Abstract
Objectives
This study aimed to compare the epigenetic changes via hypermethylation status of TIMP-3, GSTP-1 and 14-3-3σ genes, between healthy subjects and patients with reversible chronic inflammatory disease, and between healthy subjects and patients with irreversible malignant disease, to highlight the genetic changes that occur in the progression from an inflammatory condition to irreversible genetic changes commonly observed in cancer patients.
Methods
DNA was extracted from the blood of 680 healthy subjects, and tissues and blood of 110 patients with chronic inflammation disease of the gums, as well as neoplastic tissues of 108 breast cancer patients. Methylation-specific polymerase chain reaction (PCR) for TIMP-3, GSTP-1 and 14-3-3σ was performed, and hypermethylation status was analyzed and compared between the 3 groups.
Results
The hypermethylation frequencies of TIMP-3 and GSTP-1 of reversible chronic inflammatory gum disease and the control group were similar, but both were significantly lower than those for malignant disease patients (p<0.0001). The methylation frequency of 14-3-3σ in chronic inflammatory gum disease was higher than in the cancer and control groups (p<0.0001). The methylation of CpG islands in TIMP-3 and GSTP-1 in chronic inflammation patients occurred as frequently as in the control group, but less frequently than in breast cancer patients. However, the epigenetic silencing of 14-3-3σ occurred more frequently in the chronic inflammation group than in cancer patients and healthy controls.
Conclusions
The epigenetic silencing of 14-3-3σ might be essential for chronic inflammatory gum disease. The epigenetic changes presented in chronic inflammation patients might demonstrate an irreversible destruction in the tissues or organs similar to cancer.
Introduction
DNA methylation is a commonly occurring epigenetic modification, which is defined as heritable changes at both genetic and genomic levels without changes to the DNA sequences (1, 2). The mechanisms of epigenetic regulation include DNA methylation, amongst other functions. The methylation of DNA is defined as the covalent attachment of a methyl group to the C5 position of cytosine residues in CpG dinucleotide sequences that are called CpG islands, located in the promoter regions of genes. Methylation of these sites can alter gene transcription, and is involved in normal cellular control of gene expression (3).
The methylated cytosines make up about 3%-4% of all cytosines in normal human DNA (4). The methylation patterns in cancer cells, including breast cancer, bladder cancer and cervical carcinogenesis, are significantly abnormal compared with those of normal tissues and cells, and they typically cause genome-wide hypomethylation and regional hypermethylation (5). This aberrant methylation may have a role in the abnormal expression of certain genes, such as tumor suppressor gene or oncogene (6). The mechanism of this epigenetic change is still unknown. Recently, evidence shows that the hypermethylation of CpG region is closely linked to various conditions such as chronic inflammation, tumorigenesis and intestinal metaplasia (7).
Inflammation is the primary response of the body to different intruders such as infections, injuries and viruses. Normal inflammation is a self-limited process, in which the production of antiinflammatory cytokines follows that of the proinflammatory cytokines. Compared with the normal process of inflammation, chronic inflammation can be regarded as a distorted progression due to the persistence of initiating factors (infection, antigen and tissue injury). It is now widely accepted that persistent inflammation is connected with the incidence and progression of cancer (8-9-10). Based on the complex relationship between inflammation and cancer, a great deal of interest has been directed toward research into the epigenetic changes in inflammation. And there is emerging evidence about epigenetic modulation in tissue-specific chronic inflammation (7, 11-12-13).
Periodontitis is a chronic inflammation for which the outcome is determined by interactions between multiple bacteria and the host response to those bacteria. The inflammation in periodontitis may lead to formation of periodontal pockets, alveolar bone resorption and eventually loss of teeth (14, 15). Owing to the complexity and multifactorial features of periodontitis, epigenetic changes might contribute to the disease development just like it does in carcinogenesis. The possible association between chronic periodontitis and cancer is worth exploring further at the molecular level (16, 17).
Ample research has demonstrated that epigenetic events occur both in the progression of cancer and in chronic inflammation, and that methylation can be considered as an indicator of cancer progression (18-19-20). But methylation as an indicator of periodontitis has rarely been studied. Tissue inhibitors of metalloproteinases-3 (TIMP-3), glutathione transferase class P-1 (GSTP-1) and 14-3-3σ may provide links to chronic inflammation (periodontitis). TIMP-3, binding to the extracellular matrix, is a secreted 24-kDa protein that acts as an inhibitor of metalloproteinases (MMPs). It has been shown that the overexpression of TIMP-3 can induce apoptosis and suppress tumor growth and angiogenesis (21, 22). GSTP-1, an important tumor suppressor gene, is one of the genes that express detoxification enzymes that protect cells from electrophilic or oxidant carcinogens. It is 14-3-3σ that is the only isoform of 14-3-3 to show overexpression after DNA damage (23). The methylation patterns of these 3 genes have been extensively investigated in diagnosing and treating cancers, while research into methylation patterns in periodontitis has yet to be conducted.
The hypermethylation of E-cadherin and cyclooxygenase 2 (COX-2) genes was determined to be related to periodontitis, as demonstrated in our previously published study (24). This present study aimed to investigate and compare the hypermethylation status of TIMP-3, GSTP-1 and 14-3-3σ genes in gingival tissues isolated from periodontitis patients and in neoplastic tissues of breast cancer patients.
Materials and Methods
Selection of control and patient samples
This study utilized 680 randomly obtained blood samples from disease- and periodontitis-free subjects who underwent health screening at the UNIMED Medical Institute, Hong Kong, between September 2011 and March 2013, as a control population. The controlled case comprised 290 female and 390 male healthy subjects, ranging from 18 to 60 years old, who were matched and compared with the disease group (Tab. I). After blood was drawn, these healthy subjects underwent a routine dental examination at UNIMED, and were determined to be free from systemic or chronic disease; current and past nonsmokers; without lymph node swelling; and without temporal mandibular joint disease, soft tissue abnormalities or severe dental caries and furcation involvement or generalized gingival recession. An intraoral soft tissue examination was conducted to assess mean probing depth, dental calculus and bleeding on probing (BOP) (Tab. I).
BIOGRAPHICAL DATA OF HEALTHY CONTROLS AND PERIODONTITIS PATIENTS
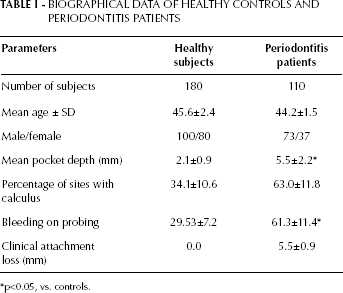
p<0.05, vs. controls.
Measuring the level of inflammation and progression in periodontal disease has been well documented and is accurate; thus chronic periodontitis patients with measurable levels of inflammation were selected as the group to represent reversible chronic inflammatory disease.
From 2011 to 2013, a total of 110 periodontitis patients were recruited from West China College of Stomatology, Sichuan University. Among these subjects, 37 were women and 73 were men, between 18 and 65 years old, with a mean pocket depth of 5.5 mm (Tab. I). They had been suffering from periodontal disease for over 5 years and had received scaling and root planing every 6 months. Patients whose teeth showed third-degree mobility had those teeth extracted. Blood was drawn from these patients, and associated periodontal tissues were sampled during the tooth extraction, for DNA analysis.
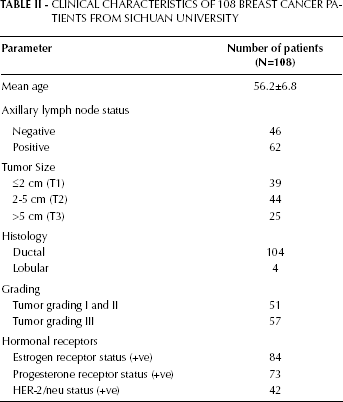
From 2011 to 2013, the specimens from a total of 108 pathologically confirmed invasive ductal carcinoma breast cancer patients (mean age of 56.2) were obtained from Huaxi Hospital of Sichuan University (Tab. II). Based on the advice of the patient's attending oncologist, the women received either 4 or 6 cycles of 5-fluorouracil (600 mg/m2), epirubicin (75 mg/m2) and cyclophosphamide (500 mg/m2) every 3 weeks after modified radical mastectomy was performed. Of 108 patients, 62 women had positive lymph node involvement (1 grade I, 4 grade II, and 57 grade 3). All patients underwent a modified mastectomy. Written informed consent was obtained from all participants, and the study was approved by the ethics committee of Sichuan University (reference number: 20100181110059).
CLINICAL CHARACTERISTICS OF 108 BREAST CANCER PATIENTS FROM SICHUAN UNIVERSITY
Preparation of blood samples
Blood was collected (10 mL) from all 3 groups of subjects, in lithium heparin tubes (Vacuette, Austria), and centrifuged for 10 minutes at 1,500 rpm, after which the plasma was removed. The cell pellet was transferred to a 50-mL centrifuge tube, and red blood cell lysis buffer was added to a final volume of 45 mL. The mixture in the tube was inverted several times and centrifuged again for 10 minutes at 1,500 rpm. The supernatant was discarded, and the cell pellet was washed with 0.9% phosphate-buffered saline (PBS) to be used for DNA extraction.
Sample preparation and tissue collection
DNA from the healthy subjects’ blood cells, periodontitis patients’ blood and tissue samples, and breast cancer patients’ blood and tumour samples was extracted by Geneaid DNA Mini Kit (Tissue) (Geneaid, Taiwan). The provided micropestle was used to grind the tissue to a pulp. Then 200 μL of GT Buffer was added to the tube, and the sample tissue was continually homogenized with grinding. Then 20 μL of proteinase K (10 mg/mL) was added to the sample mixture and mixed by vortexing. The sample was lysed by incubation at 60ºC for 30 minutes, and the tube was inverted every 5 minutes. At this time, the required elution buffer was preheated at 70ºC. Ethanol (200 μL) was added to the sample lysate and mixed immediately by vortexing for 10 seconds. Pipetting was performed to break up the precipitate formed. A GD column was placed in a 2-mL vial. The entire mixture from the previous step (including any precipitate) was applied to the GD column, then centrifuged at 13,000 rpm for 2 minutes. The vial containing the flow-through was discarded, and the GD column was transferred in a new 2-mL vial. Five hundred microliters of wash buffer (ethanol added) was added into the column and was centrifuged at 13,000 rpm for 30 seconds. The flow-through was discarded, and the GD column was placed back into the vial. The wash step by adding wash buffer was performed once again. The flow-through was, again, discarded, and the GD column was placed back in the vial and was centrifuged at full speed for 3 minutes to dry the column matrix. The dry GD column was transferred to a clean 1.5-mL microcentrifuge tube. Preheated elution buffer (100 μL) was added into the center of the column matrix and was left to stand for 5 minutes until the elution buffer was absorbed by the matrix. Centrifugation at 13,000 rpm was performed for 30 seconds to elute purified DNA.
Methylation-specific polymerase chain reaction
The extracted DNA from blood and tissue was modified by CpG DNA Modification Kit (Chemicon International, Temecula, CA, USA). The specific hypermethylated primers for each gene were used for polymerase chain reaction (PCR). The PCR mixture consisted of 1X PCR buffer (20 mM Tris-HCl, pH 8.4; 50 mM KCl), 1.5 mM MgCl2, 0.2 mM dNTPs, 40 pmol sense and antisense primers, and 0.75 units of Taq DNA polymerase. Initial denaturation at 94°C for 5 minutes was followed by 50 cycles of denaturation at 94°C for 30 seconds, annealing at 57°C for both hypermethylated and unmethylated sequences for 30 seconds and extension at 72°C for 30 seconds, then a final extension at 72°C for 10 minutes. The 3 genes of PCR products were analyzed on 2% agarose gel stained with ethidium bromide.
The sense and antisense primers for the methylation of (i) TIMP-3 were 5’-CGTTTCGTTATTTTTTGTTTTCGGTTTC-3’ and 5’-CCGAAAACCCCGCCTCG-3’; (ii) GSTP-1, 5’-TTCGGGGTGTAGCGGTCGTC-3’ and 5’-GCCCCAATA CTAAATCACGACG-3’ and (iii) 14-3-3σ, 5’-GGTTTTTTCGGTTAGTTGCGCGGCG-3’ and 5’-CCAACGAAAACCTCG CGACCTCCG-3’.
A β-actin was used with the sense primer 5’-CCTCTATGCCAACACAGTGC-3’ and antisense primer 5’-ATAC TCCTGCTTGCTGATCC-3'and considered as constitutive housekeeping genes for PCR and used to compare changes in specific gene expressions. The PCR products were analyzed on 2% agarose gel stained with ethidium bromide. The gel was visualized using a Dolphin-DOC ultraviolet illuminator (Wealtec, South Africa).
Statistical Analysis
Comparisons of the relative risk of hypermethylation status of TIMP-3, GSTP-1 and 14-3-3σ between healthy subjects and those with irreversible malignant disease (cancer patients), and between healthy subjects and those with reversible inflammatory disease (periodontitis patients) were analyzed. A chi-square test was performed to analyze the distribution of hypermethylation in healthy controls vs. patients with chronic periodontitis, and between healthy controls and breast cancer patients, using SPSS 17.0 (SPSS Inc., Chicago, IL, USA).
Results
In this study, comparisons of epigenetic changes were made between 2 groups: healthy vs. periodontitis subjects and healthy vs. breast cancer patients, using blood only for healthy individuals and blood and tissue samples obtained from the breast cancer patients and periodontitis groups studied. There were no significant differences in hypermethylation status when blood and tissue samples were compared. The results of the hypermethylation status of TIMP-3, GSTP-1 and 14-3-3σ genes from blood and tissues samples obtained from periodontitis groups were similar (p>0.05). Similarly, using samples of tissue and blood from the breast cancer groups, the hypermethylation status of TIMP-3, GSTP-1 and 14-3-3σ genes showed similar results (p>0.05). Table III shows the status of TIMP-3, GSTP-1 and 14-3-3σ for blood drawn from healthy controls and chronic periodontitis and breast cancer groups.
STATUS OF TIMP-3, GSTP-1 AND 14-3-3σ HYPERMETHYLATION IN HEALTHY SUBJECTS, PERIODONTITIS PATIENTS AND BREAST CANCER PATIENTS (MEASURED FROM BLOOD)
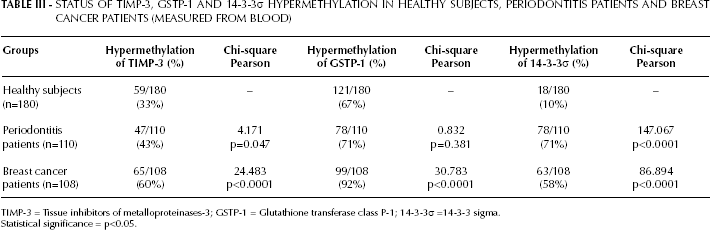
TIMP-3 = Tissue inhibitors of metalloproteinases-3; GSTP-1 = Glutathione transferase class P-1; 14-3-3σ =14-3-3 sigma.
Statistical significance = p<0.05.
For breast cancer subjects, hypermethylation of TIMP-3 was observed to be 58% in breast cancer tissue and 60% in blood, while hypermethylation of GSTP-1 was expressed at 88% in tissue and 92% in cancerous breast patient blood (Tab. IV). The methylation frequency of 14-3-3σ was 56% and 58% in tissue and blood, respectively (Tab. IV).
COMPARISON OF TIMP-3, GSTP-1 AND 14-3-3σ HYPERMETHYLATION BETWEEN THE TISSUE AND BLOOD SAMPLES IN PERIODONTITIS PATIENTS AND BREAST CANCER PATIENTS
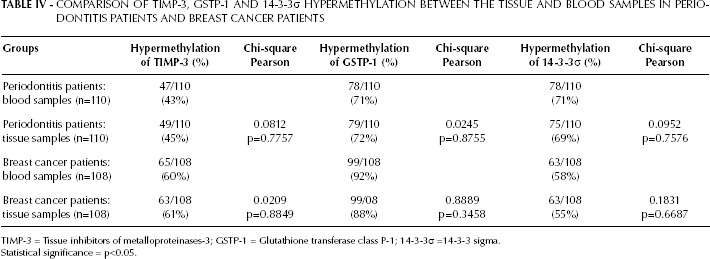
TIMP-3 = Tissue inhibitors of metalloproteinases-3; GSTP-1 = Glutathione transferase class P-1; 14-3-3σ =14-3-3 sigma.
Statistical significance = p<0.05.
TIMP-3 was expressed at 45% and 43% in tissue and blood of chronic periodontitis patients, respectively (Tab. IV). Hypermethylation of GSTP-1 in tissue was 72% and 71% in blood (Tab. IV). Similarly, methylation frequency of 14-3-3σ was overexpressed as 69% and 71% in tissue and blood.
In healthy patients, the methylation frequency of TIMP-3, GSTP-1 and 14-3-3σ was 33%, 67% and 10%, respectively, measured in blood (Tab. III).
The hypermethylation frequency of TIMP-3 (X2=4.171; p=0.047) and GSTP-1 (X2=0.832; p=0.381) was similar between the periodontitis and control groups. But the hypermethylation status of TIMP-3 (X2=24.483; p<0.0001) and GSTP-1 (X2=30.783; p<0.0001) in breast cancer patients was shown to be statistically significantly different in comparison with the periodontitis and control groups (Tab. III). A comparison of the hypermethylation frequency of 14-3-3σ between the periodontitis and control groups (X2 =147.067; p<0.0001), and the breast cancer patient and control groups (X2=86.894; p<0.0001) proved to be similar (Tab. III). The methylation frequency of 14-3-3σ in periodontitis patients (71%) was higher than that in the breast cancer patient group (58%) and the healthy control group (10%) (Tab. III). The methylation of CpG islands in TIMP-3 and GSTP-1 in periodontitis patients occurred as frequently as it did in the control group, but less frequently in breast cancer patients. In contrast, the epigenetic silencing of 14-3-3σ occurred more frequently in the periodontitis group than in breast cancer patients and the control group.
Discussion
In this study, the hypermethylation patterns of TIMP-3, GSTP-1 and 14-3-3σ changed in the periodontitis group in comparison with the breast cancer patient group and healthy control group. The hypermethylation of TIMP-3 showed a similar frequency in periodontitis patients compared with healthy controls, but was less than that in the cancer group. A similar change in hypermethylation was also noted for GSTP-1 (Tab. I).
Typically, DNA methylation undergoes a simultaneous global hypomethylation and a local hypermethylation of the CpG islands rich in the promoter region of genes in cancer. Hypomethylation of promoter regions can result in gene reactivation, up-regulation or overexpression, while promoters of transcriptionally silenced or suppressed genes are normally hypermethylated (25). The results of this study demonstrated that the hypermethylation frequencies of the 3 factors in breast cancer patients were higher than in the periodontitis and control groups (Tab. I). The findings of this study were in line with the results of previously conducted studies. The aberrant functions of DNA methyltransferase enzymes result in altered DNA methylation patterns in cancer, which can be used as a valuable marker of malignant transformation. Methylation of the TIMP-3 gene was shown to predict prolonged disease-free interval, and hypermethylation of the GSTP-1 gene was also shown to be a highly specific marker of prostate cancer (26, 27). The expression of 14-3-3σ can also be down-regulated by aberrant CpG methylation in breast cancer (28).
However, the occurrence of hypermethylation of these 3 genes in the periodontitis group requires further attention. The hypermethylation of TIMP-3 and GSTP-1 occurs in periodontitis patients and healthy subjects with a similar frequency. However, methylation of 14-3-3σ is much higher in periodontal tissues. Periodontitis is a chronic inflammation, usually initiated by pathogenic microflora, while host factors contribute to the progression (29). The inflammatory responses during periodontitis may actually be a trigger for epigenetic reprogramming by producing reactive nitrogen species (RNS) and reactive oxygen species (ROS), which mediate many important physiological functions including modification of extracellular matrix, apoptosis, cell proliferation, repair and immune modulation. But persistent production of RNS and ROS can lead to DNA damage in proliferating cells (9). There are 2 typical types of chemical modifications of methylated DNA: oxidative stress and reactive halogen compounds (13). Oxidative stress can oxidize methyl groups through agents such as ROS, RNS and nitric oxide (NO), which leads to loss of methylation. The other type of DNA methylation alteration is caused by reactive halogen compounds which can lead to an increase of methylation. Both DNA methylation modifications contribute to disruption of routine epigenetic events involved in activating and silencing of genes.
There might be an explanation for what was observed in this study. The methylation data for 14-3-3σ differed in comparison with TIMP-3 and GSTP-1. Its percentage of hypermethylation frequency in the periodontitis group was the highest, even much higher than in the breast cancer group, which might indicate that the functions of this gene could be silenced. Evidence shows that 14-3-3σ is a negative regulator of cell cycle at several key points, and epigenetic aberration of 14-3-3σ may be linked to carcinogenesis (30). Considering the relationship between chronic inflammation and cancer, as well as the findings of our study, periodontal disease might be related to cancer risk through plausible biological pathways.
The TIMP-3 and GSTP-1 are also critical for normal tissue functions. The balance between TIMPs and MMPs plays a critical role in maintaining the integrity of healthy tissues. TIMP levels are normally higher than MMP levels in healthy periodontal tissue, while its levels are lower than MMP levels in periodontitis (31). GSTP-1 belongs to the family of phase II detoxification enzymes that protect DNA against genome damage mediated by oxidants or electrophiles from inflammation exposures (32). The hypermethylation patterns of these 2 factors indicate that the epigenetic events might contribute to the regulation of periodontitis.
Some studies have already demonstrated that epigenetic aberrations may be connected to tumorigenesis, and that this is a characteristic feature in cancer (4), and periodontitis has recently been implicated as being related to the significant increase in overall cancer risk (33, 34). Indeed, growing evidence has shown that many malignancies are initiated by infections (35-36-37). Repeated damage and regeneration of tissue under the presence of RNS and ROS produced by inflammatory cells can result in irreversible genomic alterations including point mutations, deletions or rearrangements (9). The results of the present study demonstrated that epigenetic changes also occurred in chronic inflammation, resembling a similar pattern of disease development to that that occurs in cancer patients. Hypermethylation might contribute to periodontitis, which could cause damage to adjacent tissue, the same as with cancer.
Conclusion
The present data confirm that hypermethylation in the TIMP-3, GSTP-1 and 14-3-3σ genes is related to chronic inflammatory disease (chronic periodontitis). Further studies are needed to explore CpG alterations of more, related factors, which might be important for diagnosing and/or treating periodontal diseases.
