Abstract
The aim of this study was to evaluate tumor markers of molecular abnormalities that display tissue specificity, as to detect circulating tumor cells (CTCs) in breast cancer patients. Quantitative real-time RT-PCR was used to determine h-MAM, BCSG1, CK19, and c-erbB2 mRNA levels in peripheral blood (PB) of breast cancer patients. Results were compared with other epithelial cancers (lung or esophagus cancer), benign breast disease, and healthy individuals. We found that h-MAM mRNA was only detectable in the PB of patients with breast cancer (49 of 65, 75.4%), but not in patients with other epithelial cancers, benign breast disease, or healthy individuals. No significant differences in the expression level and positive detection rate of BCSG1, CK19, and c-erbB2 mRNA were observed between breast cancer and other epithelial cancers. Furthermore, the expression level and positive detection rate of h-MAM mRNA in PB were significantly correlated to the breast cancer pathologic stage (p=0.012 and p=0.015, respectively). Chemotherapy, radiotherapy, or total tumor resection (after 7 days of treatment) resulted in a significant decrease in the expression level of h-MAM mRNA in PB compared to the levels prior to the treatment (p<0.001). Importantly, an increase in h-MAM mRNA expression was detected in patients immediately after surgery, as well as 3 days post-surgery. These results indicate that the quantitative analysis of h-MAM mRNA is a useful tool for detecting CTCs in breast cancer patients, and can have a potential diagnostic utility in early micrometastasis, clinical evaluation of cancer treatment efficacy, and post-treatment monitoring of breast cancer patients.
Introduction
Breast cancer, as one of the leading causes of cancer-related death in women, has been considered a systemic disease that must be addressed with urgency because tumor cell dissemination occurs in the early stages of tumorigenesis and results in recurrent metastatic disease (1, 2). Breast cancer cells can bypass the lymph nodes and disseminate dirsectly through the blood to distant organs (3). Detection of occult epithelial tumor cells in peripheral blood, defined as circulating tumor cells (CTCs), may provide prognostic information beyond the TNM system. The detection of CTCs could have significant clinical utility in risk stratification in early breast cancer and early detection of relapse, as well as in the predictions of treatment efficacy, progression-free survival, and overall survival in primary and metastatic breast cancer patients (1, 4-5-6-7).
CTCs are rare cells present in the peripheral blood of cancer patients with the low rate of <10 cells/ml, about 1 cell per 106-107 leukocytes. Various techniques have been proposed to detect these micrometastatic cells: direct methods, mainly antibody-based assays (immunocytochemistry, immunofluorescence, flow cytometry) requiring an initial enrichment step, and indirect methods, mainly nucleic acid-based assays (mRNA transcripts by RT-PCR) (2). Quantitative real-time PCR, as a sensitive, quantitative, and contamination-free technique, can accurately estimate gene expression levels, and can differentiate between baseline gene expression in normal tissue and increased expression in cancer cells (8, 9). Molecular diagnosis using quantitative real-time PCR can detect cells expressing tumor markers and quantify tumor cells that are otherwise undetectable by other means in patients with localized or metastatic cancer. The latter may be the best method to detect micrometastases at the molecular level in various types of cancer patients (8, 10). Multimarker quantitative real-time PCR of cytokeratin 19 (CK-19), MAGE-A3, HER-2, TWIST1, hTERT α+β+, and mammaglobin gene transcripts in CTCs provides improved positive results in early breast cancer patients compared with CellSearch (11, 12). Compared with new cytometric techniques, such as positive immunomagnetic separation (IMS) with laser scanning cytometry (LSC), cell filtration, and LSC, the multimarker real-time RT-PCR assay of CK-19, mammaglobin, and prolactin-inducible peptide (PIP) was demonstrated to be the most sensitive and efficient technique for the detection of CTCs of breast cancer patients (4). As for breast cancer, tumor markers that have been investigated for their putative diagnostic and prognostic value include EpCAM, CKs (CK-7, -8, -18, -19 and -20), carcinoembryonic antigen (CEA), human mammaglobin (h-MAM), c-erbB2 (HER-2), EGF receptor (EGFR), vascular endothelial growth factor (VEGF), breast cancer-specific gene 1 (BSG1), MAGE-A3, TWIST1, hTERT α+β+, muscin-1 (MUC1), and PIP (13). Hence, the identification of the most epithelial-specific or organ-specific marker to detect CTCs by quantitative real-time PCR in breast cancer patients is of particular interest.
Mammaglobin, a mammary tissue-specific protein that is overexpressed in breast cancer tissues, has been investigated for its promising diagnostic utility in breast cancer for more than 15 years (13, 14). h-MAM has been shown to be expressed in a subset (70%-80%) of primary and metastatic breast cancer tissues, and was also detected in ovarian cancer patients (14, 15). Some controversial issues exist with respect to the clinical implication of h-MAM detection in peripheral blood. Marques et al suggested that h-MAM mRNA detection at diagnosis or during follow-up was not predictive of breast cancer recurrence (16); Benoy et al demonstrated that h-MAM expression in bone marrow had a superior prognostic significance compared with h-MAM expression in the CTCs of breast cancer patients (17). In this study, a standardized quantitative real-time RT-PCR assay was established to test and validate the prognostic value of h-MAM (compared with BCSG1, CK19, and c-erbB2) for detecting CTCs in breast cancer patients. We demonstrated that h-MAM mRNA was the most specific gene marker for detecting CTCs of patients with breast cancer. The expression level and positive detection rate of h-MAM mRNA in peripheral blood significantly correlated with the pathologic stage of the breast cancer. Furthermore, h-MAM mRNA expression levels were dynamically related to the efficacy of clinical treatments, and provided an immediate indication of the CTCs levels in the patients. Quantitative h-MAM mRNA analysis is a useful tool for detecting CTCs in breast cancer patients, and displays a potential diagnostic utility for the detection of early micrometastasis, the clinical evaluation of cancer treatment efficacy, and post-treatment monitoring in breast cancer patients.
Methods
Patients and clinical procedures
Patients diagnosed with pathologically proven breast cancer, as determined by routine imaging and cytological assessments, were eligible for the study. The clinical characteristics of the patients studied are shown in Table I. Patients with other epithelial cancers, including lung cancer and esophagus cancer, were included as controls. Among these subjects, 39 breast cancer patients, 10 lung cancer patients, and 3 esophagus cancer patients undergoing surgery for treatment were further studied. Additionally, 20 patients with breast cancer were investigated before and after treatment (Tab. II). Patients with a history of malignancy were excluded. Consent was obtained from each patient. The research was conducted in compliance with the principles enunciated in the Helsinki Declaration and approved by the ethics committee of the Anhui Provincial Hospital.
CLINICAL CHARACTERISTICS OF THE PATIENTS INCLUDED IN THIS STUDY
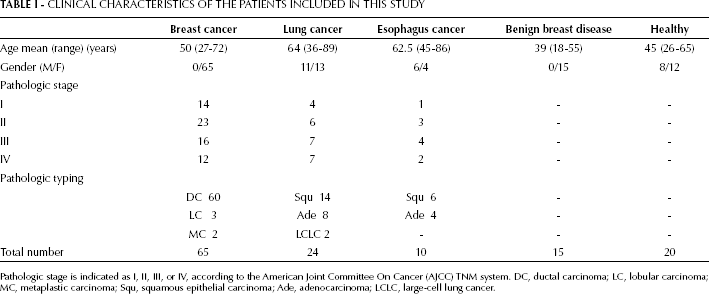
Pathologic stage is indicated as I, II, III, or IV, according to the American Joint Committee On Cancer (AJCC) TNM system. DC, ductal carcinoma; LC, lobular carcinoma; MC, metaplastic carcinoma; Squ, squamous epithelial carcinoma; Ade, adenocarcinoma; LCLC, large-cell lung cancer.
CLINICAL CHARACTERISTICS AND TREATMENT OF 20 PATIENTS WITH BREAST CANCER
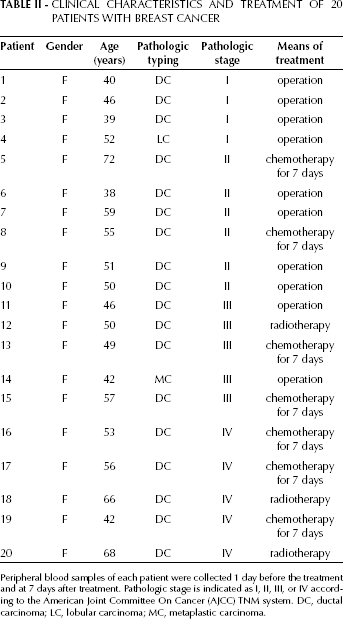
Peripheral blood samples of each patient were collected 1 day before the treatment and at 7 days after treatment. Pathologic stage is indicated as I, II, III, or IV according to the American Joint Committee On Cancer (AJCC) TNM system. DC, ductal carcinoma; LC, lobular carcinoma; MC, metaplastic carcinoma.
Collection of specimens
From each patient 3 mL of peripheral blood (the first 2 mL peripheral blood were discarded for the convenience of detection) were collected and treated with RBC lysis buffer (RX-2-1-2, U-gene, China). Nucleated cells were collected for the detection of mRNA biomarkers.
RNA extraction and cDNA synthesis
All reagents were purchased from Invitrogen. Total cellular RNA was extracted using the TRIzol reagent according to the manufacturer's protocol. mRNA was reverse transcribed into cDNA in 40 μL reaction volume, containing 4 μg of total mRNA, 5 μmol/L oligdT, 0.5 mmol/L dNTP, 8 μL 5×buffer, 10 mmol/L DTT, 56 Units RNase inhibitor, 400 Units M-MLV, and distilled water (ultrapure, DNase and RNase-free). The RT reaction was performed at 37°C for 50 minutes, followed by heating at 70°C for 15 minutes. All the steps were performed using sterile conditions in areas designated for RNA extraction and RT-PCR.
Real-time PCR
Quantitative real-time PCR was performed using real-time Taq-Man technology and a sequence detector (ABI PRISM 7000, Applied Biosystems, Foster City, CA, USA). Gene-specific primers and Taq-Man probes are listed in Table III. The standard 50 μL volume reaction contained 25 μL 2×PCR buffer (ABsolute™ QPCR Mix, AB-1140/b), 0.5 U uracil N-glycosylase (UNG) erase enzyme (Invitrogen, Cat NO.18054-015), 5 μL cDNA template, 0.25 μmol/L hybridization probe and 0.4 μmol/L forward and reverse primers. Initially, the reaction was incubated at 37°C for 10 minutes for UNG erase activation, followed by 95°C for 15 minutes. A total of 50 cycles consisting of 15 seconds melting at 95°C, and 1 minute annealing/extension at 60°C were performed. During the DNA polymerization, the Taq-Man probe was hydrolyzed and fluorescence was subsequently emitted. When the fluorescence signal reached 10 SDs of background, the threshold cycle (Ct) was noted. Each sample of unknown quantity was analyzed for the target gene in triplicate. Quantification of the detected mRNA was achieved using the standard curve method.
PRIMER PAIR AND PROBE FOR EACH BIOMARKER ASSESSED BY REAL-TIME PCR

The forward primer is shown as the upper sequence in the primer pair, while the reverse primer is shown as the lower sequence. Primers and probes for h-MAM (NM_002411.1), BCSG1 (NM_003087.2), and c-erbB2 (NM_004448.2) were designed using the Primer Express Software (Applied Biosystems). All primers were desalted when purified and the probes were HPLC-purified.
Preparation of the standard curve
h-MAM, CK19, BCSG1, C-erbB2, and β-actin cDNAs were generated from SK-BR-3, SK-BR-3, MCF-7, SK-BR-3, and A549 cells (American Type Culture Collection, Rockville, MD) using the respective primers (Tab. III). The PCR reaction consisted of an initial denaturation step at 94°C for 3 minutes, followed by 35 cycles of denaturation at 94°C, annealing at 60°C and extension at 72°C. After purification using a gel band purification kit (Invitrogen), the amplicon was ligated into the TA-cloning vector pCR 2.1 (Invitrogen, Groningen, The Netherlands) and transferred into DH5α-cells. The expected insert sequence was verified by sequencing using the ABI Prism BigDye terminator cycle sequencing ready reaction kit (PE Biosystems, Foster City, CA) on an ABI Prism 377 DNA sequencer. The number of plasmid copies was determined according to the formula
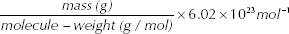
Serial dilutions ranging from 1×102 to 1×108 copies of plasmids per microliter were made in TE buffer (10 mM Tris, 0.1 M EDTA, pH 8.0) in tubes lubricated with silicon, and were detected by real-time PCR as described above. Standard curves were determined by plotting the Ct value against the initial copy number of standards (serially diluted plasmids in this case) for h-MAM, CK19, BCSG1, C-erbB2, and β-actin mRNA. The copy number of each gene marker of interest was calculated using standard curves by comparing the sample Ct value with the Ct values of serial dilutions of the standards. Furthermore, the h-MAM, CK19, BCSG1, and C-erbB2 copy numbers were all normalized as a ratio to the β-actin copy number in each sample.
Statistical Analysis
SDS 1.0 software was used to analyze the results of real-time PCR. Regression analysis was applied to the standard curves. The K independent samples test (Media Test) was used to compare the gene expression levels in peripheral blood among breast cancer patients of different pathologic stages. The K related samples and Wilcoxon signed ranks tests were used to analyze the gene expression levels in the peripheral blood of breast cancer patients before and after clinical treatment. When the gene expression results were negative, the data were treated as 0 for statistical convenience. The χ2 test was used to analyze the positive detection rate. A p<0.05 (2-tailed) was considered significant.
Results
Establishment of real-time RT-PCR procedures for the gene markers
The standard curves were established using specific plasmids for h-MAM, BCSG1, CK19, c-erbB2, and β-actin mRNA as described in the Methods' section. The amplification efficiency was 100% for h-MAM, 99.6% for BCSG1, 99.3% for CK19, 98.8% for c-erbB2, and 102% for β-actin. Using the established procedures, biomarkers were detected in each specimen and the products were assessed by agarose gel electrophoresis (Fig. 1). The expression patterns were consistent with the performance of each biomarker in these specimens, thereby further confirming the real-time RT-PCR procedures for each biomarker in this study.
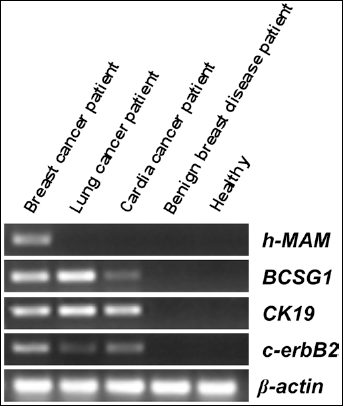
Real-time RT-PCR products for each gene marker. Each biomarker was detected by real-time RT-PCR in the indicated samples, as described in the materials and methods' section. Products were separated by electrophoresis and visualized in ethidium bromide-stained agarose gels.
h-MAM mRNA is the most specific gene marker for CTCs in breast cancer patients
As shown in Figure 2, h-MAM mRNA was exclusively detected in the peripheral blood of breast cancer patients (49/65), and not in the peripheral blood of patients with other epithelial cancers (0/34), benign breast disease (0/15), or in healthy volunteers (0/20). However, BCSG1, CK19, and c-erbB2 mRNA were detected in the peripheral blood of breast cancer patients (39/65, 48/65, and 19/65, respectively) as well as in patients with other epithelial cancers, such as lung cancer and esophagus cancer (22/34, 28/34, and 9/34, respectively). No significant difference in BCSG1, CK19, and c-erbB2 mRNA expression levels was observed between breast cancer patients and patients with other epithelial cancers (p=0.994, p=0.521, and p=0.656, respectively; Fig. 2).
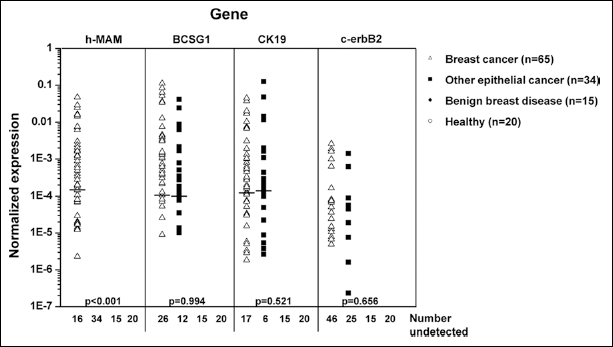
h-MAM mRNA is the most specific gene marker for CTCs in breast cancer patients. h-MAM, BCSG1, CK19, c-erbB2, and β-actin mRNA expression was determined by real-time RT-PCR in each peripheral blood sample; copy number was determined using a standard curve, as described in the methods section. The copy number of each gene was normalized to obtain the ratio to the copy number of β-actin in all samples. When the copy number of a gene marker was less than 100, it was considered as a non-detectable negative case. The numbers of negative results for each gene are displayed under “number undetected”. In each group the median value is marked using the indicator “-”. When the negative cases were >50%, the median could not be shown. The K independent samples test (Media Test) and Mann-Whitney U test were used to analyze gene expression levels among the 4 groups. The p values shown here indicate differences between the breast cancer group and other epithelial cancer groups. A p<0.05 (2-tailed) was considered significant.
Furthermore, the positive detection rate of h-MAM mRNA was 75.4% for breast cancer patients, and was similar to that of CK19 mRNA (73.8%) (p=0.840), and BCSG1 mRNA (60.0%) (p=0.061), but much higher than that of c-erbB2 mRNA (29.2%) (p<0.001) (Tab. IV). The positive detection rate for h-MAM mRNA in breast cancer patients was significantly different that in patients with other epithelial cancers (p<0.001); however, no significant difference was noted in the positive detection rates of BCSG1, CK19, and c-erbB2 mRNA between patients with breast cancer and other epithelial cancers (p=0.648, p=0.341, and p=0.772, respectively), which further indicated that BCSG1, CK19, and c-erbB2 mRNA were not specific to breast cancer cells.
NUMBER OF POSITIVE CASES FOR EACH BIOMARKER IN PERIPHERAL
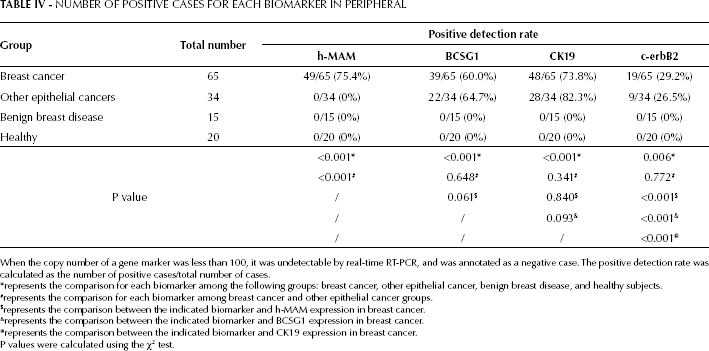
When the copy number of a gene marker was less than 100, it was undetectable by real-time RT-PCR, and was annotated as a negative case. The positive detection rate was calculated as the number of positive cases/total number of cases.
represents the comparison for each biomarker among the following groups: breast cancer, other epithelial cancer, benign breast disease, and healthy subjects.
represents the comparison for each biomarker among breast cancer and other epithelial cancer groups.
represents the comparison between the indicated biomarker and h-MAM expression in breast cancer.
represents the comparison between the indicated biomarker and BCSG1 expression in breast cancer.
represents the comparison between the indicated biomarker and CK19 expression in breast cancer.
P values were calculated using the χ2 test.
Expression of h-MAM mRNA in peripheral blood correlates with breast cancer pathologic stage
We further investigated the correlation between h-MAM mRNA expression in the peripheral blood of breast cancer patients and a number of clinical factors (Tab. V). Age and pathologic typing were not associated with the positive detection rate of h-MAM mRNA in peripheral blood (p=0.424 and p=0.413, respectively), whereas a significant difference was observed among different pathologic stages. The positive detection rate of h-MAM mRNA increased with higher pathologic stages: 50.0% for stage I, 69.9% for stage II, 87.5% for stage III, and 100% for stage IV (p=0.015). Furthermore, h-MAM mRNA expression levels were positively associated with the progression in pathologic stages (p=0.012). The most clinically severe breast cancer patients displayed the highest h-MAM mRNA expression in their peripheral blood. Particularly, a significant difference was observed in h-MAM mRNA expression levels between stage II and III tumors (p=0.011) (Fig. 3). Similar results were also observed for CK19 mRNA (p=0.013, Tab. V; p=0.012, Fig. 3). Overall, the positive detection rates of BCSG1 mRNA were also significantly correlated to the pathologic stages of breast cancer (p=0.002) (Tab. V), despite being low in patients with stage I and II breast cancer. Besides, no significant difference was observed in the expression levels of BCSG1 between stage II and III cancer patients (p=0.377) (Tab. V, Fig. 3). These results indicate that BCSG1 mRNA is not sensitive enough to quantify CTCs in breast cancer patients. The positive detection rate and expression of c-erbB2 mRNA in peripheral blood did not correlate to the pathologic stages of breast cancer (p=0.482, Tab. V; p=0.482, Fig. 3).
CORRELATION BETWEEN GENE MARKERS IN PERIPHERAL BLOOD AND CLINICAL FACTORS IN BREAST CANCER PATIENTS
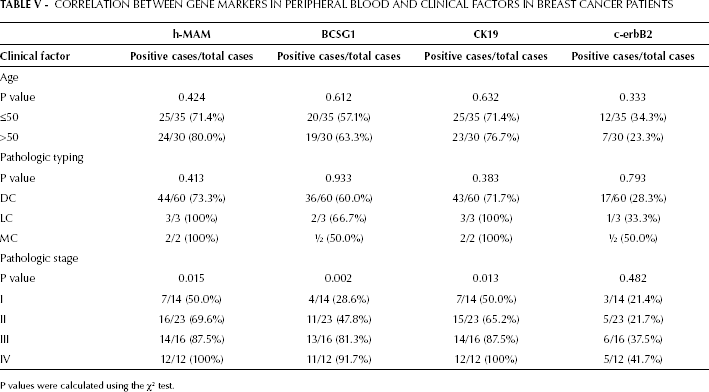
P values were calculated using the χ2 test.
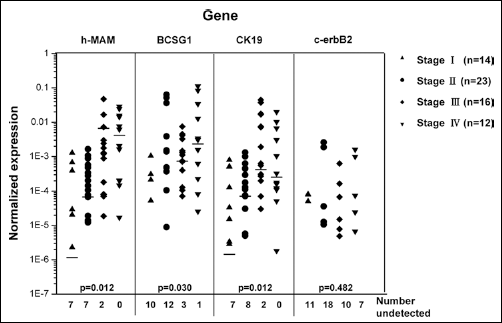
h-MAM mRNA expression in the peripheral blood correlated with the pathologic stage of breast cancer. h-MAM, BCSG1, CK19, c-erbB2, and β-actin mRNA expression was detected by real-time RT-PCR in each peripheral blood sample, and the copy number was determined using a standard curve. The copy number of each gene was further normalized to obtain the ratio to the copy number of β-actin. When the copy number of a gene marker was less than 100, it was considered as a non-detectable negative case. The numbers of negative results for each indicated gene are displayed under “number undetected”. In each group the median value is marked using the indicator “-”. When the negative cases were >50%, the median could not be shown. The K independent samples test (Media Test) was used to analyze gene expression levels among breast cancer patients from different pathologic stages. A p<0.05 (2-tailed) was considered significant.
Detection of h-MAM mRNA dynamically quantified CTCs in breast cancer patients during therapy
Expression of h-MAM mRNA in the peripheral blood of 20 breast cancer patients was tracked before and after treatments and compared with the BCSG1, CK19, and c-erbB2 mRNA levels (Tab. II). As shown in Fig. 4, the mRNA expression of h-MAM, BCSG1, CK19, and c-erbB2 in peripheral blood was significantly decreased after treatment (p<0.001, p<0.001, p<0.001, and p=0.018, respectively). As shown in Table IV, the positive detection rate of c-erbB2 mRNA was much lower than that of the other 3 biomarkers (Fig. 4). Furthermore, h-MAM mRNA expression in peripheral blood significantly increased immediately after surgery (p<0.001, Fig. 5), indicating that surgery caused h-MAM mRNA-expressing cells to enter the peripheral blood. On the third postoperative day, the h-MAM mRNA expression was still increased compared with pre-surgery levels (p<0.001, Fig. 5). As shown in Tab. VI, breast cancer cells entered the peripheral blood immediately after surgery in 9/39 (23.1%) patients who had no detectable CTCs before surgery. Among these patients, the CTCs remained detectable in 5/39 (12.8%) patients on the third postoperative day, while for 4/39 (10.3%) of them CTCs were only detectable at day 1 after surgery. Given the specificity of h-MAM mRNA for breast cancer cells, and its correlation with the pathologic stages of this disease, h-MAM mRNA seems to be a useful gene marker for detecting CTCs for the evaluation of cancer treatment efficacy and monitoring of breast cancer patients post-treatment.
h-MAM mRNA EXPRESSION IN THE PERIPHERAL BLOOD OF BREAST CANCER PATIENTS UNDERGOING SURGERY
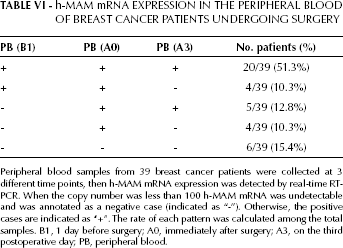
Peripheral blood samples from 39 breast cancer patients were collected at 3 different time points, then h-MAM mRNA expression was detected by real-time RT-PCR. When the copy number was less than 100 h-MAM mRNA was undetectable and was annotated as a negative case (indicated as “-”). Otherwise, the positive cases are indicated as “+”. The rate of each pattern was calculated among the total samples. B1, 1 day before surgery; A0, immediately after surgery; A3, on the third postoperative day; PB, peripheral blood.
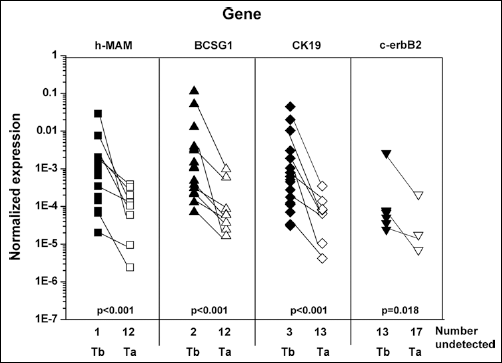
h-MAM mRNA expression in the peripheral blood decreased shortly after breast cancer treatment. Peripheral blood samples from 20 breast cancer patients were collected 1 day before the administration of treatment and 7 days after the treatment, as shown in Table II. The expression of h-MAM, BCSG1, CK19, c-erbB2, and β-actin mRNA was detected by real-time RT-PCR, and the copy number was determined using a standard curve. “Tb” indicates the time point 1 day before treatment, and “Ta” indicates 7 days after treatment. The copy number of each gene was further normalized to obtain the ratio to the copy number of β-actin. When the copy number of a gene marker was less than 100, it was considered as a non-detectable negative case. The negative results for each indicated gene were displayed as the number undetected. The Wilcoxon signed ranks test was used to analyze the gene expression levels before and after clinical treatment. A p<0.05 (2-tailed) was considered significant.
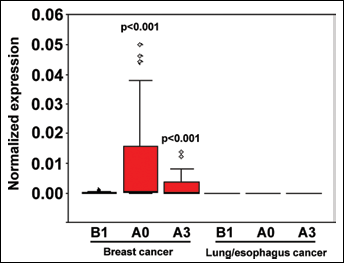
Breast cancer surgery dynamically altered the number of h-MAM mRNA-positive cells in the peripheral blood. Peripheral blood samples from 39 breast cancer patients were collected before surgery (B1), immediately after surgery (A0), and on the third postoperative day (A3). The expression of h-MAM mRNA was detected by real-time RT-PCR and the copy number was determined using a standard curve and further normalized to obtain the ratio to the copy number of β-actin. When the copy number of a gene marker was less than 100, it was considered as a non-detectable negative case. Samples from 10 lung cancer patients and 3 esophagus cancer patients were collected as controls. K related samples and Wilcoxon signed ranks test were used for the comparison of gene expression levels among the B1, A0, and A3 groups. A p<0.05 (2-tailed) was considered significant.
Discussion
In this study we established a standardized quantitative real-time RT-PCR assay for detecting CTCs in breast cancer patients. The data obtained in breast cancer patients were compared with those of patients with other epithelial cancers and benign breast disease, as well as healthy volunteers. In comparison to the 3 other breast tumor markers BCSG1, CK19, and c-erbB2 mRNA, h-MAM mRNA was the most specific gene marker for detecting CTCs in breast cancer patients. The expression levels and positive detection rate of h-MAM mRNA in the peripheral blood significantly correlated with the pathologic stages of breast cancer. CTC levels could be tracked immediately and dynamically in patients undergoing treatments, by detecting h-MAM mRNA expression in the peripheral blood. These results indicate that the quantitative analysis of h-MAM mRNA is a useful tool for detecting CTCs in breast cancer patients, and displays potential diagnostic utility not only in the detection of early micrometastasis, but also in the clinical evaluation of cancer treatment efficacy and in the post-treatment monitoring in breast cancer patients.
Cancer cells can be released from a primary site and can spread via the bloodstream to form a micrometastatic deposit in distant organs. Thus, the issue of cancer circulation is a hot topic in cancer research (18). In particular, breast cancer cells disseminate directly through the blood to distant organs during the early stage of tumorigenesis, which makes breast cancer a systemic disease with a high mortality in women, even at the earliest stage (pathologically confirmed stage I) (1-2-3). The number of CTCs prior to treatment is an independent predictor of progression-free survival and overall survival in patients with metastatic breast cancer (5). CTCs are detectable in 10%-30% of breast cancer patients with stage I to III disease, and in 50%-70% of women with metastatic breast cancer (19). It was speculated that undetected metastases were present at the time of surgery, given that conventional staging techniques lack the necessary sensitivity to properly characterize patients. Thus, the establishment of a sensitive and specific method for CTC detection has important diagnostic, prognostic, and therapeutic implications. Quantitative real-time PCR is applicable because cancer cells continue to express genetic markers specific to their tissue of origin. The following tumor markers have been investigated by quantitative real-time PCR in breast cancer: EpCAM, CKs, CEA, h-MAM, c-erbB2 (HER-2), EGFR, VEGF, BSG1, MAGE-A3, TWIST1, hTERT α+β+, Survivin, MUC1, and PIP (6, 20-21-22-23-24). In this study, h-MAM, BCSG1, CK19, and c-erbB2 were assessed to determine the most organ-specific marker for detecting CTCs in breast cancer patients by quantitative real time PCR.
As for breast cancer, the gene marker CK19 mRNA was firstly assessed to evaluate its clinical potential for the molecular detection of CTCs by quantitative RT-PCR, and further studies demonstrated its specificity and sensitivity for breast cancer cells (25-26-27-28-29-30). CK19-mRNA-positive cells were detected in 31.2% of patients with early breast cancer (stage I-II), and in 40.4% of patients with verified metastases (stage IV) (29). In our study, the established quantitative real-time PCR for CK19 mRNA was more sensitive, with a positive detection rate of 59.5% (22/37) in early breast cancer patients (stage I-II) and of 100% (12/12) in metastatic patients (stage IV) (Tab. V). Compared with CEA and maspin, CK19 mRNA is the most sensitive detection marker for occult tumor cells in operable and metastatic breast cancer (31). h-MAM is the most widely studied marker in breast cancer patients after CK19, which is a demonstrated specific marker for the identification of occult cancer cells in the peripheral blood of breast cancer patients with a positive detection rate of approximately 10%-30% (32, 33). In our study, the established quantitative real-time PCR for h-MAM mRNA was more sensitive, with a positive detection rate of 75.4% (49/65 patients) (Tab. IV). By comparison, no significant difference in the positive detection rate between h-MAM and CK19 mRNA was observed (75.4% vs 73.8%; Tab. IV). However, CK19 is overexpressed in many epithelial cancers. As shown in Fig. 2, CK19 mRNA-positive tumor cells were detectable not only in the peripheral blood of patients with breast cancer, but also in those with lung cancer and esophagus cancer, thus limiting the marker's specificity for detecting breast cancer cells in the peripheral blood.
In addition, the breast cancer-specific gene BCSG1 is undetectable in normal or benign breast lesions. BCSG1 showed partial expression in ductal carcinoma in situ, and was expressed at an extremely high level in advanced infiltrating breast cancer (34). BCSG1 has significant sequence homology with the Alzheimer's disease-related neural protein synuclein (SNC); thus, BCSG1 was also named SNC-gamma (SNCG) (35). SNCG regulates many pathways involved in breast cancer growth and progression. Furthermore, BCSG1 is a novel biomarker for clinical prognosis of breast cancer (36, 37). BCSG1 expression strongly correlates with stage, lymph node involvement, metastasis, tumor size, and Her-2 status. For example, 71.4% or 81% of stage III/IV breast cancers were positive for BCSG1 expression, while only 26.8% or 15% of stage I/II breast cancers were positive for BCSG1 expression. Patients with a BCSG1-positive tumor had a significantly shorter disease-free survival and a high probability of death compared with patients with BCSG1-negative tumors (36, 37). In these studies the tumor tissue samples were analyzed, but data regarding the utility of BCSG1 expression in the detection of CTCs in breast cancer patients were not reported. In our study, we demonstrated that 60% (39/65) of breast cancer patients were positive for BCSG1 expression in peripheral blood; of these, 85.7% (24/28) of stage III/IV breast cancers were BCSG1-positive, while 40.5% (15/37) of stage I/II breast cancers were BCSG1-positive (Tabs. IV and V). No significant difference was observed between BCSG1 and h-MAM for the detection of CTCs in breast cancer patients (Tabs. IV and V). However, BCSG1 mRNA expression was not specific to breast cancer cells, as it was found to be a novel prognostic biomarker in uterine papillary serous carcinoma (38). In our study, we also observed BCSG1 expression with a high positive detection rate in the peripheral blood of lung cancer, cardia cancer, and esophagus cancer patients (Figs. 1 and 2). No significant difference in BCSG1 expression was observed between breast cancer and other epithelial cancers (Fig. 2). These results indicate that BCSG1 mRNA expression is not specific to breast cancer cells. Furthermore, compared to h-MAM mRNA, the identification of BCSG1 mRNA is not sensitive enough to quantitatively detect CTCs in breast cancer patients, although the positive detection rates of BCSG1 mRNA also significantly correlated with the pathologic stages of breast cancer (Tab. V). We also compared the prognostic value of c-erbB2 (HER2) for the detection of CTCs in breast cancer with 3 additional biomarkers. c-erbB2 mRNA expression was not specific to breast cancer cells, and its positive detection rate and expression level in the peripheral blood did not correlate with the pathologic stages of breast cancer (Tab. V and Fig. 3). Furthermore, in breast cancer patients, the positive detection rate of c-erbB2 mRNA was significantly lower than that of h-MAM mRNA, CK19 mRNA, and BCSG1 mRNA (Tab. IV). These results demonstrate that h-MAM mRNA is the most specific and sensitive gene marker for detecting CTCs in breast cancer patients.
A good marker for breast cancer would promptly establish the patient's status both before and after relevant clinical treatments. In this study we further evaluated the utility of h-MAM for detecting CTCs in the therapeutic assessment of breast cancer patients. Chemotherapy, radiotherapy (more than 7 days of treatment), or total tumor resection significantly decreased h-MAM mRNA expression in the peripheral blood compared with samples obtained prior to treatment (p<0.001) (Fig. 4). Importantly, an increase in h-MAM mRNA expression was detected in patients immediately after surgery (Fig. 5), indicating that surgery caused breast cancer cells to enter the peripheral blood. In addition, h-MAM mRNA expression remained high on the third postoperative day compared with the levels prior to surgery (Fig. 5). This study firstly demonstrates that h-MAM mRNA, detected by quantitative real time PCR, is a specific and sensitive biomarker that can dynamically indicate the status of breast cancer patients via detection of CTCs during clinical treatment. Esophageal cancer surgery results in tumor cell dissemination and a significant increase of CTCs in the peripheral blood (39). Detection of CEA mRNA-positive cells in peripheral blood led to positive detection rates of 28.3%, 60.4%, and 42.9%, respectively obtained before surgery, immediately after surgery, and on the third postoperative day (39). By contrast, in this study the detection of h-MAM positive cells in peripheral blood led to positive detection rates of 61.5%, 84.6%, and 64.1%, respectively obtained before surgery, immediately after surgery and on the third postoperative day (Tab. VI). Breast cancer cells entered the peripheral blood immediately after surgery in 23.1% of patients (9/39) who had no detectable CTCs before surgery. Among these patients 12.8% (5/39) still had CTCs on the third postoperative day. These results might help determining the causes of the high recurrence and poor survival observed in breast cancer patients despite undergoing surgery, and should be further evaluated in follow-up studies.
Conclusions
By using a standardized quantitative real-time RT-PCR assay this study demonstrates that h-MAM mRNA is the most specific and sensitive gene marker for detecting CTCs in early stage breast cancer patients compared with BCSG1, CK19, and c-erbB2. Furthermore, the actual CTC levels could be tracked dynamically via assessment of h-MAM mRNA expression in the peripheral blood of patients undergoing clinical treatment. CTC detection by quantitative assessment of h-MAM mRNA levels has potential diagnostic utility in the diagnosis of early micrometastasis and in the therapeutic assessment of breast cancer patients.
