Abstract
Background
Peripheral blood mitochondrial DNA (mtDNA) copy number alteration has been suggested as a risk factor for several types of cancer. The aim of the present study was to assess the role of peripheral blood mtDNA copy number variation as a noninvasive biomarker in the prediction and early detection of renal cell carcinoma (RCC) in a cohort of Egyptian patients.
Methods
Quantitative real-time polymerase chain reaction (qPCR) was used to measure peripheral blood mtDNA copy numbers in 57 patients with newly diagnosed, early-stage localized RCC and 60 age- and sex-matched healthy individuals as a control group.
Results
Median mtDNA copy number was significantly higher in RCC cases than in controls (166 vs. 91, p<0.001). Increased mtDNA copy number was associated with an 18-fold increased risk of RCC (95% confidence interval: 5.065-63.9). On receiver operating characteristic curve analysis, it was found that mtDNA could distinguish between RCC patients and healthy controls, with 86% sensitivity, 80% specificity, 80.3% positive predictive value and 85.7% negative predictive value at a cutoff value of 108.5.
Conclusions
Our results showed that increased peripheral blood mtDNA copy number was associated with increased risk of RCC. Therefore, RCC might be considered as part of a range of potential tumors in cases with elevated blood mtDNA copy number.
Introduction
Renal cell carcinoma (RCC) is an extremely aggressive tumor. It represents 90% of renal tumors and 2%-3% of all cancers. RCC incidence has been increasing by about 2% annually in the last 2 decades. Several established risk factors have been identified including obesity, smoking, hypertension and family history. At diagnosis, about 30% of patients present with a metastatic disease, and these patients have a 5-year survival rate of only 5%. Despite recent improvements in treatment strategies, the prognosis remains poor, due to late presentation, tumor heterogeneity and lack of molecular biomarkers for early detection and classification (1, 2). Identification of new noninvasive molecular markers for prediction, early diagnosis and monitoring of treatment remains a major challenge in RCC management.
Mitochondria are essential for regulation of serious cellular functions such as apoptosis, calcium homeostasis and energy production, via oxidative phosphorylation (OXPHOS) reaction and production of free oxygen radicals (3). Each mitochondrion contains from 2 to 10 copies of circular 16.6 kb double-stranded maternally inherited mitochondrial DNA (mtDNA). MtDNA encodes 13 respiratory chain polypeptides, 22 tRNAs and 2 rRNAs. Each cell has from 10 3 to 10 4 copies of mtDNA, which varies according to the energy needs of the cell to maintain normal physiological functions (4). MtDNA is 10 to 200 times more susceptible to oxidative stress-mediated mutagenesis than nuclear DNA, due to absence of protective histone, deficient DNA repair system and high reactive oxygen species (ROS) exposure (5).
It was observed that altered mtDNA copy number was associated with several types of cancer, and it was suggested as a mitochondrial dysfunction marker, which predisposes to malignancy. Increased mtDNA content has been observed in lung cancer (6), pancreatic cancer (7), prostate cancer (8), endometrial cancer (9) and B-cell lymphoma (10), while decreased mtDNA content was found in hepatocellular carcinoma (11) and esophageal adenocarcinoma (12).
Contradictory results were obtained when the association between RCC and peripheral blood mtDNA copy number variation was investigated. Both an increased and decreased mtDNA copy number have been reported to be associated with RCC risk (13-16). In addition, none of these studies investigated the diagnostic value of mtDNA in early detection of RCC.
The aim of the current study was to assess the role of peripheral blood mtDNA copy number variation as a noninvasive biomarker in prediction and early detection of RCC in a cohort of Egyptian patients.
Materials and Methods
Study participants
The study was conducted on 57 patients with early localized (TNM stage I and II), newly diagnosed RCC. Their ages ranged from 50 to 77 years. They were consecutively recruited from the urology unit at the Main Alexandria University Hospital between August 2012 and March 2015. The diagnosis was confirmed by histopathological examination. Patients with advanced RCC (TNM stage III and IV) were excluded from the study to omit the effect of disease progression on mtDNA content. Patients with other malignancies, family history of kidney cancer, recurrent RCC, systemic diseases and other abnormalities of renal origin as well as patients who had received neoadjuvant therapy were excluded from the study. Sixty healthy individuals with matching age and sex were included as a control group. The ethics committee at Alexandria University approved the study, and informed consent was obtained from all patients and controls.
Relative quantification of mtDNA copy number using real-time PCR
DNA was extracted from whole blood ethylenediaminetetraacetic acid (EDTA) specimens using a PureLink® genomic DNA kit (Life Technologies, CA, USA) according to the manufacturer's instructions. The concentration and purity of DNA were measured using a NanoDrop 2000/2000c Spectrophotometer (Thermo Scientific, USA). The DNA samples were kept at −20°C until further analysis was done.
Relative mtDNA copy number was measured using quantitative real-time polymerase chain reaction (PCR) assay on an Applied Biosystems 7500 (Applied Biosystems, USA). Human hemoglobin b (HBB) gene was used as reference single-copy nuclear gene (nDNA). Real-time PCR was performed in 25 μL of reaction mixture including 2X SYBR® Green PCR Master Mix, 0.4 μM forward primers, 0.4 μM reverse primers and 50 ng DNA. A negative control (no template control) was included in each run. Thermal profile included an initial activation step at 95°C for 10 minutes, followed by 40 cycles of 95°C for 15 seconds (denaturation step) and 60° for 10 minutes (annealing/extension). For the amplification of a portion of mtDNA spanning from 3,212 to 3,300 bp, the following pair of primers was used: forward primer, 5′-CACCCAAGAACAGGGTTTGT-′3, and reverse primer, 5′-TGGCCATGGGTATGTTGTTA-′3. The sequences of forward and reverse primers used for detection of nuclear gene (HBB gene) were HBB-F 5′-GCTTCTGACACAACTGTGTTCACTAGC-′3, and HBB-R 5′-CACCAACTTCAT CCACGTTCACC-′3 (15).
Data analysis
MtDNA copy number was determined relative to the nuclear gene (HBB gene) by comparative CT method using the formula 2ΔCt, where ΔCt = CTmtDNA-CT nDNA (HBB) for each sample (17-19).
Statistical analysis
Data were analyzed using SPSS, version 20, software. Qualitative data were described as numbers and percentages and were compared using the chi-square test. Normally distributed quantitative data were expressed as means ± standard deviation (SD) and compared using Student's t-test. The Mann-Whitney test was used to compare abnormally distributed quantitative variables, expressed as medians (minimum-maximum). The association between mtDNA and RCC risk was assessed by unconditional logistic regression analysis using univariate and multivariate models to determine the odds ratios (ORs) and 95% confidence intervals (95% CIs). The multivariate model was adjusted for age, sex, smoking status, body mass index (BMI) and history of hypertension. Significance of the obtained results was judged at the 5% level. Receiver operating characteristic (ROC) curves were used to determine the diagnostic performance of mtDNA. An area under the curve (AUC) of more than 50% indicates acceptable performance, and an AUC of about 100% is the best performance for the test. The required sample size was calculated using Epi-save software. The sample size estimate was made with an assumption of 95% confidence level and 80% power of study.
Results
Characteristics of study subjects
Fifty-seven patients (36 men, 21 women) with early localized RCC were included in the current study. Fifty cases (87.7% of our patients) were histopathologically diagnosed as clear RCC, 5 cases (8.8%) as papillary RCC and 2 cases (3.5%) as chromophobe RCC. According to the TNM staging system (2), 27 cases (47.4%) were stage I, and 30 cases (52.6%) were stage II. Based on the Fuhrman grading system (2), 24 patients (42.1%) were grade 1, 14 patients (24.6%) were grade 2, 14 patients (24.6%) were grade 3 and 5 patients were grade 4.
Sixty age- and sex-matched healthy individuals (36 men, 24 women) were included in the study as a control group. All of the study participants, patients and controls, were of Mediterranean origin.
The cases were more likely to have a history of hypertension than controls (p = 0.021). No significant difference was observed between patients and controls as regards smoking (p = 0.428) or BMI (p = 0.642). Patient and control characteristics are summarized in Table I.
Characteristics of study subjects
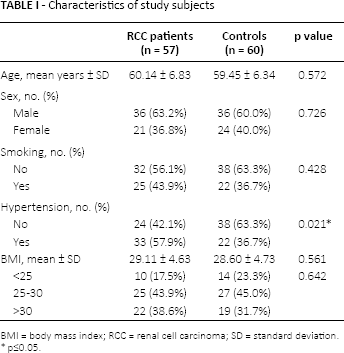
BMI = body mass index; RCC = renal cell carcinoma; SD = standard deviation.
p≤0.05.
Peripheral blood mtDNA copy numbers in RCC patients and controls
Patients had a significantly higher relative mtDNA copy number than controls (p<0.001). In the RCC group, mtDNA content ranged from 90 to 418.6 with a median of 166, while in the control group it ranged from 53.5 to 136.5 with a median of 91, as illustrated in Table II. MtDNA copy numbers reported in the present study are only applicable for assays measuring mtDNA content under the described conditions.
Peripheral blood mtDNA copy numbers in renal cell carcinoma patients and healthy controls
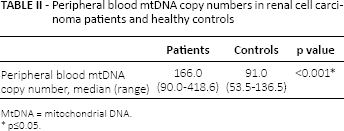

MtDNA = mitochondrial DNA.
p≤0.05.
In the RCC group, no significant association was found between mtDNA copy number and age (p = 0.064), sex (p = 0.18), smoking (p = 0.067), history of hypertension (p = 0.141) or BMI (p = 0.667). In addition, no significant difference was observed in mtDNA content between different RCC histopathological types and grades (p = 0.300 and p = 0.467, respectively) (Tab. III).
Association between mtDNA copy number and different characteristics in the renal cell carcinoma group

BMI = body mass index; mtDNA = mitochondrial DNA; RCC = renal cell carcinoma.
Similarly, in the control group, no significant difference was observed in mtDNA content as regards age, sex, smoking, history of hypertension or BMI (p = 0.604, p = 0.507, p = 0.830, p = 0.581 and p = 0.957, respectively).
Association between mtDNA copy number variation and RCC risk
The association between peripheral blood mtDNA copy number and RCC risk was evaluated using unconditional logistic regression analysis. The median mtDNA content of the control group (median 91) was used as a cutoff value to analyze mtDNA content. It was found that mtDNA content higher than 91 significantly increased the risk of RCC by 18-fold with an OR of 18.0 (95% CI, 5.065-63.9) on univariate analysis, and an adjusted OR of 18.9 (95% CI, 5.11-70.11) on multivariate analysis after adjusting for age, sex, smoking status, history of hypertension and BMI (Tab. IV).
Association between peripheral blood mtDNA content and renal cell carcinoma risk
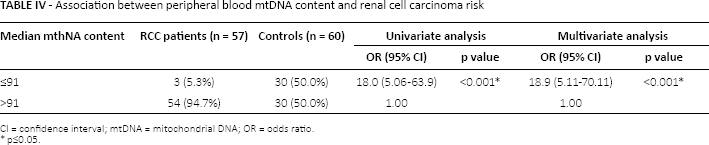

CI = confidence interval; mtDNA = mitochondrial DNA; OR = odds ratio.
p≤0.05.
Diagnostic significance of mtDNA in RCC
The diagnostic value of mtDNA content in early detection of RCC was further assessed. Using ROC curve analysis, it was found that mtDNA could detect RCC at a cutoff value of 108.5 with a sensitivity of 86%, specificity of 80%, positive predictive value of 80.3% and negative predictive value of 85.7%. The AUC was 0.926.
MtDNA copy number in patients with ccRCC
When we further focused on the group of patients (50 patients) with clear cell RCC (ccRCC), we found that the peripheral blood mtDNA copy number was still significantly higher in patients than controls. It ranged from 90.0 to 418.60, with a median of 166. Interestingly, by using logistic regression analysis, it was found that increased peripheral blood mtDNA above 91 (median mtDNA content of the controls) was associated with increased RCC risk, with an OR of 24 (95% CI, 5.343-107.81). Even after adjustment for age, sex, BMI, smoking and hypertension, the OR was 24.9 (95% CI, 5.37-115.99). By using ROC curve analysis, it was found that mtDNA at a cutoff value of 113.5 could discriminate patients with ccRCC from healthy controls with 80% sensitivity and 83.3% specificity. The AUC was 0.927.
Discussion
In the present study, the association between peripheral blood mtDNA copy number and the risk of RCC was evaluated in a cohort of Egyptian patients. A statistically significant association was observed between high mtDNA copy number and increased risk of RCC. We found that patients with peripheral blood mtDNA copy number above the controls' mtDNA median of 91, had an 18-fold increased risk of RCC. When blood mtDNA content was studied only in the group of patients with ccRCC, it was found that the risk was increased by about 24-fold.
No statistically significant difference was observed in mtDNA content between men and women. Age, smoking status, history of hypertension or BMI also did not affect mtDNA content among patients or controls.
Our study focused on newly diagnosed, localized and early-stage RCC to exclude the effect of disease progression on mtDNA copy number. Blood samples were also collected immediately after diagnosis to omit the effect of treatment. Among different RCC risk factors evaluated in our study, only hypertension was found to be significantly associated with RCC.
Our results were consistent with a previous prospective study by Hofmann et al, in which blood samples collected 6 years before diagnosis showed a statistically significant increase in mtDNA copy number in RCC patients. They suggested that increased mtDNA content might be a biomarker of RCC risk, which is related to RCC etiology and not a disease effect (13).
In previous years, numerous retrospective and prospective studies have demonstrated a powerful association between increased peripheral blood mtDNA copy number and the risk of different types of cancer (6, 7, 20-24). However, the link between blood mtDNA and carcinogenesis is not fully explained. As blood cells are transporter cells and mediators of immune response, they interact and come into contact with all tissues, carrying a lot of bioactive substances such as oxygen, nutrients and metabolites, and hormones to and from tissues (25). Thus, it was proposed that profiling of peripheral blood could represent a reflection on, and an efficient way to explore, disease pathogenesis (26).
In RCC, it was observed that ROS were significantly increased in tumor tissues as well as in blood samples. Furthermore, blood ROS increased progressively in advanced stages and decreased after curative resection. Oxidative stress was suggested as one of the factors responsible for the development of RCC (27, 28). Byproducts of oxidative stress-induced lipid peroxidation have been shown to react with DNA in renal cells to form adducts. Defects in the repair of adduct-induced DNA damage can result in DNA mutations, a crucial step in oncogenesis, especially if it occurs in proto-oncogenes or tumor suppressor genes (29).
High peripheral blood mtDNA content was suggested as an indicator of increased oxidative stress and ROS-induced DNA damage (13). ROS cause direct mtDNA damage, as mtDNA is in proximity to the inner mitochondrial membrane, where electron chain transport occurs (5, 30). MtDNA damage could result in OXPHOS system impairment with further increases in ROS production, accelerating DNA mutation rates (31, 32). MtDNA copy number was suggested to be increased in order to compensate for the metabolic defects in mitochondria harboring mutated mtDNA and a defective OXPHOS system (5, 33).
This theory is supported by the previous studies, which showed a positive association between the relative mtDNA copy number and oxidative stress markers including thiobarbituric acid reactive substances and 8-hydroxyguanosine in human leukocytes (34). In addition, when human lung fibroblasts were exposed to oxidative stress induced by sublethal concentrations of H2O2, an increase in the amount of mitochondria and mtDNA of the treated cells was observed (35).
Therefore, it is probable that a high mtDNA copy number is a marker of carcinogenesis-associated oxidative stress, rather than a cause of cancer development. However, future studies are needed to assess the correlation between mtDNA copy number in peripheral blood and renal tissue.
In contrast to our results, 3 previous retrospective studies showed an inverse association between peripheral blood mtDNA and the risk of RCC (14-16). This discrepancy may be due to study sample heterogeneity, different histological subtypes and different methods used for mtDNA calculation in their studies. These studies also measured mtDNA copy number 6 months to 1 year after diagnosis, so the results may be influenced by treatment effect, while patients included in the current study were treatment naïve.
A U-shaped association was observed between colorectal cancer and mtDNA copy number. This meant that patients with high and low mtDNA had an increased colorectal cancer risk (36).
In the present study, it was found that at a cutoff value of 108.5, mtDNA might detect early RCC with 85% sensitivity and 80% specificity. In addition, at a cutoff value of 113.5, it could discriminate patients with ccRCC from healthy controls with 80% sensitivity and 83.3% specificity. Therefore, mtDNA needs to be further evaluated as a potential biomarker for early RCC detection.
The role of mtDNA as a noninvasive biomarker has been investigated in different types of cancer. It was found that mtDNA content could differentiate between Ewing's sarcoma patients and healthy subjects with a sensitivity of 76.1 and a specificity of 68.4% (37). Moreover, Fernandes et al suggested that increased mtDNA could be used as a noninvasive biomarker for early detection of gastric cancer, with a specificity of 80% and a sensitivity of 47% (38).
Conclusion
Our results suggest that an increased peripheral blood mtDNA copy number is associated with increased risk of RCC. Therefore, RCC might be considered in the range of potential tumors in patients with an elevated blood mtDNA copy number.
Footnotes
Financial support: No grants or funding have been received for this study.
Conflict of interest: The authors declare that they have no conflicts of interest that are directly relevant to the content of this article.
