Abstract
Numerous epidemiological studies have evaluated the association between transforming growth factor beta receptor type 1 (TGFBR1) polymorphisms and the risk of cancer; however, the results remain inconclusive and controversial. To determine the association between breast cancer risk and the *6A polymorphism of the TGFBR1 gene, a case-control study of 280 breast cancer patients and 280 controls was performed in Iranian women. Our study demonstrates that women who carry the TGFBR1*6A allele are at lower risk of developing breast cancer. The highest protection against breast cancer was observed in 6A/6A homozygotes (OR = 0.32, p = 0.04). A lower frequency of the TGFBR1*6A allele in breast cancer patients may be an important genetic determinant that contributes to a lower risk of breast cancer in Iranian women. The results also showed that the allelic length of TGFBR1 polymorphisms had no significant association with the age at onset or the grade of disease, nor with the expression of progesterone and estrogen receptors and HER2.
Introduction
Worldwide, breast cancer is currently the most common cancer diagnosed in women, with over 1.3 million new cases per year, and the second leading cause of cancer deaths among women. The American Cancer Society estimates that 232,340 (29%) new cases of invasive breast cancer will be diagnosed among women in the United States in 2013 (1). In Iran (as well as in the population of Isfahan), breast cancer is the most common cancer in women, comprising 24.4% of all cancer malignancies among Iranian females (2). The incidence of breast cancer is growing in Iran and affects women at least 1 decade earlier than in the developed countries (3).
The transforming growth factor beta (TGF-β) signaling pathway has been a main focus of extensive research since it was initially discovered in 1981 (4). This signaling pathway is a very important modulator of many biological processes, such as differentiation, cell proliferation, morphogenesis, migration and apoptosis (5). The TGF-β signaling pathway plays a crucial role as a tumor suppressor in the early stages of carcinogenesis and as a pro-oncogenic mediator in the late stages. As overall TGF-β signaling could also be determined by genetic polymorphisms in many TGF-β pathway genes, growing epidemiological evidence has referred to the consequences of common variants of the TGF-β pathway for the cancer risk. TGF-β receptor type 1 (TGFBR1), the central propagator of the TGF-β signaling pathway, has been the hot spot of research.
TGFBR1 is a serine-threonine protein kinase that mediates the growth-inhibitory signals of TGF-β through a complex with TGFBR2. TGFBR1 contains a common microsatellite polymorphism at the 3′ end of the exon 1 coding sequence, resulting in the deletion of 3 alanines (GCG) within a stretch (TGFBR1*6A) of 9 alanines (TGFBR1*9A), corresponding to a variable stretch of alanine residue (6). The stretch of 9 alanines is part of the TGFBR1 signal peptide and is responsible for targeting the receptor to the membrane (7, 8). Functional investigations have suggested that TGFBR1*6A is a less effective (and impaired) mediator of TGF-β antiproliferative signals compared with the TGFBR1*9A allele (intact TGFBR1), and TGFBR1*6A has been proven to decrease TGF-β growth inhibitory signaling activity (9, 10). Furthermore, it had been shown that *6A may switch TGF-β growth-inhibitory signals to growth-stimulatory signals in cancer cells, enhancing the migration and invasion of breast cancer cells (7, 11). A number of epidemiological studies have investigated the association between TGFBR1*6A and cancer risk, but the results are somewhat controversial. Meta-analysis studies suggested that TGFBR1*6A acts as a low-penetrance tumor susceptibility allele in the development of colon, cervix, breast and ovarian cancer (9, 10, 12-16).
In the present study, we investigated the frequency of the TGFBR1*6A variant in Iranian breast cancer patients and control Iranian women to clarify the effect of this polymorphism on the risk of breast cancer in the Iranian population.
Materials and Methods
Sample Collection and Genotyping
Peripheral blood samples were collected from 280 women with breast cancer (Omid/Seyedoshohada Cancer Hospital, Isfahan University of Medical Sciences, Iran) and 280 age-matched healthy blood donors (controls) aged over 45 years. The breast cancer patients were in the age range of 26-87 years. Control subjects were randomly selected among women visiting hospitals for normal health checks (Clinical Laboratory, Alzahra Hospital, Medical Sciences University of Isfahan, Iran). DNA was extracted by the salting-out method consistent with the Fisher protocol. Extracted DNA was diluted in TE (10 mM Tris-HCl, 1 mM EDTA, pH = 8.0) and maintained at −20°C for additional analysis.
The TGFBR1 exon 1 region known to contain the GCG repeats was amplified by the PCR technique using the forward primer 5′-AGG TTT GCT GGG GTG AGG CAG-3′ and the reverse primer 5′-ATG TTT GAG AAA GAG CAG GAG-3′. A 25 μL sample of the PCR mixture contained 100-250 ng of genomic DNA, 0.2 μM each of the forward and reverse primers, 200 μM dNTPs, 5 mM MgCl2, 2.5 μL 10 X PCR buffer AMS, 2.5 μL DMSO, 5 μL betaine and 0.04 U SmarTaq DNA polymerase, all obtained from CinnaGen Inc, Iran. PCR conditions were as follows: 1 cycle of 97°C for 5 minutes followed by 33 cycles of 97°C for 1 minute, 57°C for 1 minute, 72°C for 1 minute and finally 1 cycle of 72°C for 10 minutes. Polyacrylamide gel (10%) fractionation was used to determine the length of the amplified fragments after silver staining. Alleles of variable sizes were selected and sequenced directly. The sequenced alleles were used as allele-specific markers for exact determination of the number of GCG repeats in other samples.
Statistical Analysis
Statistical analyses were carried out using the SISA (http://www.home.clera.net/sisa/) and SPSS (ver. 19) software. The association between different alleles and the risk of developing breast cancer was examined by Pearson's chi-square and odd ratio (OR) tests. P values <0.05 were considered statistically significant. ORs with 95% confidence intervals (CI) were calculated to assess the presence or absence of correlations between specific factors and therefore the risk of developing breast cancer.
Results
Polymorphism of TGFBR1
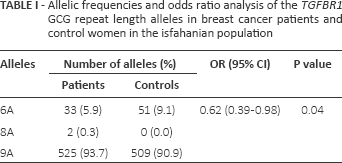
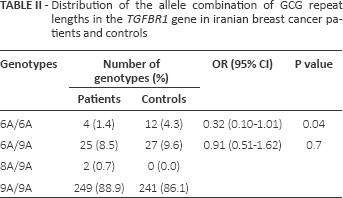
Three different alleles of the GCG (alanine) repeat in the range of 6-9 were identified in exon 1 of the TGFBR1 gene (Tab. I). The most frequent allele in both the case and control groups was the (GCG)9 (9A) repeat with frequencies of 93.75% and 90.9%, respectively. In our study population, 4 different genotypes were observed in both groups. The frequency distribution of the genotypes in patients and controls is listed in Table II. The most common genotype in patients (88.9%) and controls (86%) was homozygous 9A.
Allelic frequencies and odds ratio analysis of the TGFBR1 GCG repeat length alleles in breast cancer patients and control women in the isfahanian population
Distribution of the allele combination of gcg repeat lengths in the TGFBR1 gene in iranian breast cancer patients and controls
GCG Repeat Length and the risk of Breast Cancer
Compared with the control group, patients had a significantly lower frequency of the 6A allele. The mean frequency of the 6A allele was 5.9% in patients and 9.1% in controls. Our results demonstrate that women carrying the 6A allele are at a significantly lower risk of developing breast cancer with an OR of 0.62 (p = 0.04) (Tab. I). Homozygous 6A conferred the greatest protective effect against breast cancer with an OR of 0.32 (p = 0.04) (Tab. II). The correlation between the allelic length of GCG repeats in TGFBR1 and the age at onset, grade of disease and expression of estrogen receptors, progesterone receptors and ErbB2 (HER2) was also analyzed. No significant association was observed between the allelic length of TGFBR1 polymorphisms and age at onset, lymph node metastasis or expression status of estrogen or progesterone receptors (data not shown). To obtain precise results, additional studies with a larger sample cohort are needed.
Discussion
As one of the key effectors of TGF-β signaling, TGFBR1 mediates the growth-inhibitory signals of TGF-β through a complex with TGFBR2. Over the past few years, epidemiological studies have indicated that genetic variants affecting TGF-β production or signaling components may be related to the overall risk and survival of breast cancer patients. Several single-nucleotide polymorphisms (SNPs) and/or mutations in TGFBR1 have been associated with increased breast cancer risk and remain to be thoroughly evaluated (17). Until now, most epidemiological studies have confirmed an association between the common polymorphism in TGFBR1 (TGFBR1*6A) and an increased risk of breast cancer (12, 13, 17, 18), but others have failed to establish any correlation (19-21).
In a population-based case-control study in Iran, we studied the GCG allelic length polymorphism in exon 1 of the TGFBR1 gene and its association with breast cancer risk. In contrast to previous studies, our results suggest a negative association between the (GCG)6 allele and breast cancer risk. Women carrying this allele, especially those with a homozygous genotype (OR = 0.32), are at significantly lower risk of developing breast cancer. Compared to previous reports, our results show much higher frequencies of the 6A genotype in controls. The frequency of the 6A genotype in Isfahanian controls was 4.3%, compared to 0-1.8% in other populations (12-18). We have previously observed such discrepancies in allelic distribution and cancer association with CA repeats in the EGFR and IGF-1 genes and AAAG repeats in the ERRγ gene (22-24). Such inconsistencies might be due to racial differences as well as complex expressions of these genes. There is also a possibility of environmental influences which may give rise to such ethnicity-specific genetic variation.
In addition, it is believed that the signal peptides of wild-type TGFBR1 and TGFBR1*6A were cleaved at the same site, leading to identical mature receptors and therefore the biological effects of *6A are mediated by the signal sequence instead of the mature receptor (7). Once cleavage has occurred, the signal sequence remains within the cytoplasm, where it is possible to regulate specific gene expression or induce other cellular functions. Hence, *6A is the first example of signal sequence secondary signaling in cancer (8). The secondary signals generated by the *6A signal sequence modulate the gene expression of known effectors of the TGF-β-induced cell cycle arrest (25) and also have an effect on the expression levels of TGF-β receptors and SMADs. These mechanisms have been shown to underlie the cancer cells’ loss of TGF-β-mediated growth inhibition (26). It is unclear why the same allele length in different populations has different impact on cancer etiology. One possible explanation could be individual differences in gene expression patterns or genetic variation in other related genes. Despite all the evidence, the actual mechanism of this polymorphism remains unknown and controversial.
In summary, our results suggest that the TGFBR1*6A polymorphism may be associated with a lower risk of breast cancer in Iranian women. The relation between the *6A polymorphism and the risk of breast cancer seems to differ according to race and ethnicity. However, further studies are required to ascertain a definite relationship between this variant and the risk of cancer development.
This study was also coupled with the assessment of clinical data in the context of the age at onset or grade of the disease, as well as the expression of progesterone and estrogen receptors and HER2. Our results revealed that there is no statistically significant correlation between the (GCG)6 allele and these clinical parameters.
Our observations are based on a small sample size. Multiple testing may further affect the study power. Thus, the study results should be interpreted with caution.
Footnotes
Acknowledgment
The authors wish to thank all patients who contributed to this investigation.
Financial support: This work was supported by the University of Isfahan.
Conflict of interest: The authors have no financial or non-financial competing interests.
