Abstract
Background
MicroRNAs (miRNAs) have been emerging as valuable prognostic biomarkers of breast cancer. We therefore summarized recent research into miRNAs involved in human breast cancer and, further, completed a meta-analysis to predict the role of specific miRNAs in the survival of breast cancer patients.
Methods
Studies were identified by searching PubMed, Embase and Web of Science. Descriptive characteristics for studies were described, and an additional meta-analysis for specific miRNAs was performed. Pooled hazard ratios (HRs) and their corresponding 95% confidence intervals (CIs) were calculated.
Results
A total of 41 articles including 27 types of miRNAs were found regarding prognostic biomarkers for breast cancer survival, of which, micRNA-21 (miR-21) was the most-studied specific miRNA that appeared repeatedly among the selected classifiers. For the studies evaluating miR-21's association with clinical outcomes, the median HR in the studies was 2.32 (interquartile range [IQR] = 1.04-3.40), and the pooled HR suggested that high expression of miR-21 has a negative impact on overall survival (OS; HR = 1.46, 95% CI, 1.25-1.70; p<0.05) and disease/recurrence-free survival in breast cancer (HR = 1.49, 95% CI, 1.17-1.90; p<0.01). We also found that higher expression levels of miR-210 significantly predicted poorer outcome, with median HR in the reported studies of 4.07 (IQR = 1.54-4.43) and a pooled HR of 2.94 (95% CI, 2.08-4.17; p<0.05).
Conclusions
These results indicate that miRNAs show promising associations with prognosis in breast cancer. Moreover, specific miRNAs such as miR-21 and miR-210 can predict poor survival rates in breast cancer patients.
Introduction
Breast cancer is a significant health problem and ranks as the second most common cause of cancer death in female patients (1). There has been an increasing incidence of breast cancer in developing countries, resulting from urbanization and adoption of Western lifestyles, and reduced life expectancy (2). The heterogeneity of breast cancer poses serious clinical challenges. Thus, different approaches to stratifying patients with respect to their prognosis have been developed. In clinical practice, the management of breast cancer is based on established criteria such as tumor size, the presence of distant metastasis, histologic type and grade, the expression of hormone (i.e., estrogen and progesterone) receptors (ER and PR) and the human epidermal growth factor 2 (ERBB2/HER2) receptor (3). Ongoing advances in breast cancer management have significantly changed outcomes for patients (4). To further that advance, detectable and stable biomarkers are required for clinical practice.
Recently, the field of transcriptome has been further expanded through analyses of noncoding RNAs (ncRNAs). MicroRNAs (miRNAs) are small, ncRNAs of 19-24 nucleotides in length that were first discovered in Caenorhabditis elegans by Victor Ambros et al in 1993 (5). Since then, our knowledge of miRNA biology has continued to grow. More and more cancer-relevant miRNAs have been well characterized (6), and they have been critically implicated in carcinogenesis as either oncogenes or tumor suppressors (7). Due to their unique expression profile in human cancers, the fact that they are more stable than mRNAs and their levels and presence are easily assessed, miRNAs hold great promise as potential biomarkers for cancer prognosis.
Several studies have reported promising results for miRNA expression classifiers for prognosis of breast cancer (8, 9). The data from studies providing independent assessments of miRNAs in breast cancer were systematically reviewed in our study. Furthermore, we performed a meta-analysis of studies regarding specific miRNAs as important prognostic factors for breast cancer.
Materials and Methods
Search Studies and data Extract
We followed the guidelines of the critical checklist of the Dutch Cochrane Centre as proposed by the Meta-analysis of Observational Studies in Epidemiology (MOOSE) proposal (10) in our study. A comprehensive literature search of PubMed was done for original articles analyzing the prognostic value of miRNAs in breast cancer. Studies were selected using the following keywords variously combined: breast cancer, microRNA, miRNA, prognosis and prognostic. Original and review articles published until 20 December 2014 were sought, and no lower date limit was used. Additional records were found through Embase and Web of Science. Furthermore, the bibliographies reported in the selected studies were reviewed to deepen this search. Authors were contacted whenever useful data were not reported or to exclude overlapping data that may have been published elsewhere.
All studies published as a full paper in English, dealing with breast cancer only (any stage or histology) and analyzing prognosis in patients stratified by dichotomous miRNA expression levels in tissues or blood were included. Review articles, letters, conference abstracts or laboratory articles were excluded. Subtypes of of miRNAs, for example, miR-34a, miR-34b and miR-34c were defined as 1 type (miR-34). The extracted data elements included the following: (i) publication details including the first author's name and publication year; (ii) characteristics of the studied population including country, source, stage and histological type; (iii) miRNA assessments and cutoff values; (iv) hazard ratios (HRs) of miRNAs for overall survival (OS) and disease-free survival (DFS) or recurrence-free survival (RFS), along with the 95% confidence intervals (95% CIs) and p values. If the HRs and their 95% CIs were not directly provided, the numbers of deaths and total samples in each study were extracted to calculate this. If only Kaplan-Meier curves were available, data were extracted from the survival plots, and the HR was calculated using methods established by Parmar et al, Williamson et al and Tierney (11). When the authors reported results obviously obtained from the same patient population in multiple publications, only the most recent studies were included. When the authors reported the validation set results obviously inconsistent with the discovery set, the data was abandoned. Two reviewers (Y. T. and X. Z.) independently evaluated the titles and abstracts of the identified articles in duplicate. A flow diagram of the study selection process is presented in Figure 1.
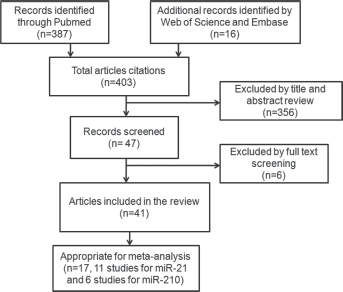
Flow diagram of the study selection process.
Statistical Analysis
Heterogeneity was assessed using the Q statistic (a p value <0.10 was considered heterogeneous). A random-effects model was used if prominent heterogeneity was observed (p<0.05 or I2>50%), otherwise a fixed-effects model was applied. Some studies did not list the HRs or 95% CI directly, instead giving Kaplan-Meier survival curves alone. The necessary statistics were performed using software designed by Tierney et al, Parmar et al, and Williamson et al (11-13). The effects of miR-21 and miR-210 expression on survival were estimated using forest plots. Pooled HR was calculated using a fixed-effects model or random-effects model as appropriate. A pooled HR >1 indicated poor prognosis for the groups with elevated miRNA expression and was considered statistically significant if the 95% CI did not overlap 1. Publication bias was evaluated using a funnel plot and Egger's test. A p value >0.05 was considered indicative of a lack of publication bias (14). All analyses were performed using STATA version 12.0 (Stata Corp., College Station, TX, USA). A p value <0.05 was considered to be statistically significant.
Results
Eligible Studies
A total of 403 studies of miRNAs and breast cancer prognosis were identified from a primary literature search in PubMed. Sixteen additional records were found in Embase and Web of Science. Of these, 356 studies were excluded based on manual screening of the title or abstract, and the remaining 47 were evaluated as full texts. Of the 47 candidate studies, 6 studies were not directly related to specific outcomes. Therefore, in the present study, we reviewed and listed a final 41 articles (9, 15-54) and identified unique miRNA classifiers associated with survival outcomes, as shown in Table I.
Main characteristics of the eligible studies
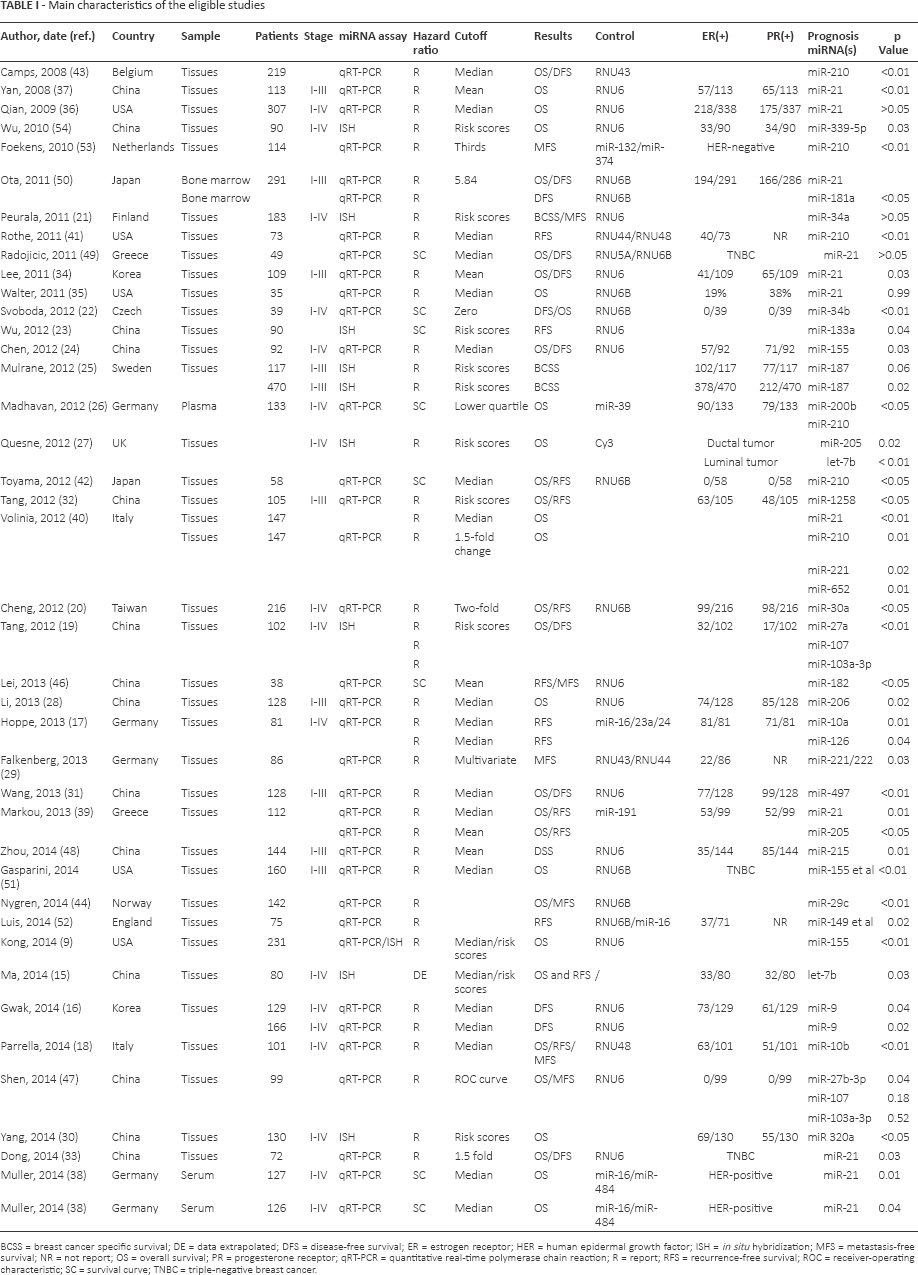
BCSS = breast cancer specific survival; DE = data extrapolated; DFS = disease-free survival; ER = estrogen receptor; HER = human epidermal growth factor; ISH = in situ hybridization; MFS = metastasis-free survival; NR = not report; OS = overall survival; PR = progesterone receptor; qRT-PCR = quantitative real-time polymerase chain reaction; R = report; RFS = recurrence-free survival; ROC = receiver-operating characteristic; SC = survival curve; TNBC = triple-negative breast cancer.
Characteristics of the Reviewed Studies
Among the 41 selected articles, there were 27 types of miRNAs within 55 study sets including a total of 5,507 patients, which were found to be prognostic biomarkers for breast cancer survival. The median study size was 113 patients (interquartile range [IQR] = 92-147 patients). Most studies (49/55) showed significant associations (p<0.05) between miRNA expression and survival of patients with breast cancer. Two studies used classifier information from more than 1 miRNA (51, 52), and another 39 studies used only a single microRNA for outcome analysis. Most studies analyzed breast cancer with frozen or formalin-fixed paraffin-embedded (FFPE) tumor tissues (49 studies), while 4 studies used blood samples and 2 studies used bone marrow. The quantitative real-time polymerase chain reaction (qRT-PCR) assay, which is widely used as the gold standard for validating microarray expression, was used in 45 studies, whereas in situ hybridization (ISH) assay was used in only 10 studies. Normalization was performed with endogenous controls in almost all studies. The most commonly used endogenous controls were U6 and U6B.
Most of the protective miRNAs (e.g., the let-7 family) were down-regulated (15), while the risky miRNAs (e.g., miR-21, miR-210 and miR-155) (24, 38, 39) were up-regulated in breast cancer and indicated poorer clinical outcomes in breast cancer patients. Moreover, up-regulated miRNAs were more common than down-regulated ones. We deduced that there may be a greater statistical ability to detect the up-regulated miRNAs as poorer prognostic biomarkers in breast cancer survival. All of the adjusted HRs (unless only an unadjusted value was available) revealed that the dysregulation of miRNAs in 6 studies was not statistically significantly associated with survival or recurrence (Tab. I). For all of the reported HRs, regardless of statistical significance in each study, the median HR was 1.31 (IQR = 0.51-2.80) and the overlapping of 1 indicated the variety role of miRNA classifiers in breast cancer. The ratio of combinations for high expression of miRNAs yielded a pooled HR of 1.07 (95% CI, 1.02-1.12; p<0.05; Fig. 2), with a prominent heterogeneity observed (p<0.01 or I2 = 90.1%). Because high expression levels of miRNAs were shown to be only a weak prognostic factor in breast cancer, subgroup analysis was completed to validate which type of miRNA underlying the studies had a better prognostic role than the others.
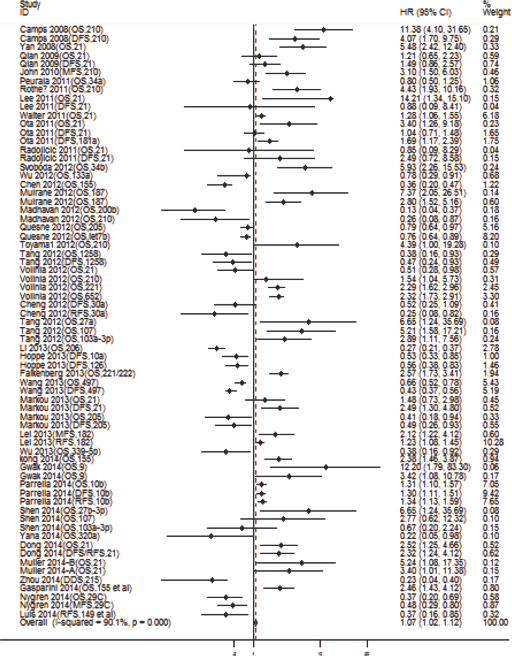
Forest plot for the association between high miRNA expression (specific miRNA indicated in parentheses next to author names and dates) and survival (overall survival [OS] and disease-free survival, recurrence-free survival or metastasis-free survival [DFS/RFS/MFS]) of patients with breast cancer. CI = confidence interval; HR = hazard ratio.
Meta-analysis Results of Specific miRNAs Associated with Prognosis
We extracted the data for the miRNA profiles from the 41 studies available and counted how many times each miRNA had been selected for survival analysis (Tab. II). A total of 27 prognostic miRNAs were identified in these classifiers, among which 6 miRNAs appeared more frequently, including miR-21 (11 times, 20.0%), miR-210 (6 times, 10.9%) and miR-10, miR-155, miR-27 and miR-30 (each 3 times, 5.5%). We performed a meta-analysis to identify whether miR-21 and miR-210 could be used as reliable prognostic biomarkers. Here we pooled the HR of miR-21 and miR-210 to look at the prognosis for breast cancer patients.
Statistical analysis of the defined miRNAs in this review, regarding how many times each miRNA was selected for survival analysis
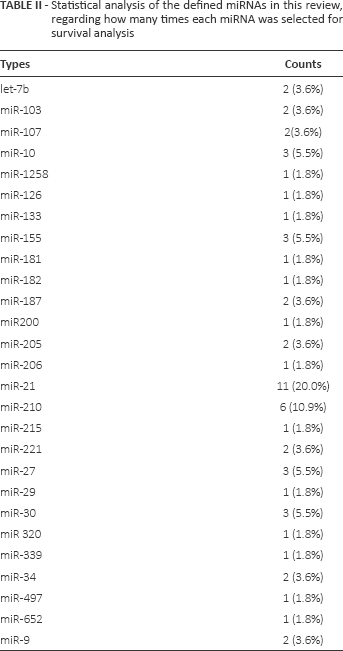
For miR-21, as shown in Table I, most studies analyzed breast cancer tissues samples (8 studies), and 3 studied serum sample. qRT-PCR was the only assay used in the 11 studies of miR-21. For each study, the p value varied from 0.99 to less than 0.01, and 3 studies had p values of more than 0.05, which were thus not statistically significant. We found that higher expression levels of miR-21 significantly predicted poorer outcomes, with the median HR in the reported studies of 2.32 (IQR = 1.04-3.40). There was heterogeneity among the studies for miR-21 evaluating OS (p<0.05, I2 = 78.9%) (Fig. 3A). Hence, a random-effects model was applied to calculate a pooled HR and its 95% CI. We found that higher expression levels of miR-21 significantly predicted poorer OS, with a pooled HR of 1.46 (95% CI, 1.25-1.70; p<0.05).
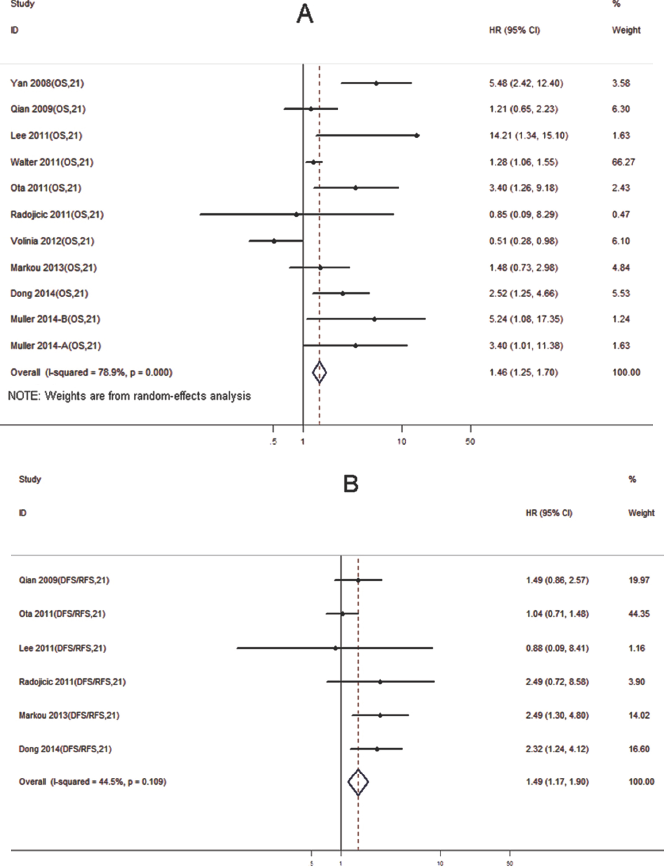
Forest plot for the association between high miR-21 expression and overall survival (OS)
To reduce the influence of heterogeneity, we reorganized the subgroup analysis according to ethnicity, internal controls, cutoff value, pathological types and material (Tab. III). In the ethnicity subgroup analysis, heterogeneity was considerably reduced between the Asian (p = 0.08, I2 = 55.7%) and Western groups (p = 0.17, I2 = 38.0%). Meanwhile, the pooled HR of elevated miR-21 of Asian cancer patients (HR = 4.08, 95% CI, 2.66-6.24; p<0.01) was higher than that of Western patients (HR = 1.33, 95% CI, 1.11-1.59; p<0.01). In the internal controls subgroup analysis, heterogeneity was not reduced between U6 and other controls (Tab. III). In the cutoff subgroup analysis, mean (HR = 7.38, 95% CI, 3.75-14.53; p<0.01) and fixed (HR = 2.76, 95% CI, 1.60-4.78; p<0.01) cutoff values were significantly associated with OS under a fixed-effects model because the heterogeneity in both cases was under 50%. OS was not significantly reduced in the median cutoff group. In the type subgroup analysis, overexpression of miR-21 was predictive of worse OS (HR = 4.23, 95% CI, 1.81-9.90; p<0.01) in Her2+ group patients by fixed-effects model with no heterogeneity among studies (p = 0.62, I2 = 0.0%). A similar result was found in the triple-negative group but not in the mixed type of breast cancer patients. In the material subgroup analysis, heterogeneity was eliminated in the blood/bone marrow group (p = 0.87, I2 = 0.0%) but not in the tumor tissue group (p<0.01, I2 = 82.1%).
Subgroup analyses of overall survival with elevated miR-21 expression
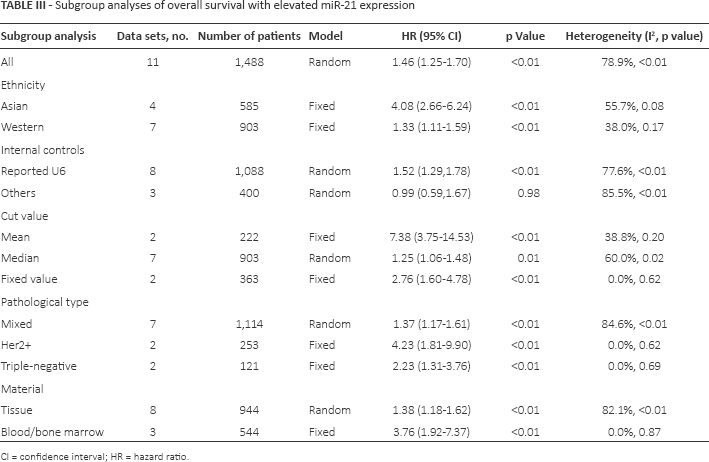
CI = confidence interval; HR = hazard ratio.
For the study evaluating miR-21 in DFS/RFS of breast cancer (Fig. 3B), a fixed-effects model was applied, because there was no heterogeneity among the cohorts (p = 0.11, I2 = 44.5%). The expression of miR-21 was also significantly correlated with DFS/RFS of breast cancer, with a combined HR of 1.49 (95% CI: 1.17-1.90, p<0.01). Finally, publication bias of the included studies for OS and DFS/RFS was evaluated by funnel plots and Egger's test, respectively. The p value of Egger's test for OS was 0.10 and DFS/RFS was 0.38, suggesting that no significant publication bias existed (Fig. 4A, B).
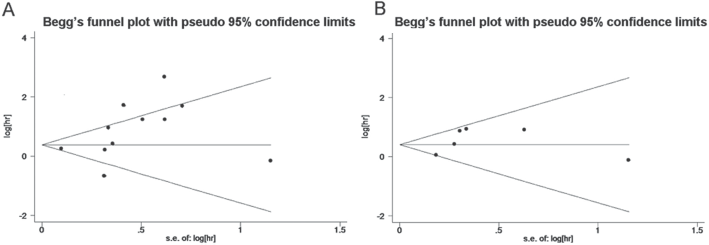
Funnel plot of high miR-21 expression on the overall survival (OS)
We also found that higher expression levels of miR-210 significantly predicted poorer outcomes, with the median HR in the studies that reported statistically significant results being 4.07 (IQR = 1.54-4.43) and a pooled HR of 2.94 (95% CI, 2.08-4.18; p<0.01), though a significant degree of heterogeneity was observed (p<0.01, I2 = 77.4%) (Fig. 5A). This heterogeneity was eliminated when the study by Madhavan et al (26) was excluded. The pooled HR was 3.69 (95% CI, 2.57-5.30; p<0.05) after eliminating the heterogeneity (Fig. 5B). Finally, publication bias of the included studies for OS/RFS/DFS was evaluated by funnel plots (Fig. 6A, B) and Egger's tests. The p value of Egger's test was 0.69 (before elimination of heterogeneity) and 0.48 (after elimination of heterogeneity), suggesting no significant publication bias existed.
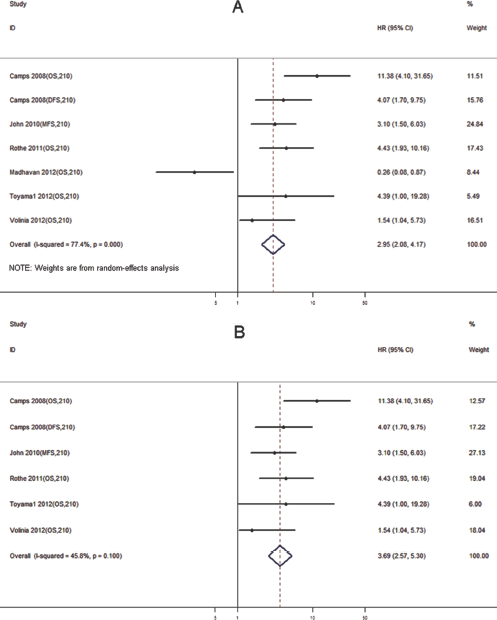
Forest plot of miR-210 expression and survival among breast cancer patients before
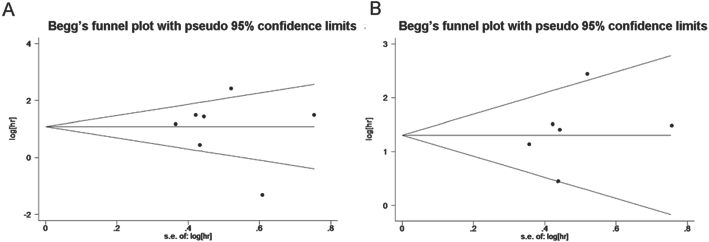
Funnel plot of miR-210 expression and survival among breast cancer patients before
Discussion
Previous reviews have shed light on the importance of associations between miRNAs and various cancers, and some studies have discussed the role of miRNAs in head and neck squamous cell carcinoma, carcinomas of the digestion system, non-small cell lung cancer and colorectal cancer (55, 56). An increasing number of studies have also demonstrated that aberrant expression of miRNAs is associated with prognosis for breast cancer, especially for invasive breast cancer (25). MiRNAs are key players in a wide array of pathological processes, which may partially explain their prognostic associations in breast cancer (42). However, the prognostic role of miRNAs in breast cancer is still a puzzle. Here we gathered more complete literature information and pooled the prognostic outcomes as compared with miRNA expression.
Both miR-21 and miR-210 were found to be highly expressed in multiple kinds of cancerous tissues including lung, pancreas, prostate, stomach, liver, colorectal and ovary cancers (57-59). Recent studies have shown that overexpression of miR-21 increases cell proliferation, migration, invasion and survival in a variety of cancer cell lines (60, 61). The oncogenic role of miR-21 has been shown by targeting several tumor suppressor genes including programmed cell death 4 (62), epithelial-mesenchymal transition breast cancer stem cell phenotypes (16) and extracellular matrix metalloproteinase inducer (15). The biological function described above may affect the relationship between miR-21 expression and cancer outcome. Meanwhile, up-regulation of miR-21 has also been observed in breast cancer tissues compared with nontumor tissues (63), and serum miR-21 was found to play a role in the pathogenesis of tumor (64), providing a clinical utility for miR-21 as an important potential diagnostic biomarker for breast cancer. MiR-210, which is up-regulated in both normal and transformed hypoxic cells has a functional role during tumor initiation (65). Tumors with high miR-210 expression may have a growth advantage via reduced mitochondrial respiration in a hypoxic microenvironment (65). Targets of miR-210 are involved in many processes, including cell proliferation, DNA repair, chromatin remodeling, metabolism and cell migration (8). But the potential diagnostic value of miR-210 in breast cancer has rarely been well elucidated.
In this study, we identified specific miRNAs that appeared repeatedly among the selected classifiers. Most of the protective miRNAs (e.g., the let-7 family) were down-regulated, while the risky miRNAs (e.g., miR-21, miR-210 and miR-155) were up-regulated in breast cancer. Two miRNAs (miR-21 and miR-210) appeared more frequently and accounted for 20.0% and 10.9% of the total miRNAs. By using a meta-analytical approach, we demonstrated that increased expression of miR-21 significantly predicted poorer OS and DFS/RFS, although a significant degree of heterogeneity was observed in the OS group. The results of the ethnicity subgroup analysis indicated that increased miR-21 expression significantly predicted low OS in Asians, and the heterogeneity was considerably reduced. Positive results (HR >2) were obtained in Her2+/ triple-negative type of breast cancer patients, the mean/fixed cutoff value and in the blood/bone marrow group, while subgroup analysis stratified by cutoff value, pathological type and source of material of breast cancer. However, due to limited numbers (n<3) of triple-negative/Her2+ studies, mean cutoff value studies and blood source studies, the results must be interpreted with caution. We also concluded that the pooled HR of miR-210 was higher than that of miR-21 in breast cancer survival. The pooled HR of miR-210 was 2.95 and increased to 3.69 when the study by Madhavan et al (26) was eliminated, suggesting that miR-210 levels have a higher statistical power to detect differences and are likely to be a stronger prognostic biomarker in breast cancer survival.
Nevertheless, the present meta-analysis had a number of limitations and needs to be refined for several reasons. Above all, empirically, HR of less than 1.5 is considered to be a weak prognostic factor (66). The pooled risks of miR-21 for survival outcomes, although statistically significant, were not strong, with an HR value for OS of 1.46 and for DFS/RFS of 1.49. Secondly, a significant degree of heterogeneity was observed among these studies. The heterogeneity was not eliminated after reorganizing subgroup analysis according internal controls, cutoff value, pathological types and source of material from breast cancer patients. Therefore, other factors such as age, sex, pathology and potentially, still other features, might have affected the reliability of the results. Meanwhile, some studies only reported unadjusted HR, which may have led to an overestimation for survival outcomes. The elevated expression of miRNA in serum may provide additional information regarding the host response and prognosis. Few studies detected miRNA expression in serum, and a positive result was obtained in the blood/bone marrow group (HR = 3.76, 95% CI, 1.92-7.37; p<0.01). Therefore, researchers should simultaneously investigate the prognostic value of circulating miRNA levels in breast cancer patients.
Conclusions
In conclusion, this study reviewed the complete literature regarding miRNAs and survival rates for breast cancer and found that high expression of miR-21 and miR-210 was significantly associated with poor survival in patients with breast cancer. However, it is necessary to collect large samples of data, perform stratified analyses and gather data from additional ethnicities to clarify the prognostic role of miR-21 and other miRNAs in breast cancer.
Footnotes
Financial support: This work is funded by National Natural Science Foundation of China (81472920; 81402518), Jiangsu Provincial Special Program of Medical Science (BL2012046) and Changzhou Scientific Project (ZD201315, CE20135050, Cy20130017 and CJ20140050).
Conflict of interest: None declared.
