Abstract
Purpose
To investigate the impact of the cytidine deaminase (CDA) A79C polymorphism on both the response to gemcitabine in non-small cell lung cancer (NSCLC) patients and the risk of hematologic toxicities in patients bearing any kind of cancer taking gemcitabine.
Methods
The PubMed and Embase databases were searched from the first available article to January 2013. Eligible studies included clinical trials that contained the keywords “gemcitabine” or “cytidine deaminase” and information about response rate of NSCLC patients or hematologic toxicities in patients with any kind of cancer. Relative risk (RR) of different genotypes and 95% confidence intervals (CI) were calculated.
Results
A total of 7 articles (623 patients from 6 studies) were included. The results showed that patients with wild type CDA (AA and AC) had a significantly lower rate of severe anemia than the homozygote mutant type CC (RR=0.308; 95%CI, 0.113-0.021, p=0.021). However, the rate of severe neutropenia, thrombocytopenia, and the response rate were identical between different CDA genotypes.
Conclusion
The A79C CDA polymorphism did not show a significant impact on the response rate to gemcitabine in NSCLC patients, while the wild type CDA genotype was indeed correlated to a lower rate of incidence of severe anemia in patients taking gemcitabine.
Introduction
Gemcitabine [2′,2′-difluoro-2′-deoxycytidine (dFdC)] is a prodrug activated in cells to become gemcitabine monophosphate, and further converted to dFdC-diphosphate and dFdC-triphosphate. The dFdC-diphosphate is an inhibitor of the ribonucleotide reductase M1 (RRM1) and can decrease the deoxyribonucleotide pools, which are necessary for DNA synthesis. The triphosphate form, dFdC-triphosphate, is incorporated into the DNA, leading to the termination of chain elongation. However, more than 90% of the gemcitabine administered is inactivated by cytidine deaminase (CDA) before being phosphorylated (1, 2).
Gemcitabine is widely used as an anti-neoplastic drug in several types of cancers, such as pancreatic, lung, and breast cancers. The toxicity profile includes nausea, asthenia, myelotoxicity, and reversible hepatic disfunction (3). Some adverse effects, especially severe myelotoxicity, which may induce severe anemia, thrombocytopenia, and neutropenia, vary wildly among individuals. Therefore it is very important to distinguish the group of patients that would probably suffer severe myelotoxicity and take preventive measures.
CDA is an enzyme encoded by the CDA gene (4, 5). Numerous studies have shown that CDA activity may affect clinical outcome (6) and may induce severe adverse reactions (7). The encoded protein forms a homotetramer that catalyzes the irreversible hydrolytic deamination of cytidine and deoxycytidine into, respectively, uridine and deoxyuridine. The human CDA gene is prone to be polymorphic (8), and this characteristic is associated with gemcitabine chemosensitivity/resistance (9); thus, we here focused on the frequency of the CDA gene polymorphism in our patients.
Three single nucleotide polymorphisms (SNPs) in the CDA-encoding gene, CDA 79A>C (rs2072671, Lys27Gln), CDA 208G>A (rs60369023, Ala70Thr), and CDA C435T (rs1048977), have been reported to be correlated with different in vitro activities and clinical outcomes of gemcitabine (10-11-12-13). The CDA 208G>A (rs60369023) SNP is related to reduced enzymatic activity with subsequent reduced drug clearance, increased drug exposure, and increased incidence of neutropenia (13, 14). However, it was only found in Asian patients and was detected only in a small number of studies. The A79C (Lys27Gln) SNP has been proposed as being a predictor of gemcitabine treatment response and toxicity (15), and can affect the CDA activity (12). Therefore, understanding the association between the CDA A79C polymorphism and gemcitabine clinical effects may be beneficial for individualized chemotherapy. Although some studies have investigated this issue, no consensus was reached. Among other studies, Ludovini et al found no significant correlation with clinical benefit and toxicity (16), while Tebaldi et al found that the CDA Lys27Lys polymorphism significantly correlated with better clinical benefit and more serious neutropenia and thrombocytopenia (17). However, Tanaka et al found that individuals with the wild type genotype had better clinical benefit and less toxicity in locally advanced pancreatic cancer patients (18).
Since no specific systematic review or meta-analysis has provided a comprehensive assessment on the association between the CDA79A>C (rs2072671) polymorphism and the clinical response of NSCLC patients or their risk of myelotoxicity, in the present work we intended to fill this gap.
Materials and Methods
Selection of studies
Literature search
A systematic literature search was performed in PubMed and Embase databases for the time period ranging from the first available article to January 2013. Publications were retrieved using the terms “gemcitabine”, in combination with the key words “cytidine deaminase” or “CDA”. Publications meeting the following criteria were included: (a) all patients included in the analysis of response had histologically proven NSCLC; (b) all patients included in the analysis of hematological toxicities had histologically or cytologically proven cancers with no limitation on the pathogenic sites; (c) trials applied gemcitabine-based chemotherapy as first line therapy; (d) trials studied the CDA A79C gene polymorphism and the data of response rate or hematology toxicity, stratified by polymorphisms, could be obtained or derived from the original article or the corresponding author; (e) patients had a measurable clinical stage III B or IV disease, an Eastern Cooperative Oncology Group performance status ≤1, were aged >18 years, had a life expectancy of >3 months, had an adequate bone marrow reserve (leukocyte count >4.0×109/L, platelet count ≥100×109/L), and had an adequate liver and renal function. Studies were excluded in any of the following cases: (a) the data of response rate in NSCLC or hematology toxicity stratified by polymorphisms could not be extracted; (b) the data could not be pooled; (c) repeated publications, abstracts, letters, or review articles. Since 2 articles (19, 20) described the same study, but showed different data (the data on the response was published in the later article [19], while the earlier one [20] published the data about hematology toxicities), we decided to include both articles (counting only once the patients included) in order to analyze both clinical response and drug toxicity.
Data extraction
Data were manually and independently extracted by 2 authors (Hui Li and Xiangling Wang), blind to each other's work, using the same data recording form. After exaction, all results were reviewed and compared by the third author (Xiuwen Wang) and discrepancies between data were discussed and solved until consensus. When necessary, we tried to contact the authors for obtaining data related to some of the studies. The following information was collected for each study: first author's name, publication year, country, median age of the subjects included, sample size, studied polymorphisms, disease and disease stage, and chemotherapeutic schedule and dose (Tab I). Clearly, the response rate of NSCLC patients and the incidence rates of severe hematologic toxicities in patients with any type of cancer and with different CDA genotypes were also extracted. The tumor response was evaluated after every 2 cycles, according to the RECIST criteria; responders included complete responders (CR) and partial responders (PR) while non-responders included patients with stable disease (SD) and progressive disease (PD). To evaluate the severity of the hematologic toxicity we collected the frequency of the following adverse events: neutropenia, anemia, and thrombocytopenia, all only in terms of high-grade events (grades 3-4), according to the National Cancer Institute's common toxicity criteria (NCICTC) (version 2 or 3; http://ctep.cancer.gov) or the WHO toxicity criteria.
GENERAL CHARACTERISTICS OF THE STUDIES
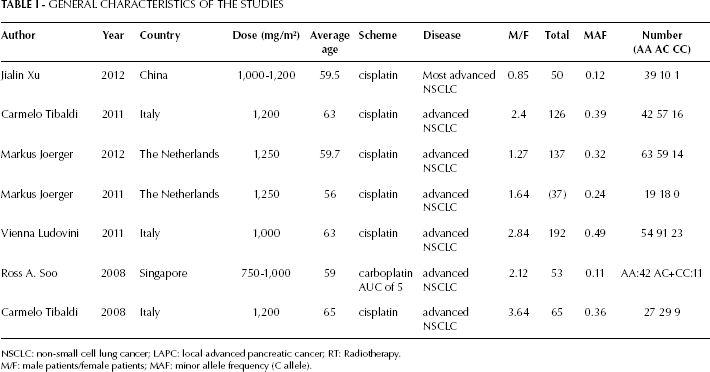
NSCLC: non-small cell lung cancer; LAPC: local advanced pancreatic cancer; RT: Radiotherapy.
M/F: male patients/female patients; MAF: minor allele frequency (C allele).
Statistical Analysis
In each study, the genotype distribution in the population was checked for departure from the Hardy—Weinberg equilibrium (HWE) using the chi-square goodness of fit.
For the incidences of response and toxicity, we calculated the relative risks (RRs) with confidence intervals (CIs) between different types of polymorphisms. We extracted the number of patients reaching a response (both PR and CR) and the number of total NSCLC patients with each polymorphism from all the clinical trials included. For the calculation of the relative risk of the 3 types of hematologic toxicities between different SNPs, we extracted the total patients' number and the number of patients suffering each kind of hematologic toxicity, and we stratified by CDA genotypes, independent of the tumor category. We had to exclude studies reporting no events during a treatment. We examined heterogeneity in results across studies using the Cochrane's Q statistic, and we quantified inconsistency with the I2 statistic [100% × (Q – df)/Q], which represents the percentage of total variation across studies that is attributable to heterogeneity rather than chance (21). A p value of less than 0.10 was considered as indicative of substantial heterogeneity. When substantial heterogeneity was not observed, the pooled estimate was calculated based on the fixed-effects model, otherwise potential sources were investigated by sensitivity analyses and the pooled estimate was calculated based on the random-effects model. To evaluate the publication bias the symmetry of funnel plots was visually inspected, and to further evaluate publication bias, the Begg's test (22) and Egger's test (23) were also performed using Stata 12.0 software (StataCorp LP, College Station, USA). We also conducted stratified analyses by ethnicity and type of partner drug: cisplatin or carboplatin. A two-tailed p value of less than 0.05 was considered statistically significant. All statistical analyses were performed using STATA v.12.0 and Revman v.5.2.
Results
General considerations
Our search yielded a total of 35 potentially relevant studies by abstracts. The selection process is represented in Figure 1. We carefully screened each one of the 35 trials: 28 trials did not present data about the response in NSCLC patients or the incidence of hematologic toxicities stratified by CDA A79C polymorphisms. Finally, a total of 623 cases included in the 7 remaining articles were evaluated (16, 17, 19, 20, 24-26). The baseline characteristics of the patients included in each trial are presented in Table I. We could calculate the allele frequency for each trial, other than one (26). The trials were in HWE (27), as illustrated in Table I. The 6 eligible studies (since 2 articles referred to the same study) were performed in 4 countries: 3 trials were from Italy, 1 from the Netherlands, 1 from Singapore, and 1 from China. We obtained the data about chemotherapy schemes and doses, disease sites, average age of the patients, and the male/female proportions (Tab. I).
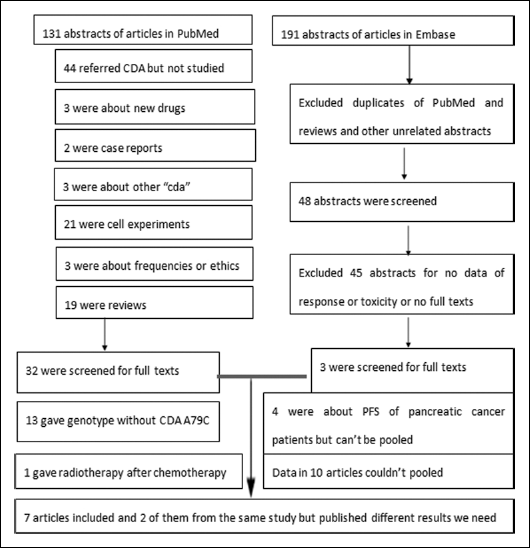
The selection process.
In the trials we selected, the response analysis was reported only for NSCLC patients, but when analyzing the hematologic toxicity we did not limit the disease sites. One trial considered for the hematologic toxicity analyses included, among others, patients with lung disease, gynecologic cancer, and sarcoma, but most patients had advanced NSCLC (24). In addition, 2 trials were carried out in Asia and had different allele frequencies (24, 26). Therefore, 4 trials were selected for inclusion in the meta-analysis of the response of NSCLC patients (16, 19, 25, 26), 4 trials were selected for analysis of the relationship between neutropenia and the A79C CDA polymorphisms (20, 24, 25, 26), 3 trials were selected for the analysis of the relationship between thrombocytopenia and the polymorphism (17, 24, 25), and 2 trials were selected for the analysis of the relationship between anemia and the polymorphism (17, 25).
According to the search results, patients were either chemotherapy-naive or previous therapy must had been completed at least 21 days before study entry with full resolution of toxicities. Patients were treated with concomitant chemotherapy of gemcitabine and cisplatin, except for one trial in which patients were treated with carboplatin (26). The dose of gemcitabine was 1,000-1,250 mg/m2, administered i.v. on days 1 and 8, with cisplatin 75-80 mg/m2 infused on day 1, or 750-1,000mg/m2 when combined with carboplatin of AUC 5 mg min/mL on day 1, every 3 weeks for a maximum of 6 courses.
Quality of the studies
For quality analysis purposes, we attempted to look at differences in both RR of responses rates and hematologic toxicity incidence rates based on the type of partner drug used and patients' ethnicity. However, no statistically significant differences were found (p>0.05, results not shown).
The RR of the response rates of advanced NSCLC patients with different CDA A79C genotypes
As the C allele was the minor allele frequency (MAF), the CC genotype only characterized few cases. We compared 2 types of genotypes: AA against AC and CC, and AA against AC. The fixed-effects model was applied and no publication bias was detected by the inverted funnel plot or the Begg's and Egger's tests in either of the 2 analyses (data not shown).
We found that the AA genotype had a higher response rate than AC and CC, but this difference did not reach statistical significance (AA vs AC+CC, RR=1.218, 95% CI: 0.912-1.627, p for heterogeneity was respectively p=0.18 and p=0.22, I2=32%, Fig. 2A). In this analysis, 4 trials and a total of 422 patients were included. Furthermore, we excluded the Asian carboplatin-using arm (Ross A. Soo 2008) (26) from the trials due to the potential difference in response associated with carboplatin use or ethnicity; nevertheless, the result did not significantly change (data not shown).
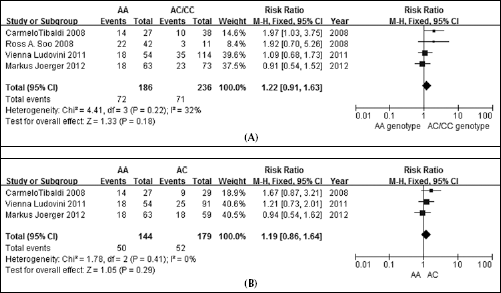
A. Response in NSCLC patients: AA versus AC/CC genotypes.
From the other pooled result (AA vs AC), we found that AA did not have a significantly higher response rate than AC alone: the RR was 1.188, 95% CI: 0.862-1.637, p for heterogeneity was respectively p=0.29 and p=0.41, I2=0% (Fig. 2B).
Relative risk of bone marrow toxicity events
The RRs of the selected bone marrow events were calculated for high-grade events (grades 3 and 4). For RR calculation we considered all trials that could be pooled, including those of different pathologies. Figures 3-5 show the results for RR of high-grade neutropenia, thrombocytopenia, and anemia stratified by CDA A79C gene type.
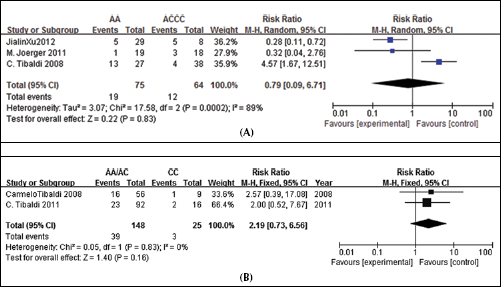
A. Neutropenia: AA versus AC/CC genotypes.
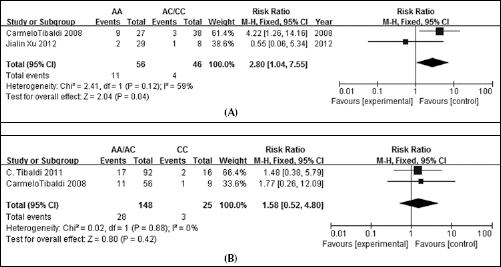
A. Thrombocytopenia: AA versus AC/CC genotypes.
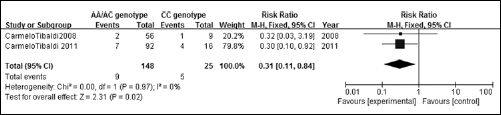
Anemia: AA/AC versus CC genotypes.
Severe neutropenia
A total of 4 trials, including 247 patients, were related to our focus on severe neutropenia (17, 20, 24, 25). No publication bias was detected by the inverted funnel plot or the Begg's and Egger's tests. From the pooled results, we could observe that the relative risk of severe neutropenia did not significantly differ in either AA/AC versus CC or AA versus AC/CC.
The random-effects model was applied to compare the AA versus the AC+CC genotypes: RR=0.79, 95% CI: 0.09-6.71 p=0.83, I2=89% (Fig. 3A). This result includes much heterogeneity since the result of Tibaldi and colleagues (17) were very different from those of other trials: the risk ratio of AA/AC+CC in the study by Tibaldi et al was 4.57, while it was 0.276 in the study of Xu et al (24), and 0.316 in the study of Joerger et al (19).
The fixed-effects model was applied to compare the AA/AC genotypes with the CC genotype: RR=2.19, 95% CI: 0.73-6.56, p=0.16; p for heterogeneity p=0.83, I2= 0% (Fig. 3B).
Severe thrombocytopenia
When considering the incidence rate of severe thrombocytopenia, we found 3 trials reporting this parameter, adding up to a total of 210 patients (17, 24, 25).
The AA genotype was compared to the AC/CC ones with a random-effects model, and the result was not significant (RR=1.93, 95% CI: 0.28-13.47, p for heterogeneity was respectively p=0.51 and p=0.12, I2=59%, using the random-effects model, Fig. 4A).
The AA/AC genotypes had a slightly higher incidence rate of thrombocytopenia than the CC one, but this difference was not significant (AA+AC vs CC: RR=1.576, 95% CI: 0.518-4.797, p for heterogeneity was respectively p=0.424 and p=0.88, I2=0%, Fig. 4B). The CI was large for both results, because of the small number of patients we could pool.
Severe anemia
For the analysis of severe anemia, we selected 2 trials including a total of 173 patients (17, 25). In both these trials the AA and AC genotypes were merged into one group to calculate the incidence rate of anemia, while the CC genotype was considered as the contrast group. The results showed that the AA/AC genotypes had a significantly lower incidence rate of severe anemia than the CC genotype: RR=0.308, 95%CI: 0.113-0.835, p=0.021 (Fig. 5).
The HWE and the Influence of ethnicity
We then calculated the allele frequency of the 3 genotypes (AA, AC, and CC) in the 6 studies that were in HWE (27); additionally, we could obtain from the other trial the small allele frequency, as shown in Table I. Different ethnicities had very different allele frequencies (28), but this did not influence the response or toxicity incidences for each genotype. Therefore, for any ethnicity the detection of polymorphisms to predict the response or incidence of severe hematologic toxicity was estimated to be practicable.
Discussion
With regard to the A79C polymorphism of the CDA gene, the evidence of its relationship with hematologic toxicity and clinical efficiency of gemcitabine had attracted many scholars' interest (29, 30); however, this issue has been controversial until today. With the present work, we aimed at summarizing the available evidence and to clarify the role of this polymorphism on gemcitabine hematologic toxicity and clinical efficiency. To our knowledge, this is the first meta-analysis considering the A79C CDA gene polymorphism focused on response and hematologic toxicity incidence in patients taking gemcitabine. By analyzing the data from 6 studies (enrolling a total of 623 cases) we interestingly found a significantly lower risk of high-grade anemia in CDA A79C wild type individuals (AA and AC) compared with CC individuals. In terms of risk of severe anemia, patients with the AA/AC genotypes were just three tenths of those bearing the CC genotype. Additionally, none of the results about response in NSCLC patients, severe neutropenia, and thrombocytopenia showed any statistical significance.
Many studies reached the conclusion that the A79C CDA wild type might have a lower activity (17, 25, 31, 32). This would lead to more severe toxicity (33) and better healing efficiency (34). Gilbert et al found that in mammary cells the Gln27 variant had 66% of the wild type activity (p<0.05), and the apparent Km value of the Gln27 allozyme was significantly higher than that for wild type CDA, p<0.025 (12). In our results, there was a trend towards a better response and an increased risk of severe thrombocytopenia in wild type individuals, but the pooled outcome showed no statistical difference. Most probably, the metabolite of gemcitabine is also affected by many other factors in the human body, and not only by CDA polymorphisms. Clearly, the clinical outcomes were more complicated than the laboratory results and could be affected by many factors. Okazaki at al (35) reported that the CDA genotype alone was not clinically relevant, and many other reports confirmed this conclusion (26, 36, 37, 38).
We could not establish how the CDA A79C gene affected red blood cells; additionally no literature or other information had reported on this question yet. However, we believe that the effect of gemcitabine may be, to a certain extent, superior in subjects with a wild type CDA genotype, so that the hematopoietic function of erythrocytes would recover better in the presence of the wild type CDA genotype. A similar mechanism appears to be true for sorafenib (39). Moreover, we must keep in mind that the analysis of severe anemia included only 2 trials and a total of 173 patients, as the available published literature was rather limited. Moreover, these 2 trials were performed by the same laboratory in different periods and their conclusions have not been proved by other studies.
In addition, a study reported that the MAFs of the A79C CDA were 0.153 in Koreans, 0.327 in Caucasian-Americans, 0.204 in Japanese, 0.155 in Chinese-Americans, and 0.087 in African-Americans (28). The trials included in our study were consistent with the abovementioned report and confirmed that the frequency of the mutant A79C CDA allele was lower in Asian patients than in Caucasians (Tab. I). Another Italian study reported a MAF of 43.7% (40); however, the Italian trials included in our analysis had a lower MAF than the value reported.
In addition, despite the fact that ethnicity was confirmed to affect the CDA alleles' frequency (28), when stratifying patients according to ethnicity, we also observed similar RRs of response rate in NSCLC patients, as well as similar RRs for hematologic toxicities, thus suggesting that the response rates and hematologic toxicities are independent of ethnicity or allele frequencies.
Despite our efforts in performing a comprehensive analysis, this study has some limitations. We obtained the data from publicly available clinical trials; thus we did not know the exact time at which adverse effects appeared in each trial or how long the responses lasted. Moreover, other limitations were: the significant heterogeneity in some analysis (Fig. 3A); the inclusion of different types of malignancies for the analysis of toxicities; the differences in the criteria of estimation of the hematologic toxicities between trials (WHO or CRETER3); the relatively low number of patients; the wideness of the CI of the combined RR; the fact that some of the trials were from the same laboratory; the different drug dosages that may affect the response and hematologic toxicities; and, finally, the fact that we could not perform the NSCLC patients' survival analysis, such as the evaluation of OS and PFS, because of insufficient data. Despite the small number of included trials, it is important to point out that the search criteria were broad and there were no other trials fitting the inclusion criteria.
On the “2011 ASCO Annual Meeting”, it was proposed to reconfirm plasma CDA as a positive predictive marker for gemcitabine efficacy, as well as to confirm the hypothesis that the A79C CDA polymorphism may be a potential genotype marker predicting gemcitabine response (41). However, from our meta-analysis we found that A79C CDA had no significant impact on the response of NSCLC patients, whereas the mutant type (CC) was related to a lower risk of severe anemia. Future prospective studies with large sample sizes and better study designs are required to confirm this issue.
