Abstract
Introduction
Prostate cancer (PCa) can be a very aggressive tumor, although often it is slow-growing (1). Many factors, including diet, habits, and genetics have been implicated in the development of PCa (2). The genetic background may contribute to the prostate cancer risk, as suggested by its association with ethnicity, family, and specific gene variants (3, 4). Men who have a first-degree relative (father or brother) with PCa have twice the risk of developing PCa, and those with two first-degree relatives affected have a 5-fold greater risk compared with men with no family history of PCa (5, 6). PCa cells may metastasize to other parts of the body, particularly lymph nodes and bones. Despite the fact that developments in early detection and treatment in these last years have reduced the mortality rates, it is estimated that approximately 27,000 patients die of PCa each year (7).
The cerebellar degeneration-related autoantigen 1 (CDR1) gene is located in the chromosomal region Xq26-q27.2 (OMIM302650), and the protein has been identified in some patients with paraneoplastic cerebellar degeneration (OMIM302650). The CDR1 gene encodes for a nuclear protein of 223 amino acids with a molecular mass of approximately 27 kDa. Over 90% of CDR1 is made up of 34 inexact tandem repeats of 6 amino acids that feature a nearly invariant core of glutamine and aspartic acid residues. Possible splice variants encoding isoforms that differ at the C-terminal end, outside the hexapeptide repeat region, have also been identified (8). RNA dot blot showed a high CDR1 expression in cerebrum and cerebellum, a low expression in lung, heart, and kidney, and none in liver. Among cancer cells lines, CDR1 expression has also been detected in neuroblastoma cell lines, as well as in the majority of renal cell carcinoma cell lines and in some astrocytoma, melanoma, and lung carcinoma cell lines (8).
Methods and Results
The purpose of this study was to investigate the expression of the CDR1 gene in the LNCaP and PC-3 PCa cell lines and in the PNT1A normal prostate cell line. The expression of CDR1 mRNA was evaluated by qRT-PCR.
The normal human immortalized prostatic cell line PNT1A provides a useful tool to study the biology and pathology of adult prostatic epithelial cells, especially for understanding the steps leading to prostate cancer transformation (9, 10). PNT1A can be considered non-tumorigenic cells showing molecular and biochemical properties close to the normal prostate epithelium. PNT1A cells express high-affinity androgen receptors. The LNCaP is a cell line that was derived in 1977 from human prostate adenocarcinoma cells of a 50-year old Caucasian male; cells were taken from a needle aspiration biopsy of a metastatic lesion in the left supraclavicular lymph node (11). The PC-3 cell line originally derived from advanced PC androgen-independent bone metastasis, and has an elevated metastatic potential (12).
The PNT1A, LNCaP, and PC-3 cell lines were purchased from the American Type Culture Collection (Rockville, MD, USA) and were cultured in RPMI 1640 supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin. All of the cells were maintained at standard cell culture conditions (37°C, 5% CO2 in a humidified incubator).
RNA extraction from all cell lines was performed using the RNeasy Mini Handbook (QIAGEN Sciences, Germantown, PA, USA), following the manufacturer's protocol. RNA quantity and quality were checked by spectrophotometry.
To avoid any genomic DNA contamination during qRT-PCR, a brief incubation of the samples at 42°C with a specific Wipeout buffer (QuantiTect Reverse Transcription Kit, QIAGEN Sciences, Germantown, PA, USA) was carried out. Retro-transcription of 600 ng of total RNA from each sample was then performed in a final volume of 50 μL (QuantiTect Reverse Transcription Kit, QIAGEN Sciences, Germantown, PA, USA) and the generated cDNA was used as a template for qRT-PCR analysis using gene expression products.
For each sample, the RT-PCR reactions were carried out in duplicates using 2.5 μL of cDNA and QuantiTect Probe PCR Master Mix Kit (QIAGEN Sciences, Germantown, PA) in a total volume of 50 μL. The assays for the target gene CDR1 and the reference gene GAPDH were obtained from Applied Biosystems (Carlsbad, CA, USA). The thermal cycling conditions consisted of 1 cycle for 2 minutes at 50°C, 1 cycle of 15 minutes at 95°C and 40 cycles for 15 seconds at 94°C followed by 1 minute at 60°C. Real time analysis was performed on an ABI PRISM 5700 Sequencer Detector (Applied Biosystems). The amplified transcripts were quantified using the comparative ΔΔCt method (13), and relative quantification analysis data were evaluated using the comparative ΔΔCt method included in the Software Version 1.3 supplied with the ABI PRISM 5700 Sequencer Detector (Applied Biosystems), which is based on the ΔΔCt algorithm. The result of higher or lower expression has been given in terms of numbers obtained by the normalized ratio (RT). The CDR1 gene expression level was normalized to GAPDH expression and the Target Mean Cp definition was used to indicate the mean normalized cycle threshold.
We found that CDR1 was overexpressed in the LNCaP and PC-3 PCa cell lines compared with the PNT1A normal prostate cell line (Tab. I).
CDR1 mRNA EXPRESSION IN PC-3, LNCaP, AND PTN1A PROSTATE CELL LINES
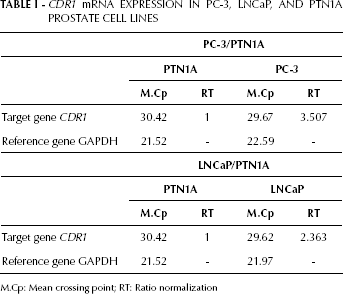
M.Cp: Mean crossing point; RT: Ratio normalization
Discussion
The results of this study suggest that the CDR1 gene may play a role in the oncogenetic process of PCa. In addition, the finding that CDR1 is overexpressed in PCa cell lines confirms previous studies showing the overexpression of this gene in other types of tumors. In addition, the CDR1 gene is a highly conserved neuroectodermal marker mapping to chromosome X in both human and mouse (8, 14, 15). Using genome-wide scans in PCa families, 40 candidate regions have been identified for PCa susceptibility loci on different chromosomes. This indicates the great heterogeneity of this disease (16, 17). One of the first loci found by linkage analysis of 360 hereditary families with PCa is HPCX1, on chromosome Xq27-q28, estimated to be responsible for 16% of the hereditary cases of PCa (18).
The finding that the CDR1 gene has a higher expression in the LNCaP and PC-3 PCa cell lines than in the PNT1A normal prostate cell line lead us to hypothesize that the CDR1 gene could be one candidate gene for the HPCX1 locus. This hypothesis does not exclude other candidate genes for the HPCX1 locus. In fact, SPANX, another important gene family, maps within the HPCX1 locus. Furthermore, CDR1 may be regarded as a new biomarker for PCa. This finding needs to be confirmed in a greater number of patients.
