Abstract
Background
Our previous study identified rs9387478 as a new susceptibility locus associated with lung cancer in never-smoking women in Asia; however, the clinical and prognostic significance of this finding is not known.
Methods
We analyzed the relationship between the rs9387478 single nucleotide polymorphism and i) clinical parameters and ii) overall survival time in 505 female nonsmoking lung cancer patients, using the chi-square test and Kaplan-Meier analysis with the log-rank test, respectively. We further established the epidermal growth factor receptor (EGFR) mutation status and assessed its association with rs9387478 genotypes as well as the efficacy of EGFR tyrosine kinase inhibitors.
Results
The frequency of the AA genotype was significantly higher in the EGFR-mutation-negative group than in the EGFR-mutation-positive group (32% vs. 16%, χ 2 = 13.025, p = 0.011). Patients with the CC genotype had a better overall survival time than patients with the AA/AC genotype (median survival time: 54.2 vs. 32.9 months, χ 2 = 4.593, p = 0.032). The distribution of rs9387478 genotypes differed according to the clinical disease stage.
Conclusions
This study indicates that the rs9387478 genotype was associated with overall survival in nonsmoking female patients with lung cancer, although it was not significant after adjusting for multiple testing. The identification of the location of the rs9387478 single nucleotide polymorphism in the genomic interval containing the DCBLD1 and ROS1 genes, together with the finding that the rs9387478 polymorphism correlates with EGFR mutation status, may have important implications for therapeutic approaches targeting EGFR or ROS1 in patients with lung cancer.
Introduction
Lung cancer is the most common cancer among men, comprising 17% of new cancer cases and 23% of cancer deaths. Furthermore, lung cancer now accounts for 11% of cancer deaths among women in developing countries (1). Although most lung cancers are the result of smoking, approximately 25% of lung cancer cases worldwide are not attributable to tobacco use (2). Our previous research suggests that patients with different smoking statuses have different variations in driver genes (3). Understanding the clinical and prognostic significance of this genetic variation will be helpful for the prevention and early detection of lung cancer and for the selection of appropriate treatment strategies for cancer patients. To improve the clinical prognosis for patients with lung cancer, several targeted therapies are being developed, particularly in relation to cancers with specific molecular features. To date, researchers have focused on understanding the molecular abnormalities of lung cancer and their repercussions for the efficacy of molecularly targeted treatments (4-5-6).
We previously identified rs9387478 as a new susceptibility locus associated with lung cancer in never-smoking women in Asia (7). The rs9387478 single nucleotide polymorphism (SNP) at 6q22.2 is located in the genomic interval containing 2 candidate genes: discoidin, CUB and LCCL domain containing 1 (DCBLD1) and ROS proto-oncogene receptor tyrosine kinase (ROS1). There have been very few functional studies regarding DCBLD1. The related gene discoidin, CUB and LCCL domain containing 2 (DCBLD2; also known as CLCP1) may have an important role in cancer metastasis. The CLCP1 gene was found to encode a protein of 775 amino acids in length, with structural similarities to neuropilins, which are cell surface receptors for VEGF165 and semaphorins. Increased expression of CLCP1 may occur in lung cancer during tumor progression, possibly correlating with the acquisition of metastatic capabilities (8). DCBLD2 is frequently downregulated and may play an important role in gastric carcinogenesis (9).
The orphan receptor ROS1 is a human proto-oncogene, mutations of which are found in an increasing number of cancers (10). ROS1 encodes a tyrosine kinase that is abnormally expressed and translocated in both brain and lung cancers (11). It has recently been shown to undergo genetic rearrangements in a variety of human cancers including glioblastoma, non-small cell lung cancer (NSCLC), cholangiocarcinoma, ovarian cancer, gastric adenocarcinoma, colorectal cancer, inflammatory myofibroblastic tumor, angiosarcoma, and epithelioid hemangioendothelioma (12). The genetic rearrangement of ROS1 was identified as a distinct molecular signature for human NSCLC (13). ROS1 gene rearrangements are reported in 1%-2% of lung adenocarcinomas and correlate with the response to the multi-targeted tyrosine kinase inhibitor crizotinib, an anaplastic lymphoma receptor tyrosine kinase (ALK)/met proto-oncogene hepatocyte growth factor receptor (MET)/ROS1 inhibitor. Administration of crizotinib led to significant tumor shrinkage in ROS1-rearranged NSCLC (14, 15). CD74-ROS1 fusion transcripts were detected in 1 of 114 (0.9%) NSCLCs (16). ROS1 is involved in the carcinogenesis of a subset of NSCLCs and represents an independent oncogenic driver and viable therapeutic target (17).
Considering the important roles of the genes related to rs9387478 in cancer development and target therapy, and considering the implications of rs9387478 for lung cancer susceptibility in female nonsmokers, we hypothesized that the genotypes of rs9387478 may correlate with clinical parameters and may have prognostic or predictive significance. We therefore analyzed the relationships between rs9387478 polymorphism genotypes and clinical parameters, EGFR mutation status, progression-free survival (PFS) in response to tyrosine kinase inhibitors (TKIs), and overall survival (OS) in female nonsmoking lung cancer patients. These data may contribute towards the development of individualized therapies for patients with lung cancer.
Materials and methods
Patient Selection and Specimens
The study population was derived from the Genome-Wide Association Study (GWAS) of the Female Lung Cancer Consortium in Asia (FLCCA), which includes studies drawn from mainland China, South Korea, Japan, Singapore, Taiwan and Hong Kong (7). There are currently 648 women enrolled in the GWAS study at Guangdong General Hospital (Fig. 1). The study was approved by the ethics committee of Guangdong General Hospital (No. 201053). All patients provided written informed consent.
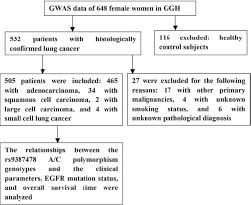

Study design and key procedures. GGH = Guangdong General Hospital; GWAS = Genome-Wide Association Study.
The eligibility criteria were a histological diagnosis of primary lung cancer; availability of demographic data including age, gender, smoking status, and histology; availability of survival data; and provision of informed consent. Patients with other primary malignancies and unclear pathological diagnoses were excluded. Clinical data were collected from the case histories of the patients in the hospital. Nonsmokers were defined as patients who had smoked fewer than 100 cigarettes in their lifetime. The patients were followed through telephone calls or re-examination of their records by the hospital follow-up group. OS was defined as the time from the first day of diagnosis to the date of death or the date when patients were last known to be alive. PFS was defined as the time from the first day of EGFR-TKI treatment to the date of progression.
DNA Extraction and Genotyping
Genomic DNA was obtained from whole-blood and normal lung tissue samples. The normal lung tissues of the patients were obtained during surgical resection and represented lung tissue located more than 5 cm from the tumor tissue. Tissue biopsies were snap-frozen in liquid nitrogen and stored at −70°C until analysis. DNA was extracted from normal tissue using the Aqua-SPIN Tissue/Cell gDNA Isolation Mini Kit (Biowatson). DNA was extracted from whole blood using a Universal Genomic DNA Extraction Kit (TaKaRa) according to the manufacturer's instructions. The integrity and quantity of DNA was assessed by gel electrophoresis and Thermo NanoDrop 1000 (Thermo Scientific) analysis. Samples were genotyped at Gene-Square Biotech in Beijing via contract, using the Illumina 660W SNP microarray. The genotype results were analyzed by the NCI team and sent to the Guangdong Lung Cancer Institute. The results were confirmed by polymerase chain reaction (PCR)-based sequencing in our research (Fig. 2). The sequences of the primers used in PCR to study the genotypes of rs9387478 were as follows: forward primer 5’-TGTGAACATACAGAAGAACAAGGGA-3’ and reverse primer 5’-AACAGGGAACTGCCAAAGACAAAAT-3’. The PCR cycling parameters were 1 cycle of 94°C for 7 minutes; 30 cycles of 94°C for 30 seconds, 58°C for 30 seconds, and 72°C for 1 minute; followed by 1 cycle of 72°C for 7 minutes. All sequencing reactions were done in both forward and reverse directions with 2 primers for PCR.
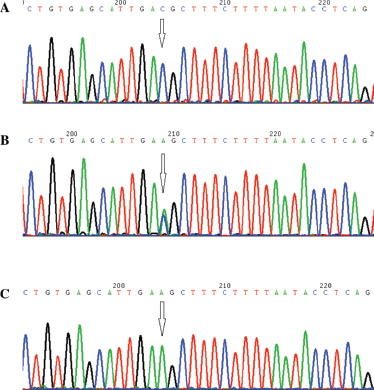
Representative images of PCR genotype analyzing of rs9387478.(
Detection of the EGFR Mutation Status
Mutation analysis of the EGFR-tyrosine kinase domain was performed by PCR-based sequencing (Fig. 3), as described previously (18). Exons 18-21 were amplified using 4 pairs of primers. Both forward and reverse sequencing reactions were performed using specific primers.
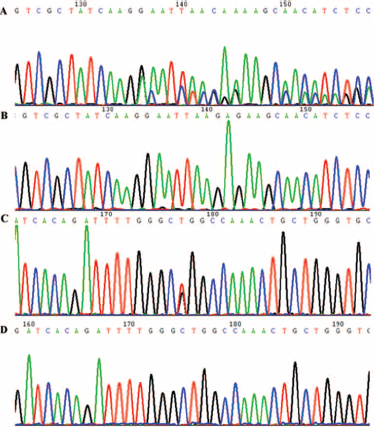
Representative images of PCR genotype analyzing of EGFR.
Statistical Analysis
Differences in genotype frequencies between groups were evaluated using the chi-squared test and Fisher's exact test, where appropriate. Survival analysis was performed using Kaplan-Meier analysis with a log-rank test. Multivariate analyses were conducted using the Cox proportional hazards model (Forward: Wald; p = 0.05, entry; p = 0.10, Removal). A 2-tailed p value of <0.05 was considered statistically significant.
Results
Patient Characteristics
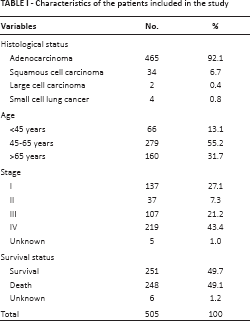
A total of 505 eligible patients were enrolled in the study. Tissue samples from these patients were obtained from 2003 to 2010 and stored in the hospital tumor bank. The patient population featured histologically confirmed lung cancer cases of all stages (Tab. I). Of these patients, 465 had adenocarcinomas, 34 had squamous cell carcinomas, 2 had large cell carcinomas, and 4 had small cell lung cancer; 137 were in stage I, 37 were in stage II, 107 were in stage III, 219 were in stage IV, and 5 were of unknown stage. The mean age of the patients was 58.3 years, ranging from 26 to 85 years. On the last date of follow-up (September 2013), there were 251 survivors, 248 patients had died of lung cancer, and the outcomes of 6 patients were unknown.
Characteristics of the patients included in the study
Frequency of rs9387478 A/C Polymorphism Genotypes and Relationship to Clinical Parameters
The frequencies of the AA, AC, and CC genotypes of the rs9387478 A/C polymorphism among the studied female nonsmoking lung cancer patients were 21.0% (106/505), 51.1% (258/505), and 27.9% (141/505), respectively. According to the NCBI SNP data bank http://www.ncbi.nlm.nih.gov/projects/SNP/snp_ref.cgi?rs=9387478, the frequencies of the AA genotype of the rs9387478 polymorphism in European, Asian, Sub-Saharan African, and African-American populations were 16.7%-26.5%, 19.3%-57.3%, 61.2%-90.3%, and 73.9%, respectively.
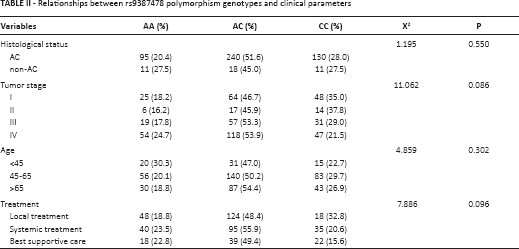
We analyzed the correlations between the rs9387478 A/C polymorphism genotypes and the patients’ clinical parameters including histological status, clinical stage, and age. The distribution of the genotypes was not related to age or tumor histology (χ2 = 4.859 or 1.195, p = 0.302 or 0.550, respectively; Tab. II). The correlation between genotype and clinical stage was marginally significant (χ2 = 11.062, p = 0.086; Tab. II). Therefore, we combined the groups with AA and AC genotypes (AA/AC) and compared this new group with CC genotype patients. The AA/AC genotype had a higher prevalence in patients with advanced tumors than in those with early tumors (χ2 = 9.836, p = 0.020). We also analyzed the rs9387478 polymorphism genotypes in relation to different treatment strategies and found no statistically significant differences between different treatments (p = 0.096).
Relationships between rs9387478 polymorphism genotypes and clinical parameters
Distribution of rs9387478 Genotypes in Patients with different EGFR Mutation Statuses
Mutations in exons 18-21 of EGFR were evaluated in the 288 patients for whom tumor tissues were available. We classified patients into 2 subgroups for further study: an EGFR mutation-positive and an EGFR mutation-negative group. The EGFR mutation rate was 56.9% (164/288) in the female nonsmoking patients with lung cancer. The proportions of AA, AC, and CC rs9387478 polymorphism genotypes were 32% (40/124), 44% (54/124), and 24% (30/124), respectively, in the EGFR mutation-negative group, and 16% (27/164), 56% (91/164), and 28% (46/164) in the EGFR mutation-positive group, respectively. The frequency of the AA genotype was significantly higher in the EGFR mutation-negative group than in the EGFR mutation-positive group (32% vs. 16%, χ2 = 13.025, p = 0.011; Fig. 4).
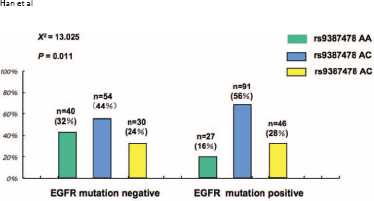
Correlation of rs9387478 polymorphism genotypes with EGFR mutation status.
Survival Analysis of Patients with different rs9387478 Polymorphism Genotypes
To assess whether the different genotypes of the rs9387478 polymorphism correlated with survival time, we classified patients into 3 genotype groups (AA, AC, and CC). There was a marginal difference in OS time between the 3 groups. The median survival times (MSTs) of patients with the AA, AC, and CC genotypes were 31.7 months, 33.6 months, and 54.2 months, respectively (χ2 = 4.653, p = 0.098, Kaplan-Meier analysis, log-rank test; Fig. 5A). When the CC genotype group was compared with the combined AA/AC genotype group, the differences in OS were statistically significant; patients with the CC genotype had a better OS time than those with the AA/AC genotype (MST: 54.2 vs. 32.9 months, χ2 = 4.593, p = 0.032; Fig. 5B).
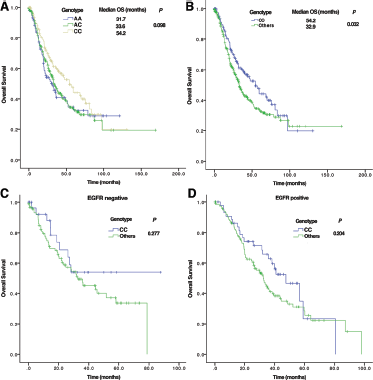
The relationship between rs9387478 polymorphism genotypes and overall survival.
We also performed subgroup analyses based on EGFR mutation status. In the EGFR mutation-negative group, no differences were found between patients with the CC and AA/AC genotypes (MST: 32.9 vs. 36.0 months, χ2 = 1.183, p = 0.277; Fig. 5C). Although it appears that patients with the CC genotype in the EGFR mutation-positive group had a longer survival time than patients with the AA/AC genotype, Kaplan-Meier analysis with the log-rank test showed no significant difference in OS between the groups (MST: 47.6 vs. 32.5 months, χ2 = 1.611, p = 0.204; Fig. 5D).
The relationship between EGFR mutation status and the OS time was also analyzed. The results showed no difference in survival time between the EGFR mutation-positive and negative groups (χ2 = 0.957, p = 0.328).
Multivariate analysis using Cox regression was used to assess the significance for survival according to EGFR mutation status, stage, histology, age, and rs9387478 polymorphism genotype, and it indicated that disease stage was the only independent prognostic factor (χ2 = 196.274, p<0.0005, Forward: Wald; p = 0.05, entry; p = 0.10, Removal).
Relationship between rs9387478 Polymorphism and PFS in Response to EGFR-TKI
The above results show that the rs9387478 polymorphism correlated with the EGFR mutation status. TKIs targeting EGFR, including gefitinib and erlotinib, have become the standard first-line therapy for patients with advanced NSCLC and activating EGFR mutations (6, 19). We analyzed the effect of the rs9387478 polymorphism on PFS in response to TKI treatment.
There were 88 patients in our study treated with TKIs for whom PFS data were available. The results indicated that the median PFS in patients with the AA, AC, and CC genotypes were 9.0, 7.0, and 6.0 months, respectively. The differences between the groups did not reach statistical significance (χ2 = 1.261, p = 0.532, Kaplan-Meier analysis, log-rank test).
Discussion
The identification of genetic biomarkers and patterns of genetic risk may enable the earlier detection and treatment of lung cancer (20). The GWAS is a promising initiative for identifying common genetic variants of diseases on the basis of millions of SNPs (21). Most of the SNPs found via GWAS are related to lung cancer susceptibility (22-23-24-25), but only a small proportion have been confirmed to correlate with clinical parameters and validated as prognostic or predictive biomarkers for lung cancer. For example, rs2282987 and rs2706748 interact with both smoking status and smoking history in contributing to lung cancer susceptibility in subjects aged 51-60 years (26). Similarly, rs4655567, rs8020368 and rs2018683 were associated with resistant relapse (27) and rs6034368T>C with survival in early-stage NSCLC in a Korean population (28).
The rs9387478 polymorphism represents a new susceptibility locus associated with lung cancer in never-smoking women in Asia (7), but the clinical and prognostic significance of this finding for lung cancer patients remains unclear. To the best of our knowledge, this is the first study to analyze the clinical and prognostic significance of this SNP. We found that patients with the CC genotype have improved OS compared to those with the AA/AC genotype. We also demonstrated that this SNP correlates with the patient's EGFR mutation status. These data suggest that rs9387478 may be a functional polymorphism in lung cancer patients.
The region where rs9387478 locates was defined by the Encyclopedia of DNA Elements (ENCODE) as containing both chromatin state segmentation and enhancer- and promoter-associated histone marks (7). Previous research has shown that SNPs in CYP3A5*3 and CYP3A5*6 can cause alternative splicing and protein truncation can result in the absence of CYP3A5 from tissues of some people (29). We hypothesize that the rs9387478 polymorphism resides within a gene response element and that the different genotypes may affect the affinity of DNA-binding proteins for the gene promoter, resulting in different transcriptional activities. This hypothesis requires further investigation.
The rs9387478 SNP is located 38,861 base pairs upstream of the proto-oncogene tyrosine-protein kinase ROS precursor. Our study suggests that patients with the CC genotype of the rs9387478 polymorphism have a better OS than patients with the AA/AC genotype, and this suggests a potential relationship between ROS and lung cancer prognosis for female nonsmoking patients. ROS1 fusions occurred in about 2.0% of Chinese patients with NSCLC, and it is likely to be associated with favorable prognostic factors for invasive ductal carcinoma of the breast (30). ROS1 fusion-negative patients may have a better survival time than ROS1 fusion-positive patients (31).
Crizotinib, an inhibitor of ALK, was also recently shown to have efficacy in the treatment of lung cancer with ROS1 translocations (32). Targeting ROS1 fusion proteins by means of crizotinib is showing promise as an effective therapy in NSCLC patients whose tumors are positive for these genetic abnormalities (33). ROS1 rearrangement is a drug target in East-Asian never-smokers with lung adenocarcinoma (33). How the rs9387478 SNP genotype relates to the outcome of the target therapy requires further analysis.
We also found that the rs9387478 polymorphism correlates with EGFR mutation status. EGFR mutation is the most reliable biomarker for EGFR-TKI treatment of patients with NSCLC. EGFR mutation detection is done on tumor specimens, but tumor specimens may not be available for all patients. SNPs can be detected in many other specimens, including peripheral blood. Although rs9387478 and EGFR are not connected at the DNA sequence level, it will be very useful if we can find the relationships between them. The SNP results can be surrogate biomarkers for EGFR mutation in some patients for whom tumor specimens are not available, so these patients will have one more chance to be treated by EGFR-TKIs. Patients with the AC/CC genotype are more likely to benefit from EGFR-TKIs. But the implication needs further investigation. The rs9387478 polymorphism is located 17,265 base pairs upstream of DCBLD1. Although there have been few functional studies on DCBLD1, the related gene DCBLD2 was identified as a novel target of gefitinib, which is a highly selective inhibitor of EGFR. These novel data will provide new insights into the complexities of EGF signaling and may have implications for target-directed cancer therapeutics (34).
Although the rs9387478 polymorphism was found to correlate with the OS of patients in univariate analysis, Cox regression indicated that clinical stage is the only independent factor that can predict patient survival. Disease stage is a well-known independent prognostic factor, and survival rates and treatment choices depend on it (35). The distribution of rs9387478 genotypes differs according to the clinical disease stage. The potential mechanisms underlying the prognostic value of rs9387478 is that patients with AA/AC genotype are more likely to develop distant metastases. Therefore, the rs9387478 polymorphism is likely a very weak prognostic factor. In addition, the different treatment strategies, especially the treatment target, may affect the survival of the patients.
We further analyzed the relationship between the rs9387478 polymorphism and the outcomes of TKI treatment in the patients. The correlation found between the rs9387478 polymorphism and the EGFR mutation status implies that the rs9387478 polymorphism may be a new biomarker for prediction of EGFR-TKI outcomes, which may aid in the selection of treatment strategies, including TKIs. However, no significant difference was found in PFS in response to TKIs among the different genotypes. A significant result may have been obscured by the small sample size or the heterogeneity of the tumors studied. Further investigation is required.
Our study is a pilot analysis of the rs9387478 polymorphism as a prognostic or predictive biomarker for female nonsmoking patients from 1 research center. The significance for all patients, including males, should be studied in the future. The molecular mechanisms underlying our findings also merit further study.
Conclusions
This study indicates that the rs9387478 polymorphism is associated with OS in nonsmoker female patients with lung cancer, although it was not significant after adjusting for multiple testing. Whether therapeutic strategies should be initiated earlier in patients with the AA/AC genotype to improve their survival time needs further investigation. The relationship between the rs9387478 polymorphism and EGFR mutation status suggests that this polymorphism is a potential surrogate biomarker for EGFR mutation detection, with the advantage of less invasive sample collection. These findings provide insight into the practical considerations for successful application of tumor genotyping in clinical decision-making.
Footnotes
Abbreviations
Financial support: This work was supported by the National Natural Science Foundation of China (Nos. 81101549, 30772531 and 81272618) and Nonprofit Research and Special Projects of National Health and Family Planning Committee of China (No. 201402031).
Conflict of interest: The authors declare they have no competing interests.
