Abstract
This technical paper describes the utilization of a new automated liquid handler from Beckman Coulter, Inc., the Biomek® NX Laboratory Automation Workstation, for genomic and proteomic applications. For genomic applications, methodology for plasmid DNA purification using Promega Wizard® SV 96 reagents was developed for the Biomek NX. A single plate of bacterial pellets can be processed to purified plasmid DNA without user interaction after initial setup. DNA quantity and quality were assessed by spectrophotometric analysis, restriction digestion, PCR (The PCR process is covered by patents owned by Roche Molecular Systems, Inc., and F. Homan La Roche, Ltd.), and capillary sequencing. Additionally, the plasmid preparation method was used to purify plasmid DNA from bacterial clones isolated in a bacterial two-hybrid screening procedure. In this case, the system quickly and efficiently prepared clones for rapid identification of target sequences. For proteomic applications, His-tag proteins were purified from bacterial cultures in a 96-well plate format. Following purification, a Bradford assay was used to determine the quantitative yields of the His-tag protein products in each of the aliquots from the purified samples. The AD 340 Automated Labware Positioner (ALP), an integrated absorbance reader, was used for absorbance measurements in the Bradford assay. Given the placement of this ALP on the deck of the Biomek NX, the entire process of protein purification and quantitation was performed in a complete walk-away automated format. Results obtained when purifying proteins, from both uninduced and induced bacterial cultures, on the worksurface of the Biomek NX will be described. (JALA 2004;9:177-84)
Keywords
Materials and methods
Biomek NX Hardware
A method for the purification of nucleic acids was developed on a stand-alone, 96-Multichannel Biomek NX. The method for the purification and quantitation of proteins was developed on a standalone Biomek NX with a Span-8 and Gripper (Figure 1). The Biomek NX uses the same ALP architecture as the Biomek® FX. Static ALPs as well as active ALPs, such as the Orbital Shaker and AD 340 ALP, can be utilized on the deck of the Biomek NX.

The Biomek NX Laboratory Automation Workstation.
Software Used on the Biomek NX
Biomek® Software, originally developed for the Biomek FX, is used to control the Biomek NX. It is common software architecture among Beckman Coulter's Laboratory Automation Workstations (Biomek 3000®, Biomek NX, and Biomek FX). Software is designed to provide the user with the capability to define critical variables in the process, establish version control, and track samples. A view of the plasmid purification method is shown in Figure 2.
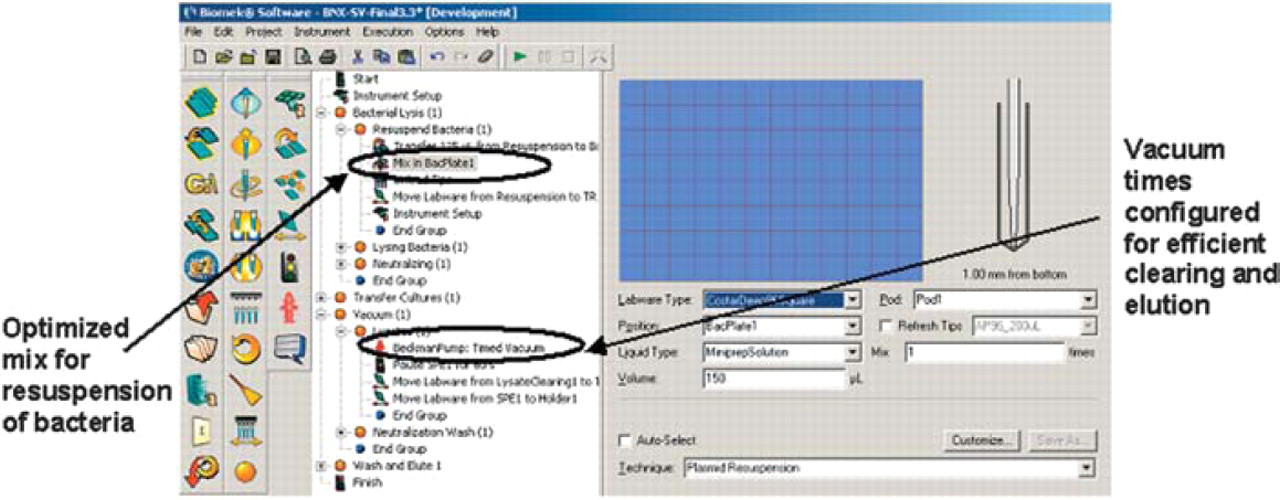
The plasmid preparation method on the Biomek NX.
Overview of Plasmid Preparation
Plasmid DNA is purified using Promega's Wizard® SV 96 (Wizard SV 96 Plasmid DNA Purification System, Technical Bulletin 272, Promega, Inc., Madison, WI, October 2002) reagents.
Briefly, bacterial pellets are resuspended, lysed, and transferred to filter plates after neutralization. The first filter plate (Lysate Clearing Plate) is used to trap cellular debris whereas DNA is bound to the second filter (DNA Binding Plate). After the Lysate Clearing Plate is discarded, the DNA Binding Plate is washed to remove impurities and dried. Purified plasmid DNA is eluted from the filter with molecular biology grade water.
Biomek NX Deck Setup for Plasmid Purification
The deck used for plasmid purification is shown in Figure 3A. A vacuum filtration manifold is affixed to the deck on the SPE site. The holder ALP is used for the assembly and disassembly of the collar and filter plates. The plasmid purification method on the Biomek NX will purify a single 96-well plate of bacterial pellets. The method has been optimized to purify high quality plasmid DNA. Critical steps in the process such as bacterial pellet resuspension and vacuum times have been optimized to deliver consistent results and minimize contamination. The starting instrument setup for Wizard SV 96 plasmid purification is shown in Figure 3B.
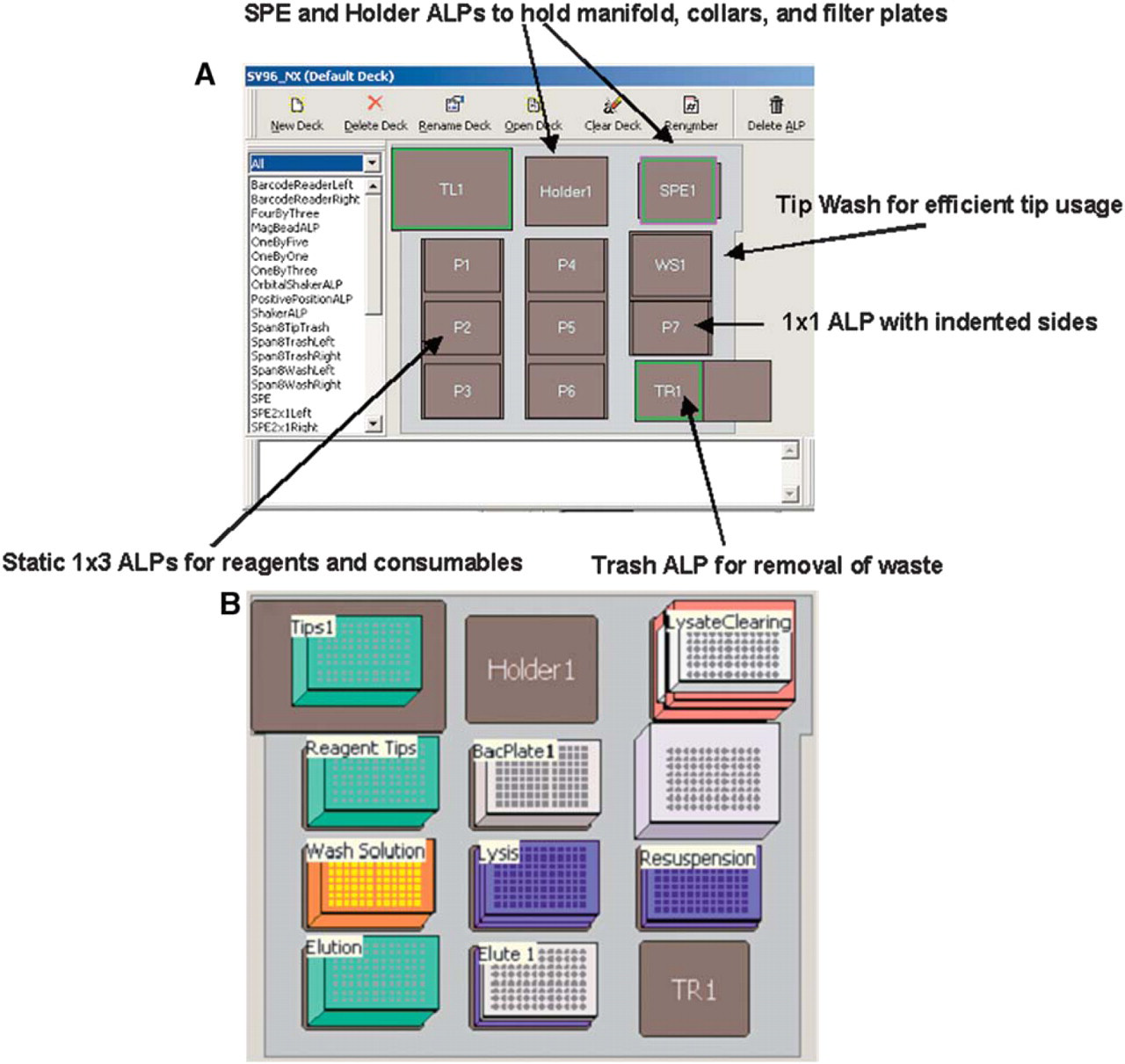
(A) The ALP positions on the Biomek NX for plasmid preparation. (B) Starting instrument setup for plasmid preparation.
Overview of Purification and Quantitation of His-Tagged Proteins
The purification and quantitation method can be used to purify up to a single plate of samples using Sigma-Aldrich's HIS-Select iLAP™ HC Nickel-Coated 96-well plates (Catalog No. H9412) from Sigma-Aldrich, Inc. (St. Louis, MO). An overview of this procedure is shown in Figure 4. Using the default settings, the entire process, of purifying proteins to getting quantitation data, takes approximately 4.5 hours. The method initiates with transferring bacterial cultures for lysis (Figure 5A). Quantitation and standard-curve plates are read on the deck using an integrated AD 340 ALP (Figure 5B). The gripper moves the plates to be read in the tray of the ALP and retrieves the plates once read. The protocol to read the plates (Bradford) is generated and saved within the method. The data are saved both within the reader software and in a .csv file as configured in the “Finish” step of the method.
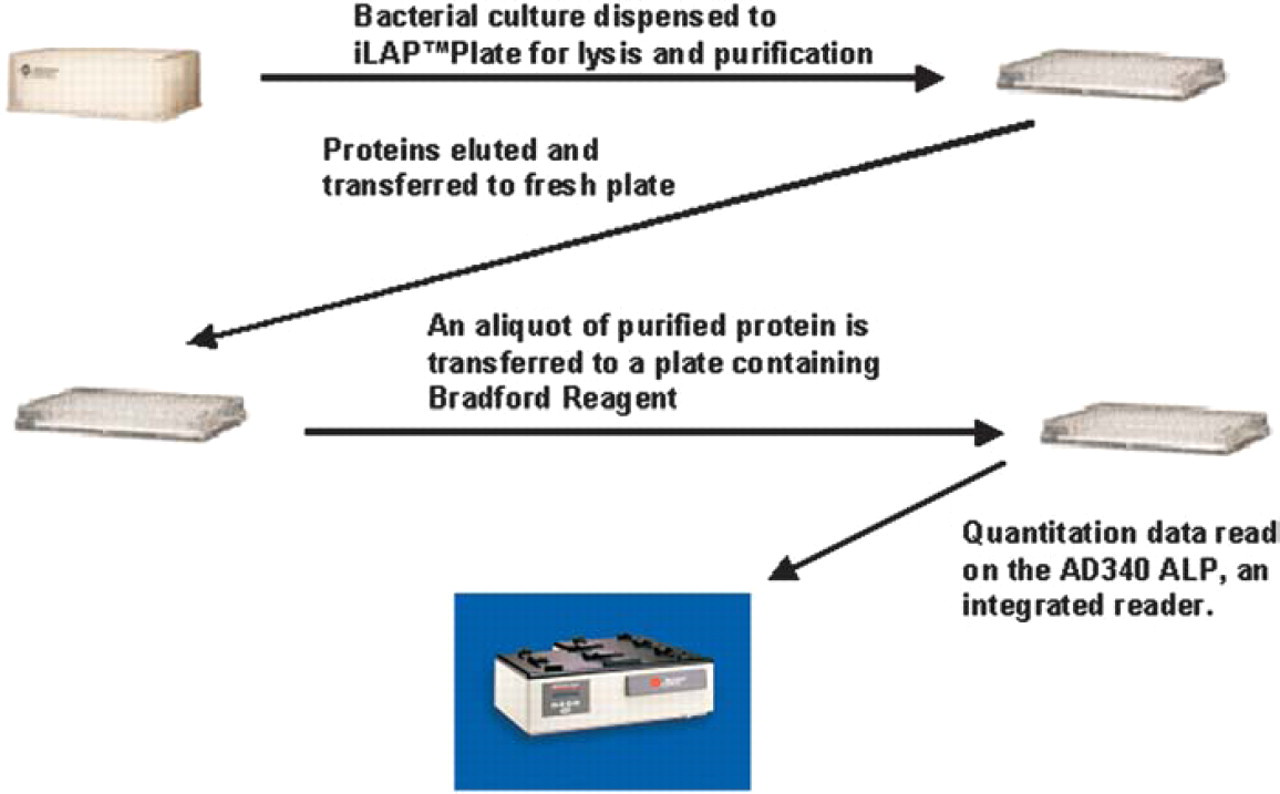
Method overview for protein purification and quantitation.
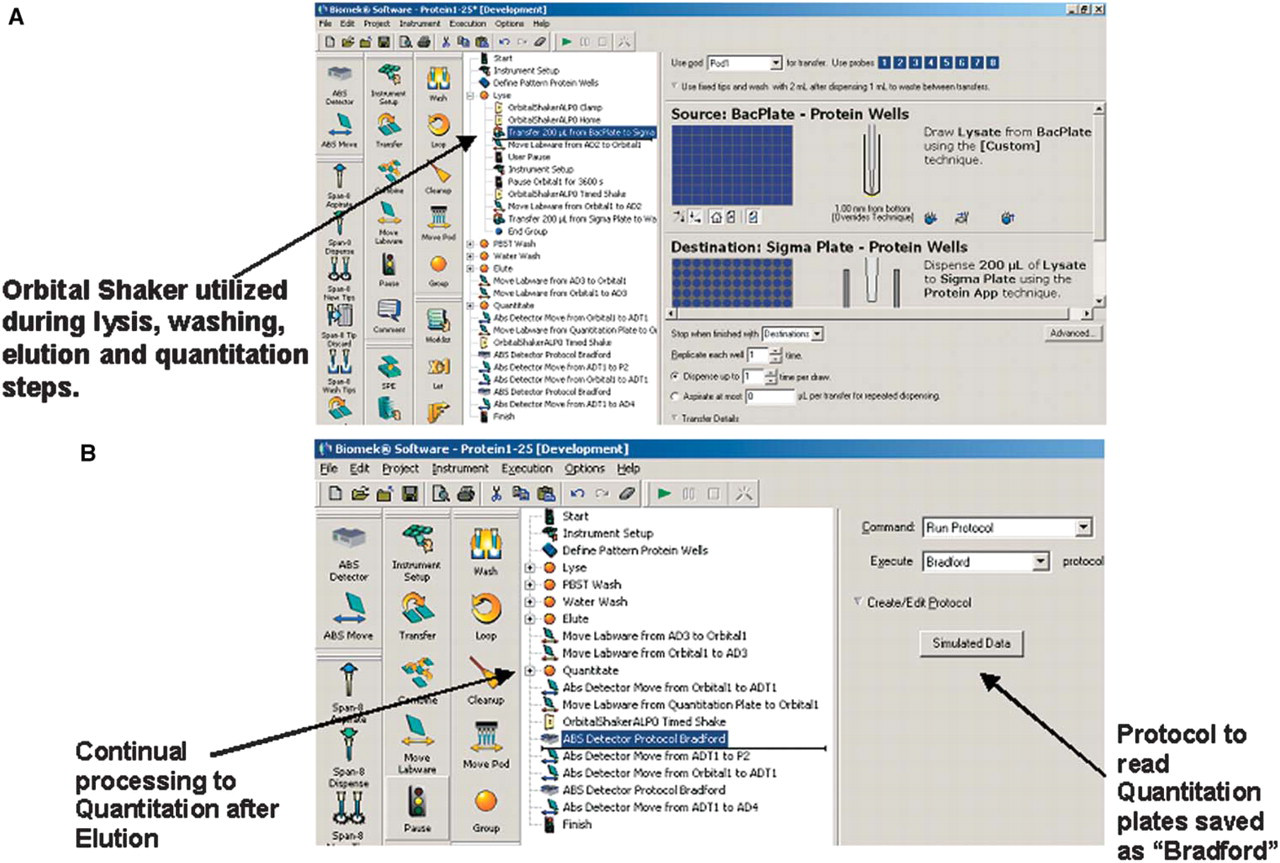
(A) A portion of the protein purification method. (B) Highlights of the quantitation portion of the method.
Biomek NX Deck Setup for Protein Purification
The deck used for the purification and quantitation of His-tagged proteins is shown in Figure 6. An AD 340 ALP, an integrated absorbance reader, is located to the back left of the deck and is used to read the final quantitation plates. An Orbital Shaker ALP is located in the back right corner of the deck. This ALP is used during the lysis and elution steps of the purification process to improve the yield of purified protein.
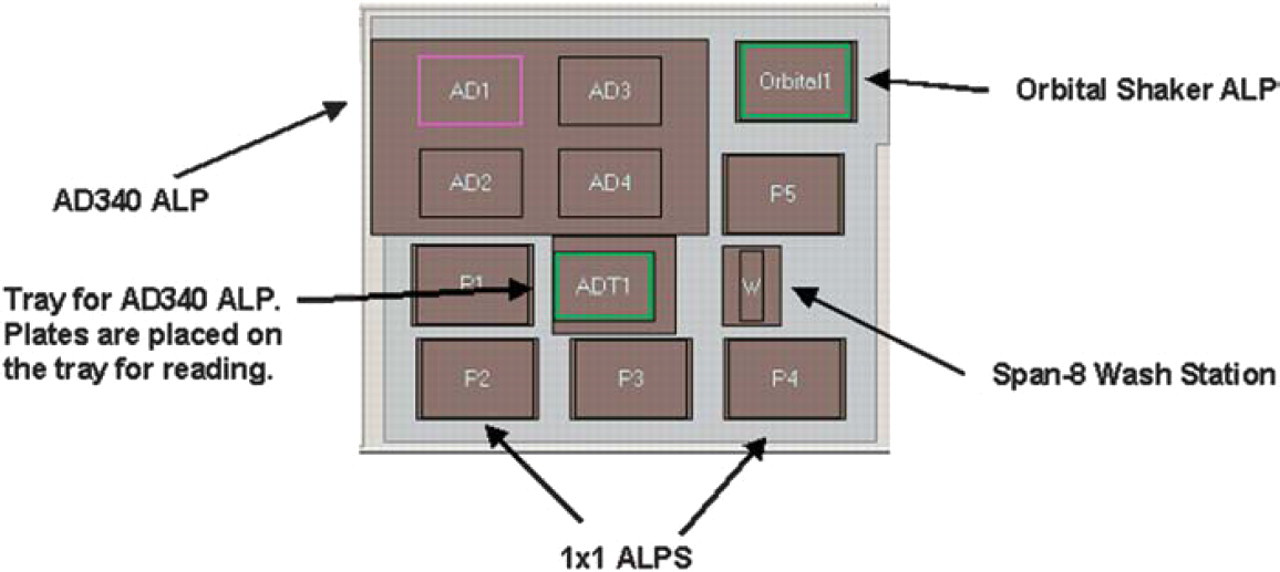
The ALP positions on the Biomek NX for protein purification and quantitation.
Starting Instrument Setup for Protein Purification and Quantitation
At the start of the method, all reagents needed to purify and quantitate proteins are located on the deck of the Biomek NX (Figure 7). After the bacterial cultures are transferred to the HIS-Select iLAP plate, the user is prompted to remove the bacterial plate to storage preventing it from being open to the surroundings for the duration of the method. The process initiates with the transfer of 200 μL of bacterial culture to a HIS-Select iLAP plate. The plates are incubated for at least 2 hours with shaking at room temperature or overnight at 4°C to lyse the cells and bind the recombinant protein to the plate. After washing the plate, proteins are eluted into phosphate-buffered saline (PBS), 300mM NaCl, and 250 mM imidazole. Purified samples are quantitated using Bradford Reagent.
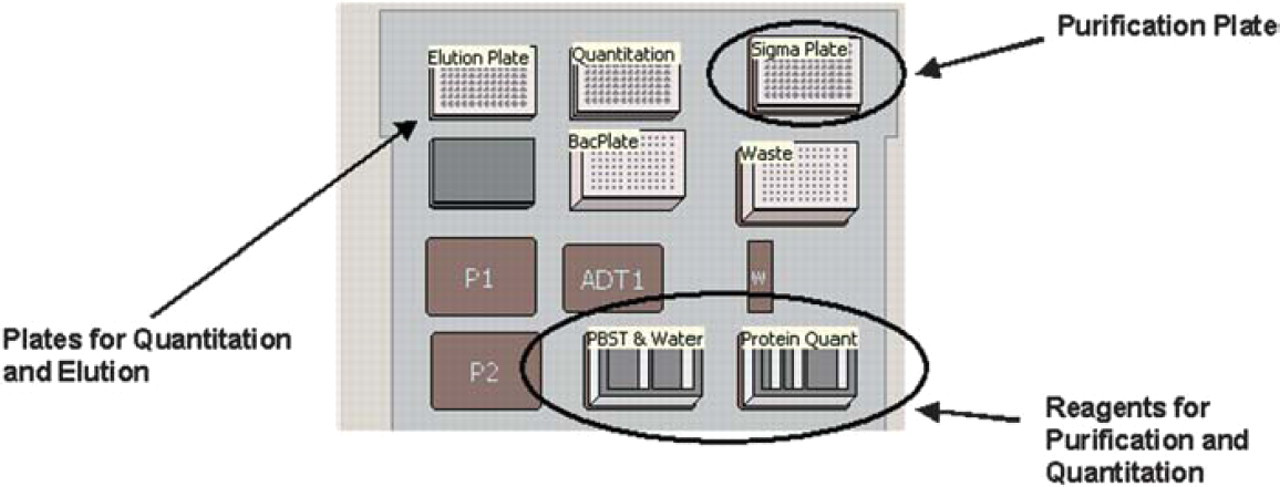
Starting instrument setup for protein purification and quantitation.
Clones Used for Protein Purification
The plasmid DNA used for the protein expression studies included a His-tagged lacZ and His-tagged human IL-8 clone. Expression of IL-8 in bacteria interfered with the growth of these cells, therefore, culture conditions and incubation times for purification were optimized to obtain detectable levels of recombinant IL-8.
Results for plasmid preparation on the biomek nx
Plasmid Yield and Quality
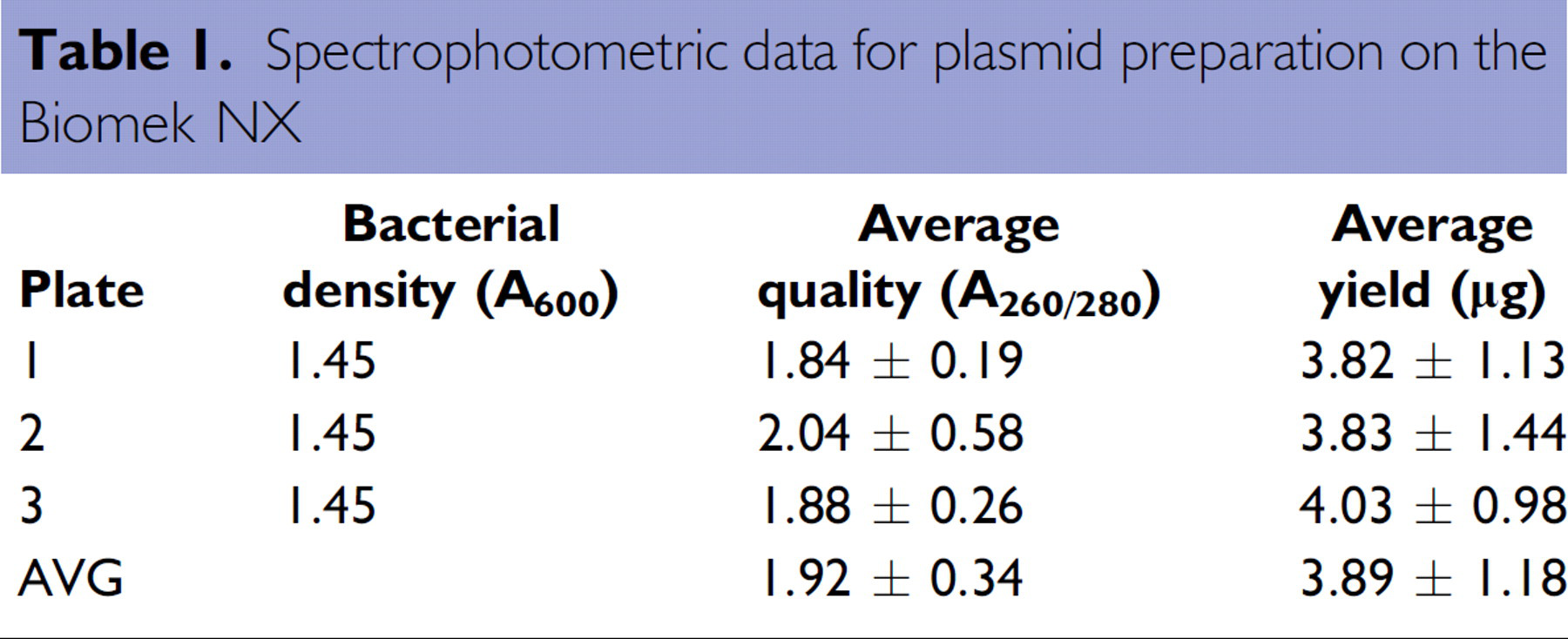
The spectrophotometric data shown in Table 1 demonstrate the quantity and quality of pGEM-3Zf(+) plasmids purified using Wizard SV 96 purification reagents on the Biomek NX. Overall yield was approximately 4 μg from a small biomass (approximately 1.5 OD600) with an average concentration of approximately 49 μg/mL using the default elution volume of 100 μL. The purity of the cultures was consistently excellent as indicated by the OD260/280 ratio.
Spectrophotometric data for plasmid preparation on the Biomek NX
PCR Amplification of Purified Plasmids
To assess the level of cross-contamination of plasmid DNA from bacterial cultures in adjacent wells, a PCR amplification-based assay was used. Bacterial cultures were seeded into every other well on a 96-well plate; this plate was processed on the Biomek NX. Eluted plasmid DNA was used in an amplification reaction with custom primers designed to generate a 752 base pair fragment. Samples were easily amplified and no product was observed in wells without bacterial cells indicating that contamination of adjacent wells did not occur during processing (Figure 8).
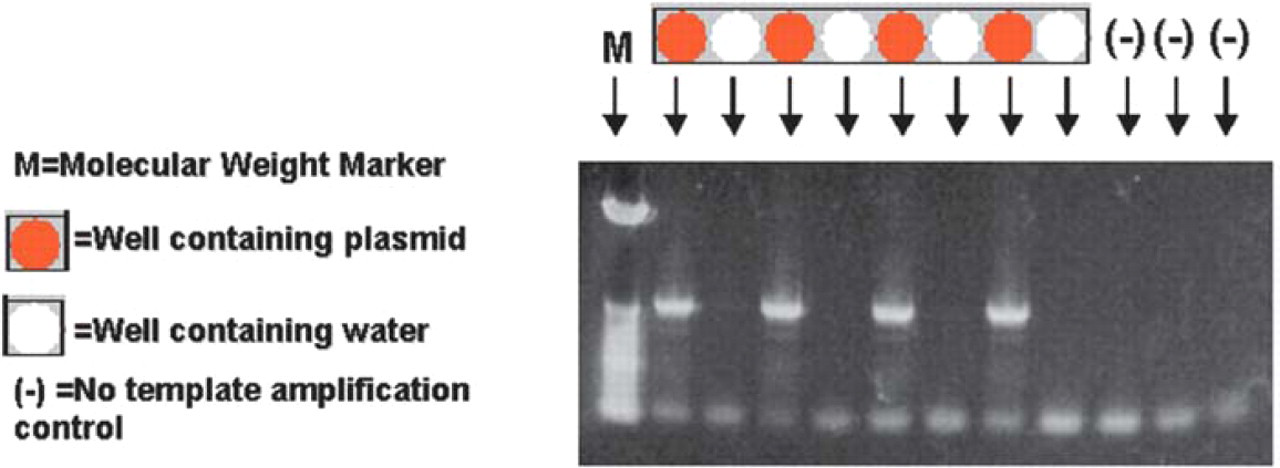
Agarose electrophoresis of PCR products from pGEM-3Zf(+). Sample order as indicated.
Restriction Digestion of Purified Plasmids
To demonstrate the suitability of the eluted plasmid DNA for typical downstream experiments, the purified pGEM-3Zf(+) plasmid DNA was subjected to restriction digestion. Three different enzymes, EcoRI, BglI, and PvuII, were used in the analysis (Figure 9).
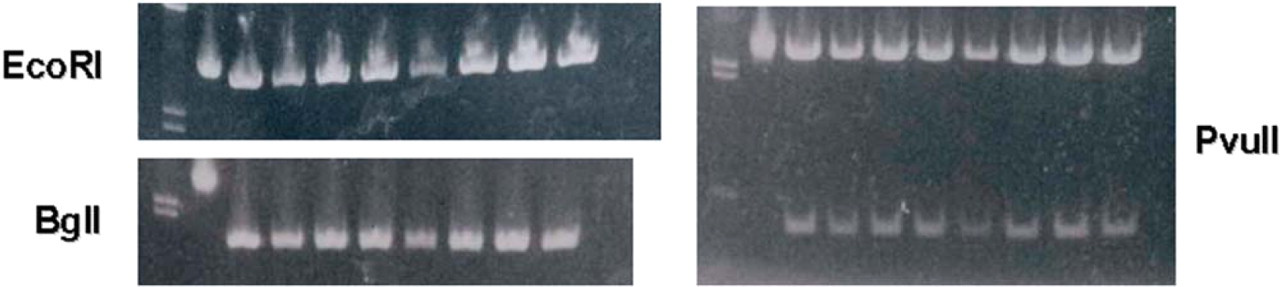
pGEM-3Zf(+) digested with enzymes as indicated. Lane order for all gels is: Lane 1=markers, Lane 2=uncut plasmid, and Lanes 3–10=digested plasmids.
Capillary Sequencing on Beckman Coulter's CEQ™ 8000 Genetic Analysis System
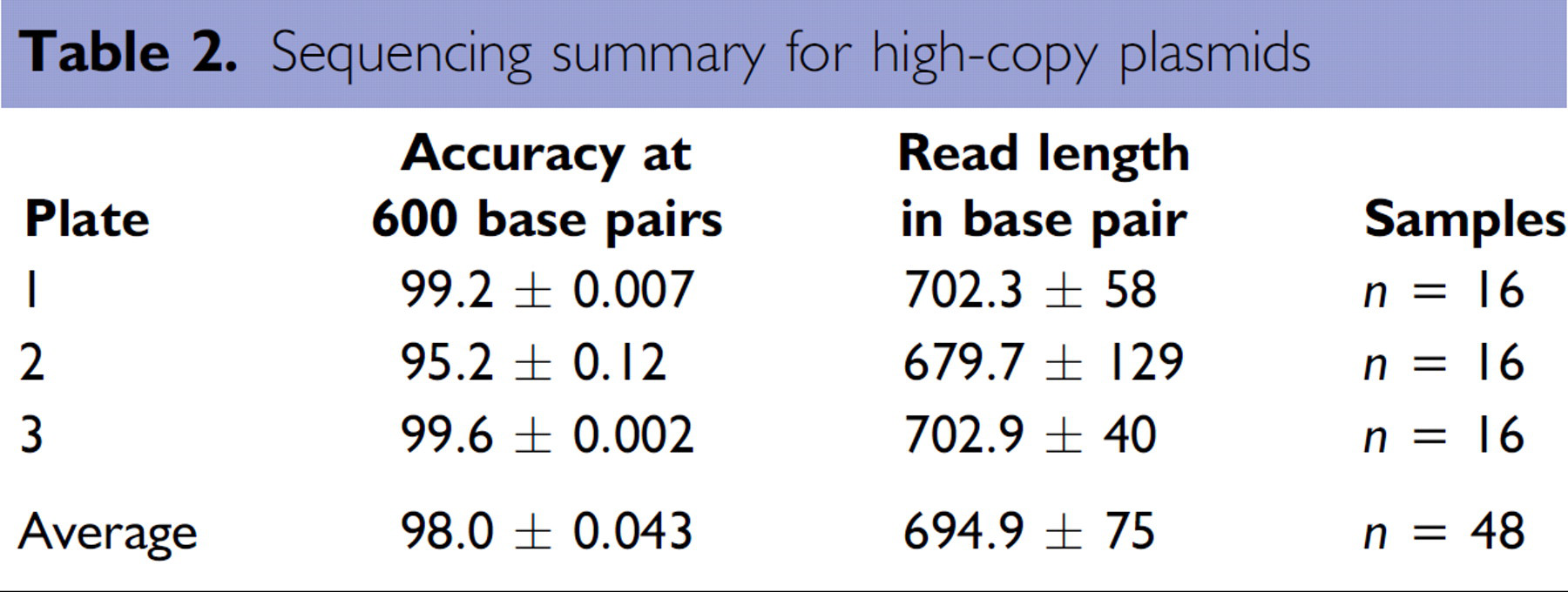
As further demonstration of the quality of the plasmid DNA generated by the Biomek NX method, the purified plasmid DNA was subjected to sequencing. Sequencing reactions and post-reaction cleanup were performed as recommended in the standard protocol (CEQ™ Dye Terminator Cycle Sequencing, P/N 608019-AK. Beckman Coulter, Inc., February 2002). Briefly, after plasmid DNA, primers, and sequencing premix were dispensed into a thermo well plate, the plate was sealed and placed in a thermocycler. After an initial denaturation for 2 minutes at 98°C, the reaction was amplified using 30 cycles of 96°C for 20 seconds, 50°C for 20 seconds, and 60°C for 4 minutes. After 30 cycles, the plate is held at 4°C until loading. The amplified products were loaded onto a Beckman Coulter CEQ 8000 Genetic Analysis System. A typical sequencing plot from plasmid DNA purified on the Biomek NX is shown in Figure 10. Composite sequencing data from random samples of plasmid DNA isolated from different plates using the Biomek NX are summarized in Table 2. The average read length obtained on the CEQ 8000 Genetic Analysis System was approximately 695 base pairs. The accuracy of the sequence data at 600 base pairs was excellent at approximately 98%.
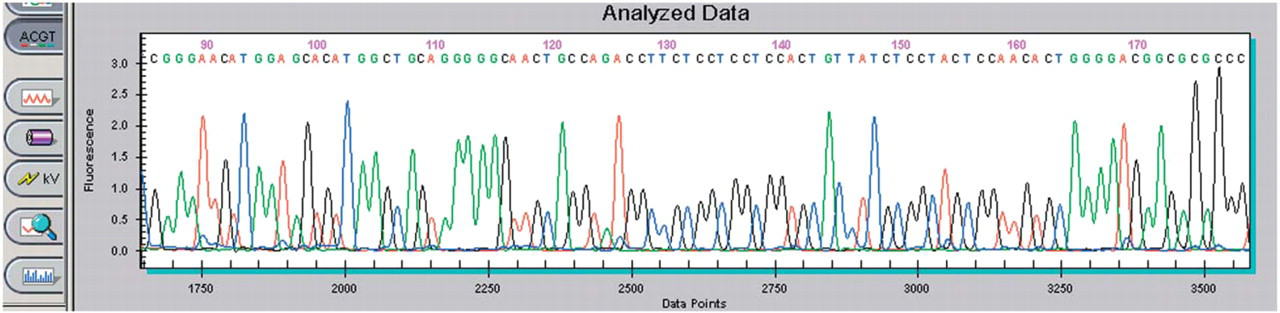
Sequencing data from purified pGEM-3Zf(+) sequenced on Beckman Coulter's CEQ 8000 Genetic Analysis System and analyzed with the analysis module.
Sequencing summary for high-copy plasmids
Plasmid Preparation for a Bacterial Two-Hybrid Experiment
Bacterial two-hybrid systems are a mechanism by which to identify protein-protein interactions. In this system, bait and target plasmids are co-transfected into bacteria and a colorimetric read out is used to identify colonies in which the bait and target proteins interact. These colonies are selected and cultured for DNA isolation and sequencing thereby identifying the target gene(s) of interest. To show the utility of the method and platform for performing experimentation beyond purifying high copy number plasmids such as pGEM-3Zf(+), bacterial colonies isolated during a bacterial two-hybrid screen were purified using the plasmid purification method for the Biomek NX. In order to accommodate the low copy plasmid in an endonuclease positive strain of E. coli: alkaline protease was added to the lysis buffer in a ratio of 40 μL alkaline protease per mL lysis buffer. To enhance the concentration of DNA in the final eluate, the vacuum time for clearing lysates was increased to 10 minutes and the elution volume was decreased to 60 μL. Purified plasmid DNA from 136 clones was used directly in a sequencing reaction. The eluted plasmid DNA was used directly in a sequencing reaction with CEQ Dye Terminator Cycle Sequencing reagents as follows. 6 μL plasmid DNA, 1.6 μM custom primer to the vector, and 12 μL DTCS premix were dispensed into thermo well plates. After an initial denaturation for 2 minutes at 98°C, the reaction was amplified using 30 cycles of 96°C for 20 seconds, 50°C for 20 seconds, and 60° C for 4 minutes followed by holding at 4° C. The amplified products were loaded onto a CEQ 8000 Genetic Analysis System.
Sequencing Results for Plasmid Isolated in the Two-Hybrid Experiment
Representative sequence generated from one of the two-hybrid clones is shown in Figure 11. Importantly, good quality sequence could be obtained with minimal changes to the methodology to accommodate purifying a low copy plasmid in an endonuclease-positive strain of bacteria. From the 136 samples purified and sequenced, results were obtained from GenBank® searches with the following definitions and summarized in Table 3. An isolate is defined as having significant homology when the majority of its sequence has homology to sequences in GenBank. A clone having isolated regions of homology to sequences in GenBank is defined as having limited homology. And, a clone having no homology or less than 25 base pairs of homology to sequences in GenBank is defined as having no significant homology.
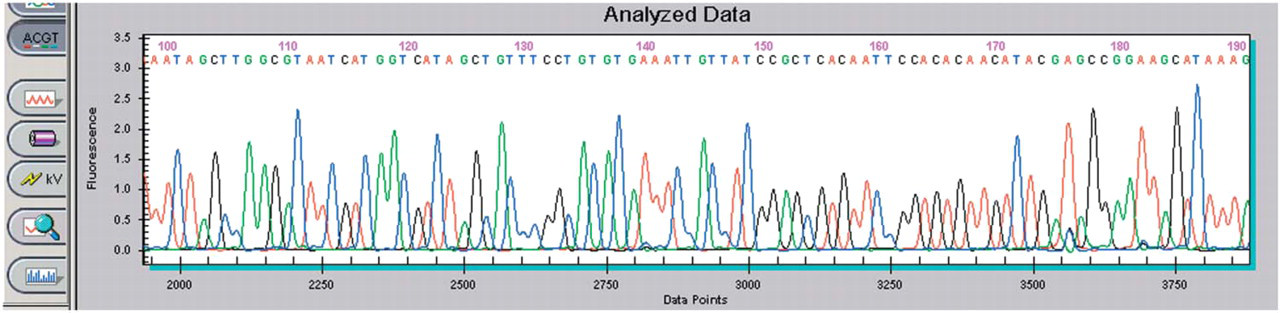
Sequencing data from a purified two-hybrid clone.
Summary of homology search for two-hybrid clones

Results for protein isolation and quantitation on the biomek nx
Cloning and Characterization of His-Tagged IL-8 Proteins
An expression plasmid was generated by cloning the coding sequence for human IL-8 into a bacterial expression plasmid (Catalog No. K100–01) from Invitrogen Corp. (Carlsbad, CA). To test for expression of the construct, individual colonies were isolated after transfection and grown in liquid culture. After induction by IPTG, cells were harvested by centrifugation, lysed in sample loading buffer and analyzed by SDS-PAGE. Figure 12 demonstrates the induction of protein in the bacterial system.
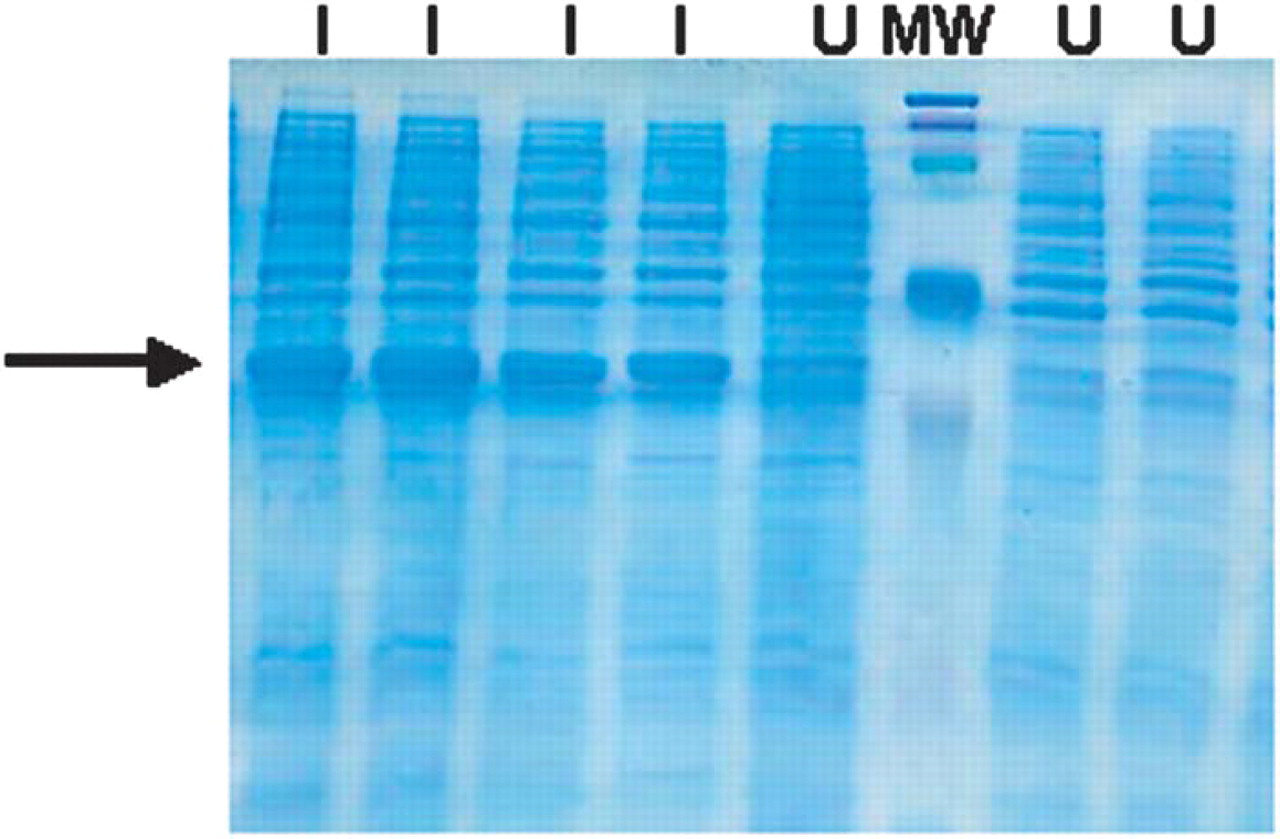
SDS-PAGE Analysis of bacterial lysates from induced and uninduced cultures. I = induced; U = uninduced; MW=molecular weight markers. Arrow indicates induced IL-8 protein.
Verification of Automated Protocol for Quantitation
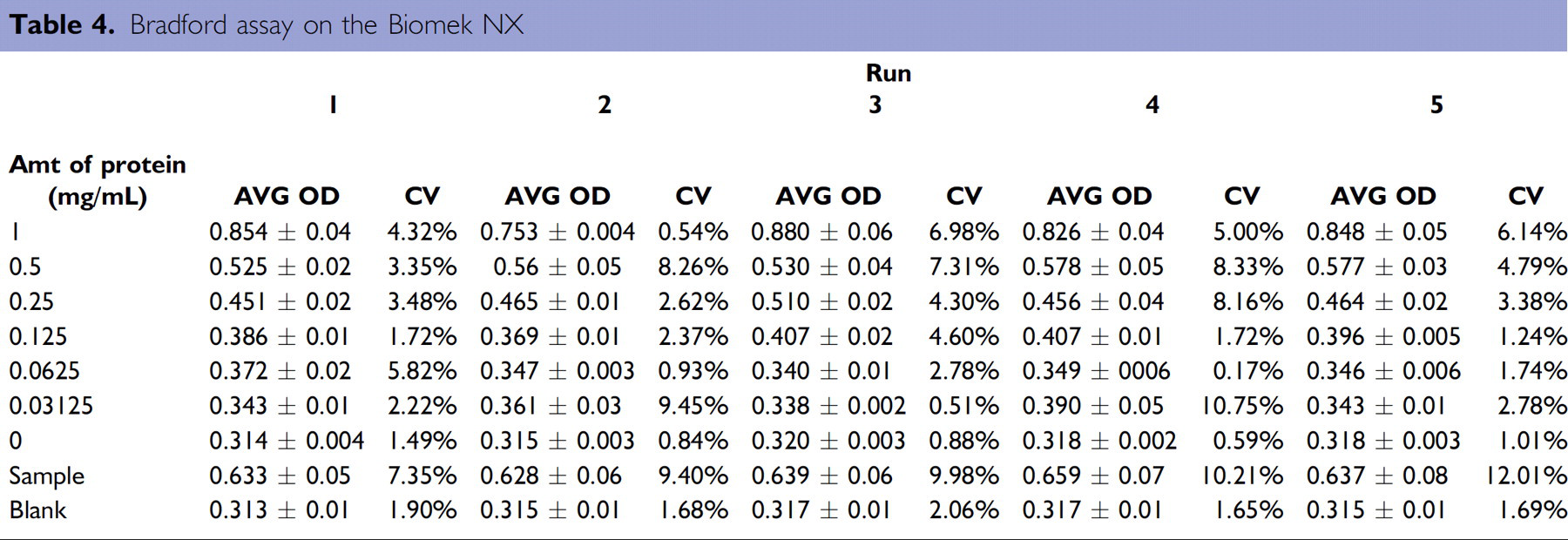
Prior to purifying proteins on the Biomek NX, the quantitation method was tested to ascertain its repeatability and robustness. To do so, a stock solution of bovine serum albumin (BSA) was seeded into even numbered columns of a 96-well plate and used as a source plate in the quantitation method. A standard curve was generated in conjunction with each experimental plate. Table 4 includes the average O.D.s and the C.V.s obtained for the standard curves for 5 separate runs of the method. The sample concentrations were calculated from the standard curves generated for each execution of the method; a representative standard curve is shown in Figure 13A. The calculated concentration of protein ranged from 0.599 to 0.692 mg/mL with the average concentration being 0.635 mg/mL (Figure 13B). Now verified, the quantitation portion of the process was included in the final protein purification and quantitation method on the Biomek thereby allowing for complete processing from bacterial samples to quantitation information.
Bradford assay on the Biomek NX
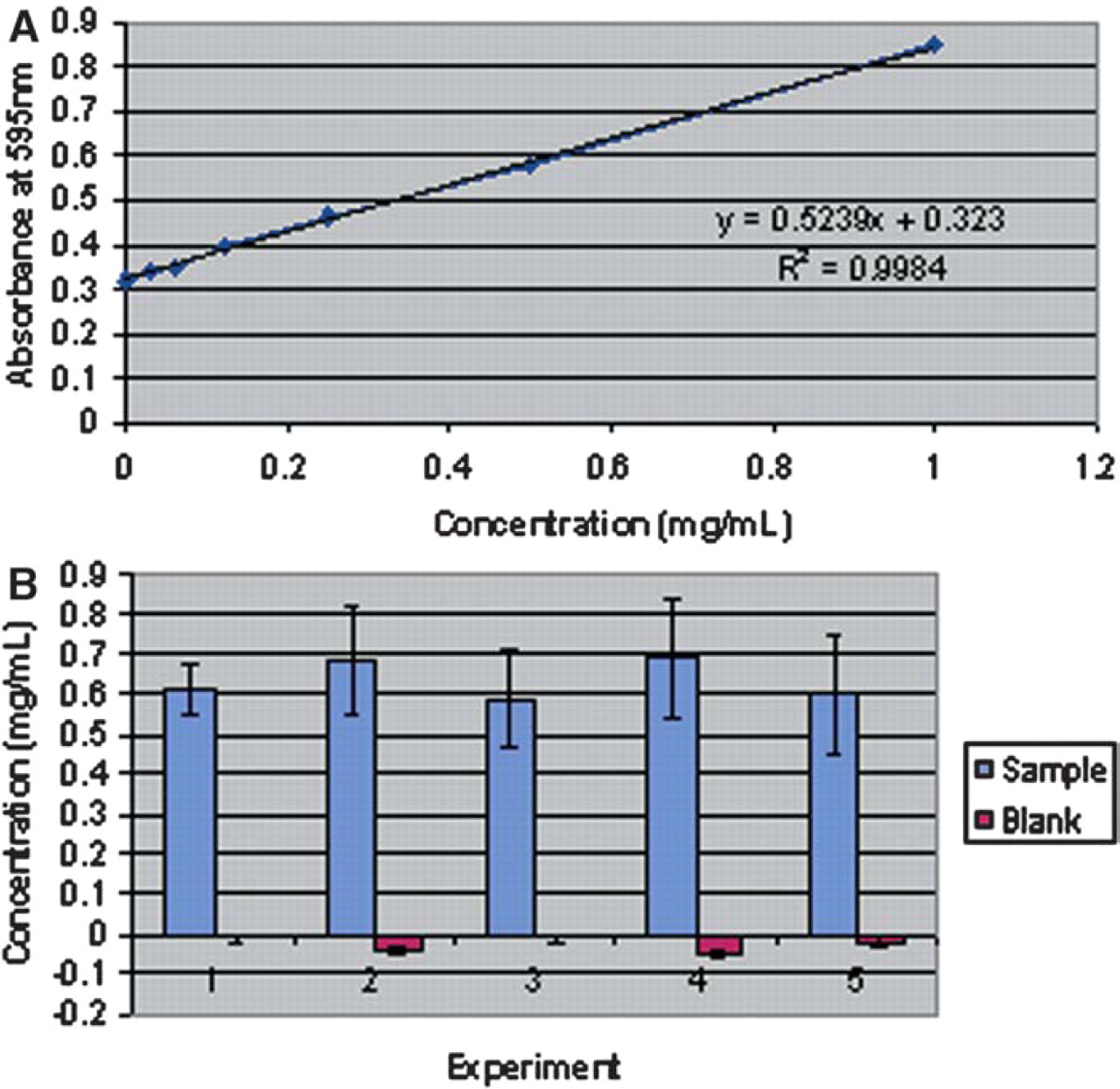
(A) A representative standard curve. (B) The calculated concentration of sample wells containing BSA and blank wells containing PBS.
Data from Automated Purification and Quantitation of His-Tagged Proteins
A deep-well plate with bacteria containing the His-tagged lacZ and IL-8 plasmid constructs was processed on the Biomek NX. The samples were lysed for two hours and eluted in 75 μL elution buffer. A 5 μL aliquot was used for quantitation and that data shown in Table 5.
Concentration of proteins using a two-hour lysis incubation
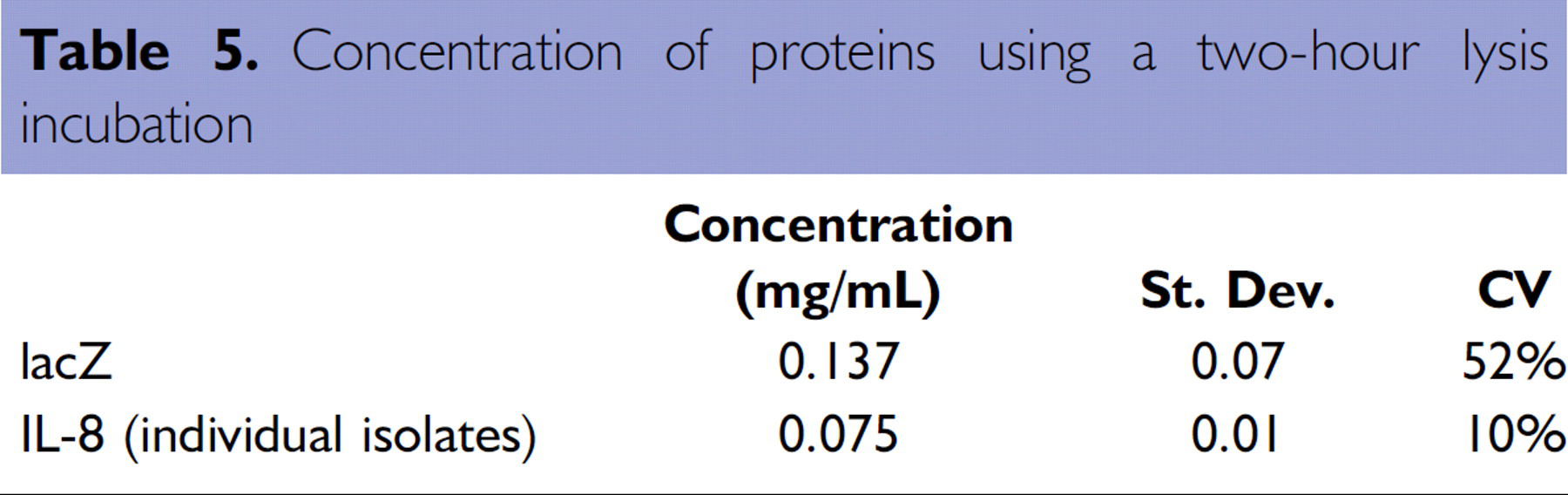
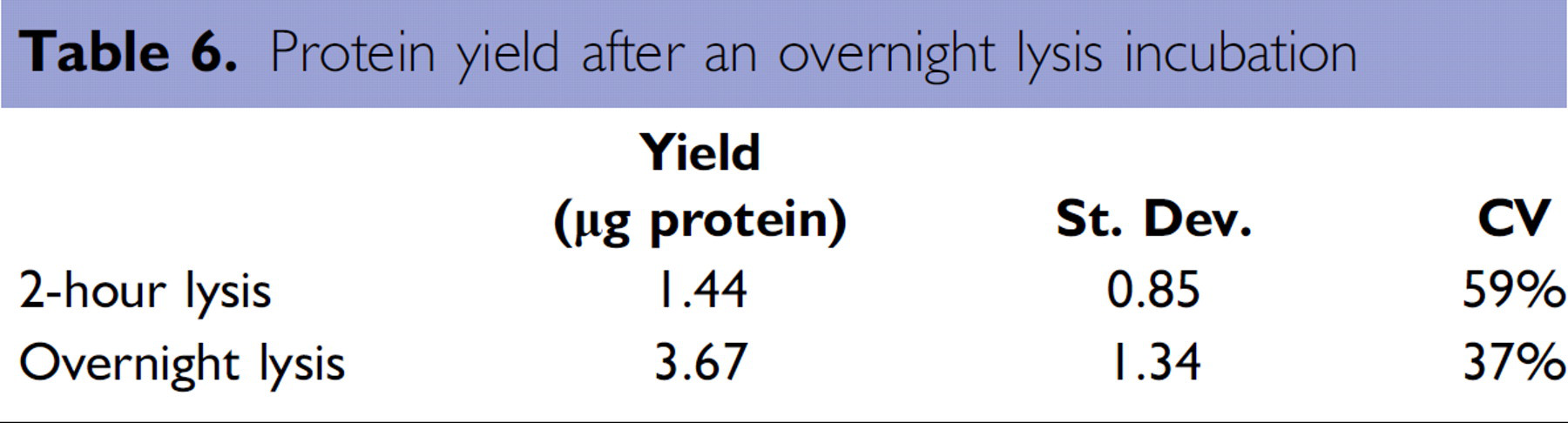
A larger amount of His-tagged lacZ was recovered as compared to the recombinant IL-8 protein. This was an expected result as the growth of the IL-8-producing bacteria cultures was slow compared to the lacZ-producing bacteria cultures. To maximize the lysis of the bacterial cells, cultures were dispensed into a HIS-Select iLAP HC Nickel-Coated plate and incubated overnight at 4°C. As a consequence, the extended overnight lysis incubation enabled a greater than two-fold increase in recovery as compared to a 2-hour lysis incubation (Table 6).
Protein yield after an overnight lysis incubation
SDS-PAGE Analysis to Demonstrate Enrichment of IL-8 Protein after Purification
Purified IL-8 proteins were analyzed by SDS-PAGE to assess the quality of the purified protein. As shown in Figure 14, purified His-tagged IL-8 eluates were enriched for the IL-8 protein as compared to bacterial lysates.
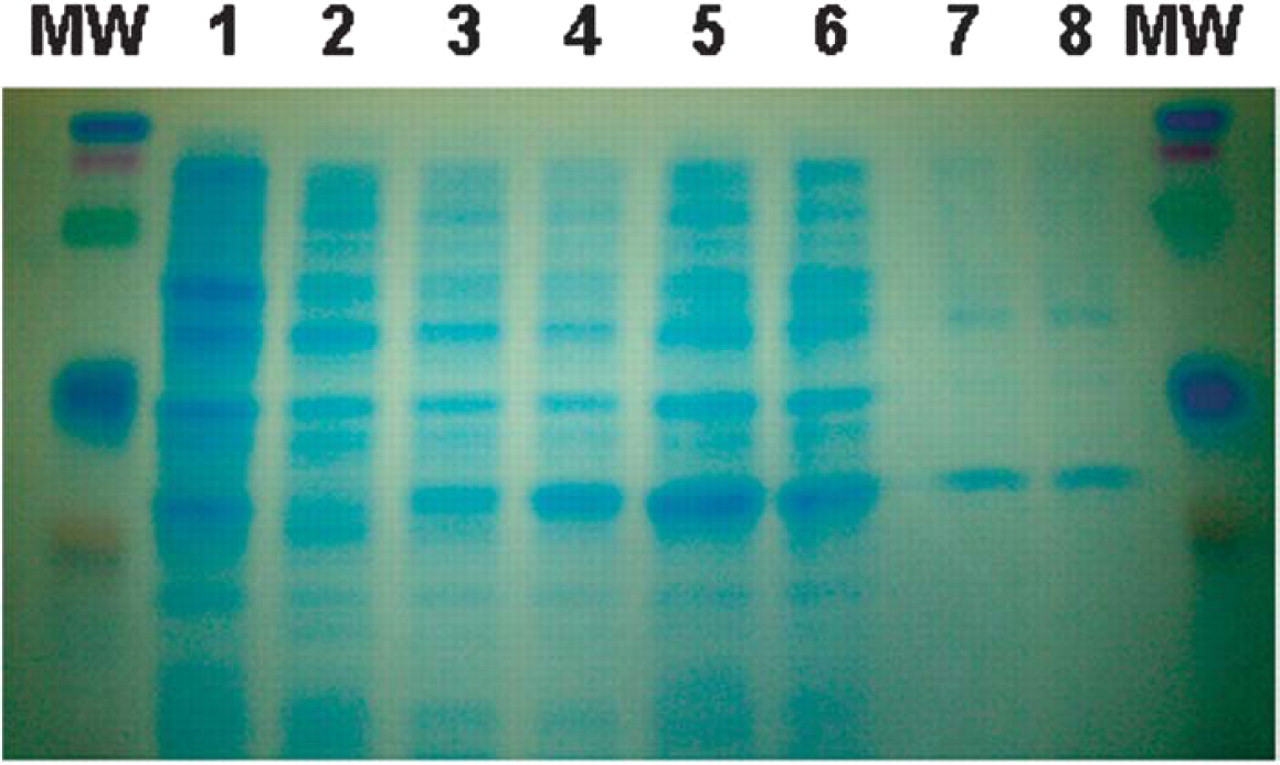
SDS-PAGE Analysis of purified IL-8 proteins. Arrow indicates induced IL-8 protein. Lane Order: 1 =host bacterial cells and Lanes 2–6 = IL-8 producing bacteria harvested after induction for 0, 1, 2, 3 hours, and overnight, respectively; Lanes 7–8 = purified IL-8; MW=molecular weight standards.
Utilization of Purified IL-8 Proteins in Downstream ELISA Assay
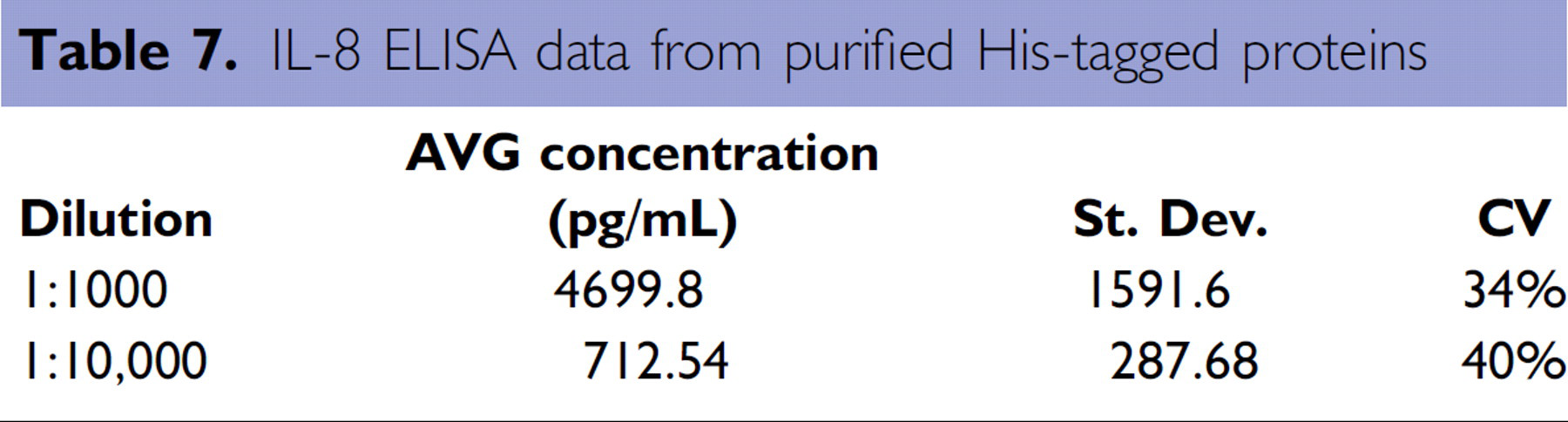
Purified IL-8 protein was used in an automated IL-8 ELISA assay to demonstrate the utilization of purified proteins in a downstream assay. To this end, the purified IL-8 samples were run in a dilution series on the Biomek 3000 with Beckman Coulter's Immunotech™ IL-8 ELISA kit (Catalog No. IM2238, Beckman Coulter, Inc., Fullerton, CA). The results from this assay are shown in Table 6 and illustrate that the proteins purified on the Biomek NX may be used in downstream assays. Additionally, the variability in the ELISA assay is reflective of the variability observed in purification of bacterial cultures (compare Tables 6 and 7).
IL-8 ELISA data from purified His-tagged proteins

CONCLUSION
The data presented here demonstrate that the Biomek NX can be utilized in genomic and proteomics processes such as the purification of plasmid DNA and proteins. The plasmid purification method detailed here generated high quality plasmid DNAs from standard growth conditions employed when generating templates for sequencing reactions. Furthermore, the same method produced plasmid DNA from colonies isolated in a bacterial two-hybrid screening experiment that was directly sequenced after purification. The methodology on the Biomek NX only required slight modifications to accommodate the low copy number plasmid and the endonuclease positive strain of bacteria. Additionally, the Biomek NX can be utilized for the purification and quantitation of proteins. This was achieved using a single, continuous method. An integrated reader, the AD 340 ALP, allowed for complete processing of samples to data without user intervention. The isolated His-tagged proteins were used in a functional assay, the IL-8 ELISA, thereby demonstrating the utility of proteins isolated on this platform for such downstream assays.
Acknowledgments
The authors would like to thank David Daniels, Ph.D., and the members of the Applications Group of Beckman Coulter, Inc. for their review and comments. The work on the bacterial two-hybrid screen was done in collaboration with Loren J. Field, Ph.D., Professor of Medicine, Indiana University School of Medicine, Indianapolis, IN. We would also like to thank Clay Watkins for his assistance with instrumentation.
