Abstract
Background
Transrenal DNA can potentially be useful in the disease management of colorectal cancer (CRC) and has potential diagnostic utility. Our study aimed to conduct preoperative and postoperative analysis of KRAS-positive patients, using transrenal DNA compared with plasma DNA. This is critically needed for disease diagnostics and treatment response monitoring.
Methods
CRC patients at different stages of the disease and with different molecular profiles were recruited for serial time-point analysis. Preoperative and postoperative urine specimens were extracted and compared with tumor tissues and plasma DNA.
Results
Our analysis demonstrated that transrenal DNA offered comparable sensitivity and specificity to plasma DNA analysis. Collection of transrenal DNA, being noninvasive, is an attractive assay and easily allows serial monitoring of the disease. Results from preoperative detection showed a close correlation to tumor tissue profiling and demonstrated close associations to the disease. We also observed significant decreases in mutant KRAS DNA concentration after surgery, which confirmed transrenal DNA's sensitivity to treatment response.
Conclusions
The use of transrenal DNA offers a possible alternative method of disease profiling, detection and tracking. Our study is one of the first to systematically analyze a relatively large number of different CRC patients using transrenal DNA. The positive correlation with disease demonstrates the assay's viability in the clinical setting, and our method further opens up the possibility to use transrenal DNA for clinical intervention investigations.
Introduction
Colorectal cancer (CRC) is one of the more prevalent cancer types and a leading cause of cancer deaths worldwide (1, 2). Several risk factors have been linked to the disease – namely, age, family history, diet, obesity, smoking and alcohol consumption (3, 4). Greater public awareness has led to more patients being detected early (5). and CRC can be cured by surgical means if the disease has yet to metastasize (6). Cur rently, colonoscopy is the gold standard to confirm CRC. Stool and blood–based assays complement the screening process by picking up asymptomatic individuals. Nonetheless, a significantly large proportion of CRC patients are still being diagnosed with distant metastases (7). This highlights the need for alternative detection methods that can aid in better early diagnosis.
A promising candidate is circulating tumor DNA (ctDNA) present in bodily fluids. For instance, ctDNA has been detected in the plasma or serum of patients with different types of cancer including CRC (8, 9). It contains signatures of driver mutations associated with the disease and is closely linked to disease outcome (10, 11). Given that plasma can be extracted from blood specimens, this makes it ideal for sampling, unlike tissue extraction that requires invasive surgery (12). The origins of these DNA materials appear to be from apoptotic cells that shed nucleic acid into the body's systemic and pulmonary circulatory system (13). The literature has shown that the DNA consists of small molecular weight fragments (14, 15). Given the small size of these fragments, they easily pass through the kidney barrier (16) and are excreted in the urine. Su et al, for instance, showed very sensitive detection of mutated KRAS DNA in urine could be significantly higher than in serum or plasma in certain conditions, which opens up the possibilities for noninvasive disease screening, diagnosis and prognosis (17). Transrenal DNA therefore may offer a new perspective for cancer diagnostics.
Our study aimed to investigate the clinical relevance of transrenal DNA in CRC patients. The main reasons for using urine specimens are the ease in sample collection and also the fact that it is technically easier for DNA isolation. Compared with surgery or blood extraction, urine collection is more straightforward and patient friendly. The technical challenges, however, are the low mutant DNA frequencies and the harsh environment of urine samples for DNA (18). Here the potential of transrenal DNA in CRC detection will be explored using the KRAS mutation, which is a fairly prevalent mutation in CRC. KRAS is a part of the RAS protein group of GTP/GDP binding proteins. The incidence of KRAS mutations in CRC is approximately 40% (19), and recent analysis has shown that it is linked to resistance to cetuximab and panitumumab treatment (20, 21). Besides characterizing the molecular signature, our study aimed to ascertain the stability and specificity of transrenal DNA to enable its use in a clinical setting.
Materials and methods
Study design and patient profiles
For the current study, 150 CRC cases were analyzed. The characteristics of the patients are shown in Table I. Informed consent was sought from each of the participants, and the methodologies used were approved by the institutional review board (IRB, Jingzhou Central Hospital). Altogether, the study involved 30 patients with stage I carcinomas, 40 with stage II, 40 with stage III and 40 with stage IV. Patients were selected randomly but followed certain inclusion and exclusion criteria. To be included, patients had to be treatment naïve and to have tissue specimens available for mutational analysis. Tumor tissue samples were extracted via either colonoscopy or surgical resection. Patients with insufficient or low quality samples were excluded. This was important as the tissue specimens served as the reference mutational profiles for subsequent transrenal and plasma DNA comparisons. Peripheral blood and urine samples were collected at the same time during each visit. For postoperative sampling, serial specimens were collected for a total duration of 3 months where possible. We noted that 32 stage IV CRC patients did not undergo surgical procedures as their disease had progressed beyond surgical control. Instead chemotherapy was given. As a negative control for the study, 40 healthy volunteers were recruited and their blood and urine specimens analyzed.
CRC patient and healthy volunteer characteristics
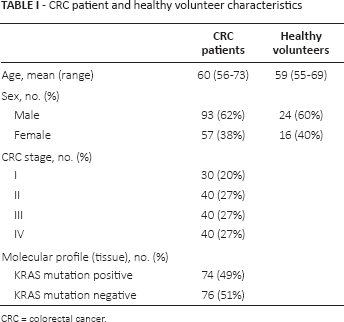
CRC = colorectal cancer.
Sample collection and DNA isolation
Altogether, 4 blood and urine samples were taken from each patient during the trial. Serial samplings were conducted at 1-month intervals. For peripheral blood collection, 10 mL of blood was taken. Blood was drawn in ethylenediaminetetraacetic acid (EDTA) vacutainer tubes and processed within 3 hours after collection. To extract plasma from whole blood, a 2-step centrifugation process was performed. The blood specimen was spun down at 1,000 g for 15 minutes at 4°C, and 4 mL of supernatant was removed carefully into a new centrifuge tube. A repeat centrifugation step with the same setting ensured that debris and cellular materials were excluded during DNA isolation. For urine samples, 30-50 mL of midstream urine was collected into sample holders that contain EDTA (Sigma-Aldrich, China). Samples were processed within 3 hours using high-speed centrifugation. Specimens were processed using a setting of 10,000 g for 10 minutes to remove all contaminants. Then 10 mL of supernatant was carefully removed into a fresh centrifuge tube. To sum up, 4 mL and 10 mL of supernatant from whole blood and urine, respectively, were extracted for subsequent DNA profiling.
For DNA isolation, both specimens were processed with a QIAamp Circulating Nucleic Acid Kit (Qiagen Inc., USA) following the manufacturer's instructions. The purified genomic material was eluted in 20 μL Tris-EDTA (TE) buffer. Quantification was performed on a Nanodrop 2000 (Thermo Fisher Scientific, USA). The nanodrop instrument measures DNA concentration by measuring the absorbance of the sample at 260 nm, adjusted for various parameters and using the relationship at A260: 1.0 = 50 μg/mL pure double-stranded DNA (dsDNA). The extracted DNA was stored at −20°C prior to molecular analysis. For tissue samples, DNA was extracted using a DNeasy Blood & Tissue Kit (Qiagen Inc., USA) following the manufacturer's protocols.
Detection of KRAS mutation in plasma and urine samples
Molecular profiling of tumor tissue DNA and ctDNA for KRAS mutations was done using droplet digital polymerase chain reaction (ddPCR; QX100; BioRad Laboratories Inc., USA). The protocols used strictly followed the manufacturer's recommendations. Briefly, probes for KRAS mutations (G12/G13) were obtained from BioRad's PrimePCR ddPCR mutation assays (BioRad Laboratories Inc., USA) using a multiplex approach. A reaction mix for ddPCR was prepared by adding DNA templates from plasma or urine specimens and probes to the master mix. Each reaction volume was 20 μL, and the thermocycling conditions were as follows: 95°C for 10 minutes, followed by 40 cycles at 94°C for 30 seconds and 55°C for 1 minute. Finally, the reaction mix was set at 98°C for 710 minutes for enzyme deactivation and to be ready for detection on the QX100 droplet reader (BioRad Laboratories Inc., USA). The analysis was performed using BioRad QuantaSoft software (BioRad Laboratories Inc., USA).
Statistical analysis
All categorical variables were tested with the Shapiro-Wilk normality test. Comparisons of DNA extracted from different study groups were performed using a 1-tail Student's t-test. A p value of less than 0.05 was considered statistically significant. The quantitative comparisons among patients of different stages were made using ANOVA. The agreements between different sample types were measured using Cohen's kappa coefficient, which is robust in determining inter-rater reliability. The calculation was based on the simple estimate proposed by Cohen in his original paper on kappa (22). To ascertain the clinical value of transrenal DNA in CRC patients, we constructed receiver operating characteristic (ROC) curves and calculated the areas under the curve (AUCs). We made comparisons of the AUC measurements using the Hanley-McNeil method. All statistical analyses were carried out using Prism software (Graphpad, USA).
Results
Study design and association of transrenal DNA with CRC patients
The source of transrenal DNA is suggested to be mainly cell-free DNA circulating within the pulmonary and systemic circulation (16). Small-molecular-weight DNA fragments cross the kidney barrier and are present in urine. To confirm if this phenomena was also associated with CRC patients, we examined the nucleic acids from the urine of healthy volunteers and CRC patients. The isolated DNA was quantified using spectrophotometry. For the current study, 150 CRC patients with varying stages of the disease and 40 healthy volunteers were recruited. Table I gives details of the characteristics of the study cohorts.
Figure 1A shows the baseline measurements of transrenal DNA quantity in healthy volunteers and CRC patients before treatment. As a comparison, we tabulated the quantity of the plasma DNA in these study groups as well (Fig. 1B). The results clearly highlighted the differences between healthy participants and CRC patients. For both plasma and transrenal DNA, higher quantities of genomic materials were detected in CRC patients than in healthy volunteers. Among the different patient subgroups (stages I-IV), using the ANOVA test, there was a statistically significant difference in the mean recovered transrenal DNA (p<0.01). Average DNA quantity obtained from healthy volunteers was 0.423 ng/mL (95% confidence interval [95% CI], 0.346-0.500 ng/mL). CRC patients with disease stages I-IV had 0.694 ng/mL (95% CI, 0.5380.850 ng/mL), 1.61 ng/mL (95% CI, 1.54-1.68 ng/mL), 1.80 ng/mL (95% CI, 1.67-1.93 ng/mL) and 2.70 ng/mL (95% CI, 2.61-2.78 ng/mL), respectively. Based on the mean quantity of DNA in the different groups of patients, we observed a monotonic increase with stage of the disease. Similar results were obtained using plasma DNA as well (Fig. 1B). With ANOVA, we found a p value <0.05 – i.e., significantly different.
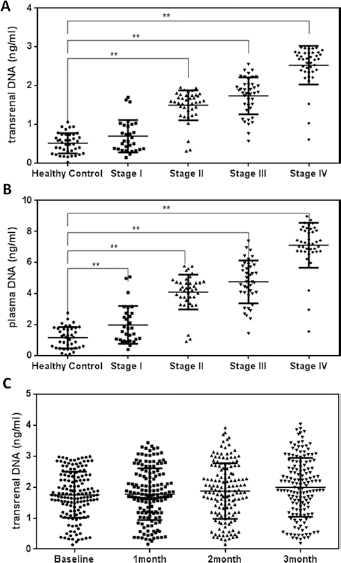
Measurement of total cell-free DNA among different study cohorts. (
Figure 1C shows the serial measurements of transrenal DNA quantities for the CRC patients. Slight variations in mean DNA quantities were observed, but these were not statistically significant. It is to be noted that this analysis did not take into consideration adjuvant therapy given to patients, but nonetheless, the results demonstrated stable release of nucleic acid into the urinary system. This provides a viable source for disease profiling.
CtDNA from transrenal DNA is strongly associated with disease
We hypothesized that transrenal DNA carries mutant DNA that is associated with CRC. To evaluate this, we compared the ctDNA analysis derived from ddPCR, with the molecular profiles from tissue samples. Table II shows the breakdown of sample types and the agreement in molecular profiles with transrenal DNA. The tissue molecular profiles of CRC patients included 49% that contained mutant KRAS. Overall concordance among CRC patients for transrenal DNA with biopsy samples was 80% (k = 0.84; 95% CI, 0.75-0.93). With plasma DNA, the concordance was also 80% (k = 0.83; 95% CI, 0.74-0.92). When we compared transrenal and plasma DNA, the agreement between them was 98%. We observed that the result was skewed due to stage I CRC patients. Excluding stage I patient outcomes, the concordance rates were 93% (k = 0.94; 95% CI, 0.88-1.00) for transrenal DNA and 90% (k = 0.91; 95% CI, 0.84-0.98) for plasma DNA.
Concordance of tissue specimens vs. urine, and tissue specimens vs. plasma for KRAS mutations in CRC patients
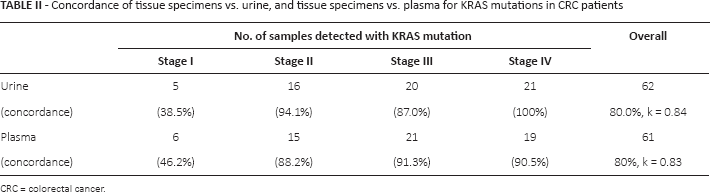
CRC = colorectal cancer.
We further performed a series of ROC analyses comparing healthy individuals and CRC patients based on staging, to determine the clinical value of transrenal DNA. The AUCs using transrenal DNA were 0.69 (95% CI, 0.57-0.82), 0.96 (95% CI, 0.91-1.01), 0.99 (95% CI, 0.97-1.01), and 0.99 (95% CI, 0.98-1.01) for stage I-IV patients, respectively, as shown in Figure 2A.
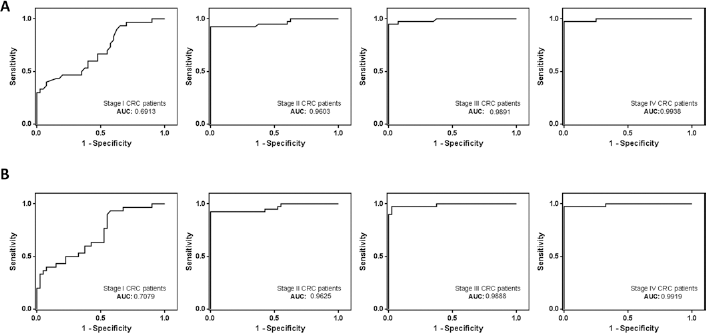
Receiver operating characteristic (ROC) curve analysis of healthy volunteers compared with colorectal cancer (CRC) patients. (
As a comparative study, we performed the same analysis using plasma DNA results. The AUCs were 0.71 (95% CI, 0.59-0.83), 0.96 (95% CI, 0.92-1.01), 0.99 (95% CI, 0.97-1.01) and 0.99 (95% CI, 0.98-1.01) for stage I-IV patients, respectively, as shown in Figure 2B. There were no statistically significant differences between AUC measurements derived from plasma and transrenal DNA (p>0.05, in all cases).
We also attempted to further define cutoff values for each condition based on our ROC curve analysis. The results are depicted in Supplementary Figure 1A for transrenal DNA (available online at www.biological-markers.com) and Supplementary Figure 1B for plasma DNA (available online at www.biological-markers.com). This may help to stratify patients against healthy individuals for possible diagnostic applications.
Trends in ctDNA from transrenal DNA associated with treatment
We performed a time-point analysis of ctDNA variations for KRAS-positive individual patients to determine if this could be associated with treatment. Baseline samples were collected prior to treatment, and 3 separate collections were made at monthly intervals after treatment. Figure 3A highlights the results from transrenal DNA analysis, and Figure 3B from plasma DNA. It was clear that immediately after surgery, there was a significant drop in ctDNA quantities for both sample types (transrenal and plasma DNA). This was confirmed using a paired Student's t-test, and irrespective of tumor staging. The decrease was most pronounced in stage III patients, who registered a drop of more than 22% from transrenal DNA analysis. This was also much higher than what was captured from plasma DNA analysis.
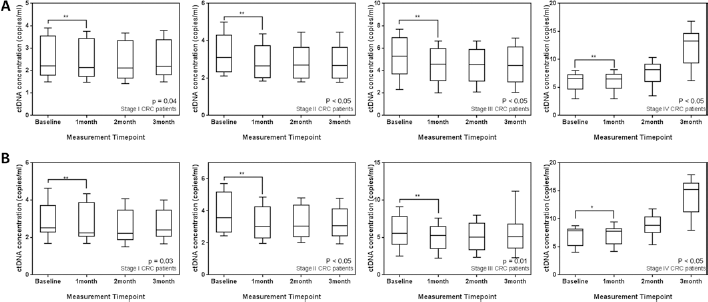
Mutant KRAS concentration in cell-free DNA. (
A secondary trend that was observed in these KRAS-positive CRC patients was that subsequent monitoring of ctDNA concentrations showed little variation except for stage IV patients. For stage I-III CRC patients, the mean concentrations of ctDNA were maintained throughout the remaining 2 measurement time points. Using ANOVA, a statistically insignificant difference was found among stage I-III patients from transrenal DNA analysis (p = 0.14, p = 0.86 and p = 0.51, respectively). For stage IV patients, we observed an increase after the second serial sampling, and this continued into the third sampling. Overall, stage IV patients showed a twofold increase above baseline measurements before treatment.
Discussion
The need for alternative methods of detection and monitoring of CRC patients cannot be overemphasized. CRC patients benefit from numerous available treatment options, but early detection of the disease is key to enhancing the survival rates of these patients (23-25). Liquid biopsies using plasma and transrenal DNA offer a relatively easy method to extract tumor material that can be used for disease correlation. Though many studies have been reported for plasma DNA (26-28), the use of transrenal DNA in CRC is relatively new. Compared with blood extraction, collection of urine specimens is noninvasive and more patient friendly. In the literature, it has been demonstrated that ctDNA could be detected in urine specimens of early and advanced stage bladder cancer with good sensitivity (29). In the present study, we demonstrated that KRAS mutations can be clearly detected in different CRC patients.
Firstly, our study demonstrated that the quantity of isolated DNA from urine was significantly higher than that from healthy volunteers (Suppl Fig. 2, available online at www.biological-markers.com). This suggests that the urine specimens of CRC patients contain materials that could potentially be related to the disease. More importantly, it was also determined that the amount of DNA material remained quite stable when grouped by patients of different tumor staging. This offers the potential to use transrenal DNA as a diagnostic marker for the disease.
Comparing the results of transrenal DNA with plasma DNA, we observed similar trends between them. The agreement with plasma DNA results indicated that transrenal DNA is closely associated with ctDNA in blood circulation. Our results finding higher DNA content in individuals with disease are consistent with numerous other studies that analyzed plasma DNA (30-32). For instance, Spindler et al found a more than sevenfold higher DNA content for positive metastatic CRC patients compared with healthy controls (30). Though their results showed much higher levels than in our current study, this can be attributed to the different patient profiles in the 2 studies and different methods of sample preparation. We observed that late stage patients (III and IV) in our study showed higher plasma DNA content akin to the results in that study performed by Spindler et al (30). More importantly, we demonstrated that transrenal DNA had similar trends and could potentially be substituted for or complement plasma DNA assays. We further provided evidence of the stable release of DNA in time-point measurements that will enable its use in clinical settings.
Molecular profiling of a disease is conventionally done using tumor tissue samples. The drawback to this is that these samples are hard to obtain and not feasible for serial time-point analysis. In the current study, we demonstrated the stable release of DNA in urine specimens that could be linked to the disease, which is suitable for molecular testing. Using ddPCR, we showed that KRAS mutations can be readily picked up from transrenal DNA and that the results are in good agreement with tissue samples in late stage CRC patients. The concordance rate for patients with stage IV CRC was as high as 100% and demonstrated the strong sensitivity and specificity of the transrenal DNA assay. Our results are consistent with those in a study by Su et al, which showed, in a smaller cohort of 5 CRC KRAS-positive patients, that tissue analysis was entirely concordant with analysis of urine specimens (16). Su et al further observed that the source of this low-molecular-weight class of urine DNA was blood circulation. Possibly, this opens up more options for molecular profiling of the disease.
Our study also further expanded on prior studies with the analyses of a wider spectrum of patients of different tumor staging and profiles. Transrenal DNA could also aid in cases where tumor tissues are hard to reach during initial colonoscopy. In our tests, stage I CRC patients did not fare very well, possibly due to the localization of the tumor and the lack of DNA releasing into the blood circulation. This in turn affected the detection in the transrenal DNA. The low concordance rate was expected for the stage I patient group, but it was surprising to observe that 5 patients were still able to be detected accurately for the mutation. We noted that this concurs with another study on non-small cell lung cancer (NSCLC) for early stage patients after surgery, that found that key driver mutations were still detectable (33). We further performed ROC analysis on total transrenal DNA, comparing patients with healthy individuals to gauge the performance of transrenal DNA in disease discrimination. AUC measurements for patients with different stages showed a value as high as 0.99, which indicated good accuracy in the tests. This supports the use of transrenal DNA in the disease management of CRC.
It is interesting that in our preoperative and postoperative serial measurements, the mutant DNA detected decreased for KRAS-positive patients in correlation with stages I-III. This could be indicative of treatment response and suggest the possibility of disease monitoring using transrenal DNA analysis. Indeed, prior studies have shown similar results for plasma DNA in NSCLC (34). Serial monitoring of ctDNA in these studies demonstrated close correlation to disease outcome, and was strongly linked to survival and disease progression. Our results showed that transrenal DNA could potentially do the same in CRC and that the decrease in ctDNA concentrations could be associated with the removal of tumors during surgery. We hypothesized that ctDNA in transrenal DNA could potentially be linked to tumor burden, and could be used for tracking treatment and disease progression in CRC. As evident from our study observations, stage IV CRC patients who were mostly on maintenance treatment without surgery saw significant changes in their corresponding ctDNA variations. This noninvasive approach for molecular profiling will benefit patients and allow longer-term serial urine samplings to track their disease progression.
The main limitation of the current study was the limited number of patients in each subset that were analyzed and the lack of consideration given to any adjuvant therapies given to different patient groups. This study, however, lays the foundation for further work on transrenal DNA for CRC patients. Our study is one of the first to address the specificity and sensitivity of using KRAS mutations to profile CRC patients, and it demonstrated that transrenal DNA can be an effective source of tumor material. It is, however, to be noted that the levels of transrenal DNA might not be just associated with cancer alone. And yet, its ease of extraction and use for serial monitoring makes it ideal to complement current disease management routines. Other issues were mainly associated with the detection limit for ddPCR, as early stage patients, particularly those after surgery, would have low or no circulating mutant DNA. ddPCR is one of the most sensitive methods for detection without the need for further mutant DNA purification or enrichment (35), which is why this was selected as our detection assay. Other techniques such as ICE COLD-PCR (36) could potentially aid in further improving the detection limits for early stage patients. Our study establishes a basis for a larger clinical study to encompass patients with different molecular profiles and to investigate clinical interventions.
Conclusions
The investigation of transrenal DNA for the diagnosis and treatment monitoring of CRC is promising. Our study clearly demonstrated the stable signatures of transrenal DNA in mutant KRAS-positive CRC patients, which can be exploited to complement current disease management routines. While colonoscopy remains the gold standard to confirm diagnosis, transrenal DNA analysis can play a role for sensitive prescreening and subsequent treatment monitoring. Our results showed a close correlation with tumor tissues, which were comparable to plasma DNA analysis. This supports the use of transrenal DNA, where specimen extraction is more patient friendly and highly responsive to disease changes. Our methods and workflow could also be applicable to other patient subtypes to better address CRC. Continued development of transrenal DNA analysis may improve current screening processes and better stratify patients to improve treatment efficiency.
Footnotes
Financial support: This work was supported by a research grant provided by Jingzhou Central Hospital (201116374).
Conflict of interest: All authors declare they have no conflicts of interest.
