Abstract
A key event in the development of cervical carcinoma is the deregulated expression of high-risk human papillomavirus (HR-HPV) oncogenes, most commonly due to HPV integration into host DNA. Here we explored whether HPV-16 E2 gene integrity is a biomarker of progressive disease with oncogenes expression. HPV-16 genome disruption was assessed by amplification of the entire E2 gene, while mRNA expression patterns of the E1, E2, E6, and E7 genes were evaluated by reverse transcription PCR (RT-PCR). As expected, E2 disruption was significantly higher among patients with cervical cancers than subjects with benign lesions (p=0.02). The status of the E2 gene correlated with tumorogenesis, and seemed also to correlate with the stage of the carcinomas, since integrated HPV-16 DNA was frequently detected in patients with advanced cancer stages (75% of stage III vs 60% stages I and II). In bivariate analysis, the lesions’ grade was most significantly associated with HPV-16 DNA disruption (p<0.05). In cervical carcinoma the deletion pattern involved more frequently the E2 gene rather than the E1 gene (62.5% vs 45.8%). The prevalence of the E6/E7 HPV-16 transcripts in cervical carcinoma specimens and in benign cervical lesions were detected with frequencies of, respectively, 91.6% and 45.4%. The mRNA levels of the HPV-16 E6/E7 genes were expressed at approximately the same levels in each physical state. We consistently observed that E6/E7 were absent or weakly detectable in the presence of E2. However, in the absence of E2 the levels of E6/E7 markedly increased (p<0.05). This study underscores the significance of investigating alternative mechanisms of E2 expression and oncogenes E6/E7 transcripts in vivo as biomarkers for disease severity in cervical carcinomas.
Introduction
Infection by high-risk human papillomaviruses (HR-HPV) is the single most important risk factor of cervical cancer (1–3). Although most women will, at some point, get infected by HR-HPV, only for very few of them the infection will progress to an invasive disease, thus indicating the involvement of other additional factors (4, 5). Since the integration of the viral genome mainly occurs at fragile sites (6), it not only affects various cellular pathways of the host cell-cycle machinery, but it also disrupts the viral E1 and E2 genes (7, 8). Some investigators reported the early integration of HPV-DNA in low-grade cervical lesions and in asymptomatic infections (9–12), whereas others observed HPV integration almost exclusively in high-grade lesions and invasive carcinomas (13, 14). In the absence of E2-driven repression, E6 and E7 are overexpressed, thus driving infected cells toward transformation (15). Moreover, integrant-derived transcripts are more stable than those originating from episomal viral DNA (16). Taken together, the malignant transformation depends on the continuous expression of HPV E6 and E7 oncogenes, which bind to and modulate a number of different gene products, particularly the tumor suppressors p53 and pRb. These interactions lead to a disturbance of the mechanisms of cell-cycle control and a deficiency in DNA repair, thus resulting in genomic instability and an increased risk of malignant transformation (17). Transcription of both viral oncogenes, E6 and E7, is under control of a common promoter (p97), which is mainly repressed by the viral E2 gene product (18). The disruption of the E2 gene abrogates this negative regulatory circuit and is considered a selective event in tumor development and progression (19, 20). E2 function can also potentially be altered by gene mutation/variation (21). The aim of this study is to analyze the integrity of the HPV-16 E2/E1 genes in all grades of cervical neoplasia and to study specific genomic expression profiles of different HPV-16 DNA physical states.
Material and Methods
Clinical Samples
A series of 24 HPV-16-positive cervical carcinoma specimens and 11 HPV-16-positive benign cervical lesions were evaluated. Cervical carcinoma patients were recruited from the Radio-oncology department of the Salah Azaiez Institute (Tunis, Tunisia). Clinically, invasive disease was staged according to the criteria of the International Federation of Gynecology and Obstetrics (FIGO) and all patients had stage I to III of the disease. One patient had stage I disease, 19 had stage II and 4 stage III disease. The subjects’ age ranged between 38 and 76 years (mean, 57.2 years). Histological classification was performed according to the World Health Organization (WHO) criteria, and HPV-16 DNA was detected in 23 patients with squamous cell cancer and in 1 patient with adenocarcinoma. Benign cervical biopsies were obtained from women with cervical inflammation attending the clinic for routine reproductive healthcare counseling, and were subsequently examined at the Obstetrics and Gynecology Department of the La Rabta Hospital (Tunis, Tunisia). Cervical inflammation was determined by cytology. The control group consisted of 29 women with a normal Pap smear and normal colposcopy, as well as a negative HPV-DNA test. For this group of women the mean age was 41 years (range, 27–56 years). Clinicopathological variables, including histological diagnosis and stage of disease, were obtained from the medical records. Biopsy specimens were resuspended in 1 mL of phosphate-buffered saline (PBS, pH 7.4) and then stored at -20°C until processed.
Nucleic Acid Preparation
DNA and RNA extractions were conducted using the QIAamp DNA Mini Kit and the RNeasy Mini Kit (both from Qiagen, Hilden, Germany) following the manufacturer's instructions.
Quality and quantity of the target nucleic acids were respectively assessed by agarose gel electrophoresis and spectrophotometry. Suitability for PCR analysis was assessed by performing PCR with the β-globin housekeeping gene, which resulted in amplicons of 268 bp (22).
Evaluation of HPV-16 Physical State
All HPV-16 E1-positive samples were subjected to PCR amplification of the E2 ORF region spanning the full length of the HPV-16 E2 gene (23). Briefly, 200 ng of purified DNA were used in 20 μL PCR reaction mixture containing 100 ng of primers, 50 mM KCl, 1.5 mM Mg Cl2, 50 mM Tris HCl (pH 8.3), 50 mM (NH4)2SO4, 25 μM of each dNTP, and 0.5 U of thermostable DNA polymerase (Fermentas). We defined an HPV as integrated if the E2 product was not amplified. When the results of E2 primers were positive, the E2 gene was considered intact.
Reverse Transcription (RT)-PCR for Detection of HPV-16 E1, E2, E6, and E7 Transcripts
HPV-16-positive samples were further subjected to amplification of E1 (24), E2 (25), E6, and E7 transcripts (26). RT-PCR was performed using primers for the constitutively expressed β-actin (26) as a loading control and sample normalization. Details of the target genes, primer sequences and expected sizes of the amplicons are shown in Table I.
Pcr and rt Primers, Product Length, and PCR Conditions
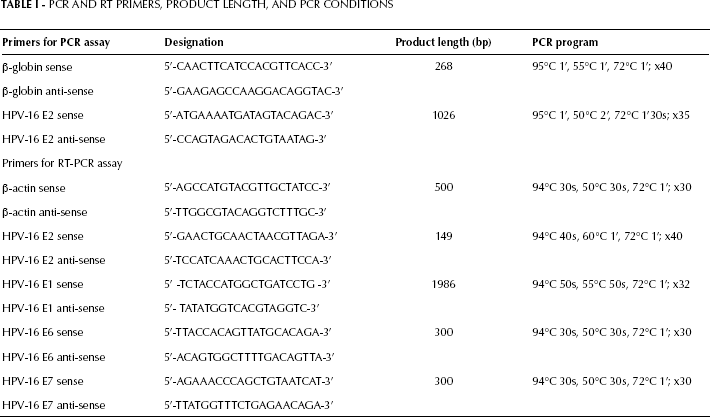
RT-PCR was performed in a 50-μL total volume, using 1 μg of template RNA and the one-step RT-PCR Kit (Qiagen). Reactions contained 0.6 μM forward and reverse primers, 1X Qiagen one-step RT-PCR buffer, 400 μM dNTP mix and 2 μL Qiagen one-step RT-PCR enzyme. The reaction was carried on for 30 minutes at 50°C. HPV-16 type-specific plasmids were used as positive controls. For each of the target genes, control reactions without templates were performed to make sure that no contamination had occurred. Cycling protocols for all RT-PCR reactions are shown in Table I. The amplicons were resolved by 1.5% gel electrophoresis, marked by 50-bp and 100-pb DNA ladders (Gene Ruler™, Fermentas), stained with ethiduim bromide, and visualized under UV light.
Semiquantitative RT-PCR Analysis
An equal amount of cDNA (100 ng) from each sample was analyzed by semiquantitative RT-PCR to measure HPV-16 E2/E6/E7 oncogenes expression (cDNA levels reflect mRNA levels).
The values corresponding to the intensity and density of the gel bands of different transcripts were obtained with the help of Doc-Print II software and Photo Capt software (VILBER LOURMAT, Marne-la-Vallée, France). Semiquantitative estimates of mRNA expression levels of the target genes were determined relatively to the β-actin expression in the same sample. Using the same amount of total cDNA (in ng) from each sample allowed accurate comparison of the target genes. Significant differences between the expression levels of the E2 and the E6/E7 genes were tested using the Student's t-test.
Statistics
The relationships between the different variables were assessed using the Fisher's exact test. Odds ratios (OR) and corresponding 95% confidence intervals (CI) were estimated by logistic regression models. Data were statistically analyzed using the Student's t-test. Values of p<0.05 were considered to indicate statistical significance. The analyses were carried out using SPSS, version 18.0.
Results
Comparison of the Physical State of HPV-16 in Cervical Lesions of Different Grades and Clinicopathological Data
Overall, 35 HPV-16-positive DNA samples from patients with different grades of cervical neoplasia were analyzed by PCR. The integration rate in case of HPV-16 infection was higher in cervical cancer samples (62.5%) than in benign lesions (18.1%). Overall, episomal HPV-16 DNA was found in 9/24 (37.5%) cases with cervical carcinoma and in 9/11 (81.8%) women with benign lesions (p=0.02, Tab. II).
Bivariate Correlations with E2 Gene Status of Hpv-16 Positive Tumors
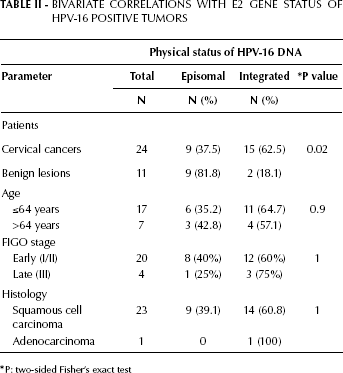
P: two-sided Fisher's exact test
There was no significant difference in patients’ age between different physical statuses of HPV-16 DNA (Tab. II). The integrated forms of HPV-16 DNA were not detected in the only patient with stage I disease, but were detected in 7/12 (58.3%) patients with stage IIp, in 5/7 (71.4%) with stage IId, and 3/4 (75%) with stage III disease (Fig. 1).
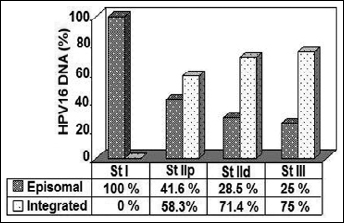
Distribution of HPV-16–DNA physical state in different tumor stages. Abbreviation: St, stage; p, proximal; d, distal.
Despite the fact that there was no statistically significant association between viral genome integration and clinical stage of the tumor, the integrated forms could be widely observed in patients with advanced stages of cervical cancer (Tab. II, Fig. 1). Integrated forms were detectable in 14 (60.8%) squamous cell carcinomas, whereas only 1 adenocarcinoma carried the integrated form of HPV-16 DNA (Tab. II). The relationship between the histologic type and detectability of episomal HPV-16 DNA was not statistically significant.
Assessment of E2/E1 HPV-16 mRNA Expression
To evaluate the quality of the RNA extracted from the specimens for further amplification by RT-PCR, all samples were tested for expression of the β-actin housekeeping gene using RT-PCR. β-actin was detected in all cervical specimens and negative controls were consistently negative for β-actin. The presence of intact HPV-16 E2 DNA, indicative of episomal forms, was confirmed by HPV-16 E2 RNA analysis (Fig. 2), as HPV-16 E2 mRNAs were consistently expressed in the presence of the virus with episomal forms (Fig. 2A, 2B).
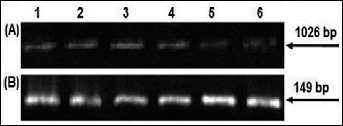
Detection of HPV-16 physical status. (A) PCR products of HPV-16 episomal forms from cervical cancer samples: lanes 1–6 are shown as examples (amplicon length 1,026 bp). (B) Transcripts product for HPV-16 E2-positive confirming the PCR results: lanes 1–6 (amplicon length 149 bp).
A lack of amplification of the E1 and/or E2 genes indicates the presence of disrupted genes, because of the HPV genome integration into the host cell genome.
In this study, the E1 RT-PCR analysis has shown integration in 1/11 (9%) benign samples and in 11/24 (45.8%) cervical cancers (Tab. III, Fig. 3). Based on these data, we found a significant difference in the physical state of HPV-16 E1/E2 genes between cases and controls (p<0.05). The combined integration results in cervical carcinomas with both the E1 and E2 integrated genes; analysis of these samples showed that the disruption occurred more frequently in the E2 gene (62.5%) than in the E1 gene (45.8%) (Fig. 4).
E1 Gene Status in HPV-16-Positive Benign Lesions and Cancers
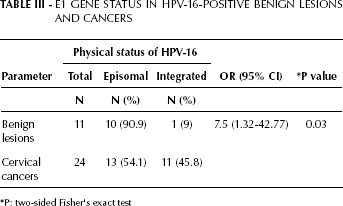
P: two-sided Fisher's exact test
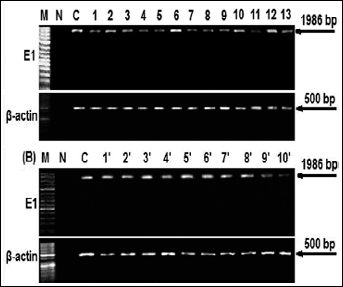
Transcripts product for the HPV-16 E1 gene. (A) mRNA expression of HPV-16 E1 from cervical cancer samples: lanes 1–13 and (B) from benign lesions: lanes 1′-10′ (amplicon length 1,986 bp); β-actin was used as control; M, size marker; N, negative control (normal cervix not infected with HPV); C, positive control (HPV-16 plasmid).
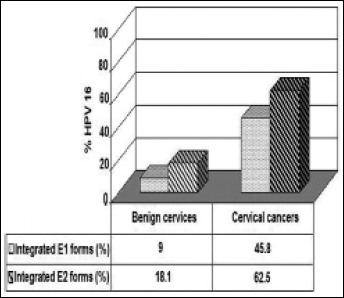
HPV-16 E1/E2 genome disruption associated with cervical malignancy.
Prevalence of Episomal and Integrate-Derived Transcripts
Using RT-PCR, we found that the major E6 and E7 transcripts were detected in 22 of the 24 (91.6%) HPV-16 cervical carcinomas, and in 5 of the 11 (45.4%) HPV-16 benign cervical lesions (Tab. IV). To assess whether the integration status might have an influence on malignant progression, we analyzed the relative transcript levels. The RT-PCR product levels were normalized to β-actin expression. Quantification was done using densitometric analysis. In the majority of cases E6/E7 transcripts originated from integrated as well as from episomal HPV-DNA with different levels of expression. Conversely, E2 transcripts originated only from the episomal forms.
Prevalence of Hpv-16 E6/E7 Transcripts in Cervical Samples
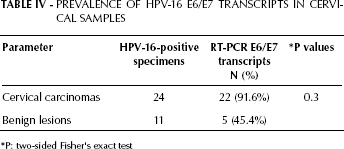

P: two-sided Fisher's exact test
As shown in Figures 5, 6A, and 6B, the mRNA levels of the HPV-16 E6/E7 genes were expressed at approximately the same levels in each physical state. We consistently observed that the E6/E7 levels were absent, decreased, or weakly detectable in the presence of E2 (p<0.05) (Fig. 6A and 6C). However, in the absence of E2 the levels of E6/E7 markedly increased (p<0.05) (Fig. 6B and 6D).
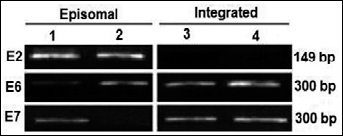
RT-PCR products of HPV genes’ transcripts. Lanes 1, 2: episomal form of HPV-16; lanes 3, 4: transcripts product for HPV-16 E6/E7 indicative of integrated form. Lane 4: more intensive RT-PCR product for E6/E7 genes derived from the integrated form.
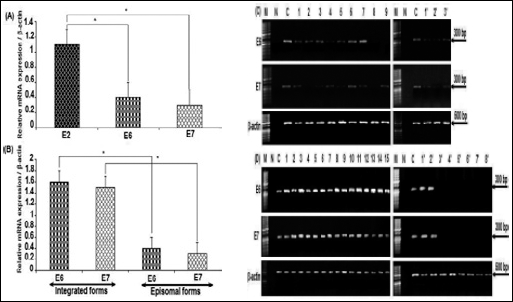
Expression and semiquantification of target genes in cervical samples. (A) Specific gene expression of target genes (E2, E6, and E7) relative to β-actin expression levels in episomal forms. (B) Comparison of specific gene expression of target genes (E6 and E7) relative to β-actin expression levels in episomal and integrated forms. Bars represent means ± SEM. *Significant elevated expression (p<0.05) according to the Student's t-test. (C) RT-PCR products of HPV-16 E6/E7 transcripts in episomal forms. Lanes 1–9: cervical cancer samples; lanes 1′-3′: benign lesions. (D) Transcripts product for HPV-16 E6/E7 indicative of integrated form. Lanes 1–15: cervical carcinoma samples; lanes 1′-8′: benign lesions. β-actin indicates positive control; M, size marker; N, negative control (normal cervix not infected with HPV); C, positive control (HPV-16 plasmid).
Discussion
In this report we investigated the degree of HPV-16 genome integration during carcinogenesis of cervical carcinoma. We showed that integration of viral DNA into the host genome takes place very early in cancer development and that the phenomenon might be a potential prerequisite for the first step in the multistaged development of cervical cancer. Disrupted viral DNA is prevalent in high-grade cervical lesions when compared to low-grade lesions. We identified disruption of the E2 gene in 62.5% of cervical carcinomas but only in 18.1% of benign cervical lesions. These data are consistent with the fact that viral integration is found more frequently in advanced tumor stages than in early ones. Integration of HPV-DNA into the human genome has been previously considered as a late event in carcinogenesis, but recent studies implicate that it occurs already in early lesions and even in clinically normal epithelium (27).
HPV-E2 mRNAs were constantly expressed in the presence of the intact virus. The PCR technique is rather time-consuming and laborious, however it allows accurate determination of the site of integration in the human genome. On the other hand, the RT-PCR technique is extremely suitable to evaluate the dynamic expression of the specific genes and seems to be more sensitive than the DNA PCR method in detecting intact genome because of the gene copy number. For that reason, the RT-PCR for HPV-16 E1 can be used for the determination of genome integration. Integrated HPV-16 E1 has been found in 11 of 24 (45.8%) HPV-16-positive samples in patients with cervical carcinoma, among which 62.5% presented disruption of the E2 gene. These combined findings indicate that HPV integration occurs frequently in the HPV-16-related cervical transformation process and that the E2 gene seems a more suitable target than E1 to identify HPV-16 DNA integration into the cellular genome. In accordance with several other investigators, this region appears to be deleted or disrupted in the majority of cervical carcinomas (28, 29) and pre-neoplastic stages (11).
The HPV-16 E2 protein has also been shown to be expressed at relatively high levels in benign lesions compared to cancers (30). Consequently, the detection of HR-HPV-E2 transcripts might serve as an additional biomarker of cervical diseases. However, expression of the early viral proteins E6, E7, and E2 play an important role in the disruption of cell-cycle control mechanisms and are, therefore, likely to be pivotal in the acquisition by infected cells of subsequent genetic abnormalities. In particular, as E2 negatively regulates transcription of the E6 and E7 genes, a loss of function of E2, through disruption, leads to an upregulation of the HPV early promoter and increased expression of the E6/E7 proteins (21, 31). The overall prevalence of E6/E7 mRNA transcripts in cervical carcinoma specimens (91.6%) is higher than that in benign cervical lesions (45.4%). Results from the current study also indicate that the transcriptional activity of HPV-16 E6/E7 is more indicative of their molecular activity. The detection of HR-HPV E6/E7 transcripts might serve as an additional factor for risk evaluation in the development of cervical disease (32–35). It will be also interesting to understand whether a correlation between the detection of integration and transcriptional activity can be demonstrated. E2 proteins are transcription factors that can repress transcription of the E6 and E7 genes of the genital papillomaviruses through several distinct cellular complexes (36). Several studies have shown that E2 proteins regulate the expression of the viral oncoproteins E6 and E7 (37). In this study, we identified a direct interplay between the HPV-16 E2 and E6/E7 proteins. E6/E7 transcripts originate from integrated, as well as from episomal HPV-DNA, whereas E2 transcripts originate only from the episomal HPV-DNA. Initially, we observed E2 inhibition of E6/E7 expression. Taken together, this research clearly shows that deregulated expression of the viral E6 and E7 oncogenes is the predominant hallmark of cervical carcinogenesis. Disruption of the viral E2 gene in cells with integrated HPV genomes may release inhibition of the viral promoter that controls the oncogenes E6 and E7. This implies that reintroduction of an intact E2 gene in cervical cancer cell lines (HeLa) results in growth inhibition due to inhibition of E6/E7 expression, apoptosis, and senescence (38). HPV-16 harboring intact E2 in the episomal form is often found to coexist with disrupted forms in cervical carcinomas. However, after the initial integration event, the remaining episomes can still repress the E2 protein, and repression of the integrated HPV genome may leave the cellular phenotype unaltered. Thus, the total loss of E2 expression may lead to malignant lesions that contain only integrated HPV genomes (39).
Describing the presence of intact E2 genes in almost 37.5% of HPV-16-positive cervical cancer cases has subsequently paved the way for new paradigms of cervical carcinogenesis. However, E2 disruption is not a prerequisite for the development of an invasive disease, as intact E2 genes have been identified in invasive lesions (40). The genetic instability of the E2 DNA sequence may in some manner enhance the overexpression of E6/E7 oncoproteins resulting in rapid progression to aggressive malignancy. Different studies showed the association of E6/E7 expression with either methylation or sequence variations in LCR. Moreover, during progression of cervical cancer, integration of HPV increases with a simultaneous decrease in methylation. An higher expression of E6/E7 in episomal unmethylated samples than episomal methylated samples suggests that expression of these genes is under the control of a promoter/enhancer. However, in the integrated form, this type of controlling mechanism is lost, as the viral genome is under the control of the stretch of host DNA where it integrates (41).
The present study may provide valuable information for the evaluation of cervical cancer development: measurement of the HPV-16 E6/E7 transcripts levels and evaluation of the E2 physical status could serve as an improved risk factor rather than the detection of the physical state of viral DNA.
In summary, the results presented here suggest that physical status and expression of the viral oncogenes can be assessed in a high-throughput manner, thus facilitating the detection of markers that predict cervical progression. The transcriptional function of the viral E2 gene appears to be the most appropriate target for the development of therapeutic strategies aimed at the treatment of viral infection and virus-induced cancers. Gene delivery may provide new therapeutic approaches for the treatment of cancer. Therefore, future therapies directed against the viral oncoproteins may be effective.
Footnotes
Acknowledgments
Special thanks to Dr. Alison McBRIDE, Chief, DNA Tumor Virus Section, Laboratory of Viral Diseases, at the National Institute of Allergy and Infectious Diseases, NIAID, NIH, Bethesda, Maryland, USA, for the constructive comments and linguistic revision of the manuscript. Financial support for this work was provided by a grant from the Ministry of Higher Education and Scientific Research in Tunisia.
