pathogen and toxin detection in foods. For example, a novel method has been reported to incorporate dye molecules into a nanostructure to create dye-doped silica nanoparticles. 21 When conjugated with antibodies for E. coli O157:H7, these immunofluorescent nanoparticles were capable of detecting 1 CFU/g of this bacterium in ground beef within 20 min. 21 In addition, the coupling of nanotechnology with microfluidic systems has been successfully applied in molecular biology and can be adapted to detecting pathogens and toxins. A good example is the micro-PCR where 40 cycles of PCR can be performed in less than 6 min, 22 significantly reducing the total analysis time. However, the possibility of clogging microfluidic system by food particles is a major concern for such technologies.
Advanced Technology Application Examples
This section discusses three examples of advanced technology applications. One is using a combination of PCR and oligonucleotide-labeled nanoparticle detection system (Luminex, Austin, TX) for molecular serotyping of Salmonella used by the Centers for Disease Control and Prevention. 23 24 The second one is the utilization of an isothermal amplification technology, transcription-mediated amplification assay (TMA) for the rapid detection of Listeria genus, Salmonella genus, and Campylobacter genus developed by Gen-Probe (San Diego, CA). 25 The third one is using a liposome-PCR assay for the ultrasensitive detection of biological toxins. 26
In the first application, PCR primers are designed to specifically target several distinct flanking sequences of the genes that code for Salmonella O antigen groups and two phases of the H antigen. Two tiers of multiplex PCR assays are conducted for either the O group or H antigen. The second step is to hybridize the PCR products to antigen-specific labeled bead sets and then detect the fluorescence of the beads on a Bio-plex platform (Bio-Rad, Hercules, CA). The Bio-plex uses the Luminex multianalyte profiling (xMAP) technology (www.luminexcorp.com), which uses up to 100 different bead sets each internally labeled with different proportions of red and infrared fluorophores, resulting in up to 100-plex analyte detection. This application provides an easy-to-use, high–throughput system for the rapid detection of Salmonella O serogroups and H antigen types.
For the second application, food samples are first lyzed to release target analytes (ribosomal RNA), which are captured using poly-A-linked specific probe, and subsequently purified from the food matrix using magnetic particles coated with poly-T oligomers. During the second step, the purified rRNA samples are amplified using real-time TMA with fluorescence dye-labeled molecular beacons for the analyte and the internal control, respectively. The TMA technique involves the sequential use of reverse transcriptase (formation of cDNA) and the RNA polymerase (synthesis of RNA) to achieve multiple copies of RNA amplification products. The realtime TMA is rapid, highly sensitive, specific, and isothermal, with billions of target RNA amplified at 42 °C for 75 min. The detection system has shown great promise in detecting Listeria, Salmonella, and Campylobacter from multiple food samples. 25
The third application involves the ultrasensitive detection of biological toxins using a liposome—PCR assay. 26 The assay adopts a familiar sandwich ELISA format. Firstly, the target analyte (biological toxins) is captured by toxin-specific antibodies coated inside the wells of a microtiter plate. Liposomes that are conjugated with toxin receptors on the surface and embedded with reporter DNA within are then added to specifically bind the toxin and form a sandwich structure. The liposomes are then ruptured and a realtime PCR is used to quantify the released reporter DNA, which correlates with the amount of target toxins bound to the liposomes. Therefore, the detection assay is quantitative. Evaluation of the assay with cholera and botulism toxins showed a greater sensitivity than current detection methods. 26
Future Perspectives
An ideal detection system should include high specificity and sensitivity; fast response time; capability for mass production; elimination or simplification of sample preparation steps; minimal perturbation of sample; and providing continuous data analysis. Much progress has been made for the last decades, including automation and high throughput for sample processing and testing. However, there is a lack of real “real-time” procedures from sampling to results. Culture enrichment step that takes most of the testing time has not been eliminated and is likely to stay due to the several advantages discussed above. Design of improved media formulations that would facilitate “flash” enrichment is needed to decrease the time-to-test results. Alternative concentration technologies other than enrichment need to be developed to achieve the goal of real-time testing. Therefore, developing improved universal sample preparation method is critical toward the next generation of pathogen and toxin detection methodologies in foods. This requires collaborative efforts among chemists, engineers, molecular biologists, microbiologists (clinical, environmental, and food microbiologists), and food scientists.
In addition, desirable features such as quantification and multiplex detection are becoming increasingly important. Technologies that are routinely used in the chemical and physical testing have adopted for microbial testing in foods, such as mass spectrometry (MS) 27 and optical scanning technology. 28 Again, collaborative efforts among scientists with expertise in multiple disciplines should lead to major technology advancements and result in the next generation of pathogen and toxin detection methodologies in foods.
