Abstract
The purpose of this study was to investigate the diagnostic value of the deletion of fragile histidine triad (FHIT) and p16INK4a (p16) mRNA in biopsies obtained by bronchoscopy. Biopsies were analyzed using RT-PCR in 52 patients with lung cancer and 19 patients with benign lung disease. The results showed that the detection rates of FHIT and p16 gene transcript deletion were significantly higher in lung cancer patients than in patients with benign lung disease (65.4% versus 10.5%, p=0.001 and 59.6% versus 5.3%, p<0.001, respectively). The sensitivities for detecting FHIT and p16 transcript deletion in biopsies were 65.4% and 59.6% (combined 80.8%), respectively, which were markedly better than those of histology and cytology (42.3% and 34.6%, respectively; combined 57.7%). In 22 lung cancer patients with negative histology and cytology at initial bronchoscopy, FHIT and p16 mRNA loss was detected in 40.9% (9/22) and 36.4% (8/22) cases, respectively. FHIT mRNA loss was associated with smoking status in lung cancer patients. In conclusion, deletion of FHIT and p16 mRNA can be identified in biopsies obtained during bronchoscopic procedures. FHIT and p16 mRNA deletion can be used as biomarkers in the clinical diagnosis of lung cancer and may serve as adjuncts to histology and cytology in lung cancer diagnosis.
Introduction
Lung cancer remains the most common cause of cancer deaths worldwide (1). The overall 5-year survival rate for stage I non-small cell lung cancer (NSCLC) patients who are typically treated with surgery is as high as 83%. In contrast, only 5% to 15% and less than 2% of patients with stage III and IV NSCLC, respectively, are alive after 5 years (2). Since there is a strong inverse correlation between prognosis and stage of the disease, efforts aimed at early identification of lung cancer are of major importance. Fiberoptic bronchoscopy is the method of choice for confirming the diagnosis of a suspected lung cancer by pathological assessment of tissue or cytology specimens obtained during the procedure. However, nondiagnostic results are common, and in about half of patients suspected of having lung cancer, the establishment of a final diagnosis after the first bronchoscopy fails (3), leading to the use of additional invasive procedures (4). The development of biomarkers that could detect tumor in tissue and cytology specimens should greatly improve lung cancer diagnostics (5, 6).
It is generally believed that the development of human cancer is a multistep process involving the clonal evolution of abnormal cell populations by accumulating specific alterations in at least 2 groups of genes, i.e. the activation of protooncogenes and the inactivation of tumor suppressor genes (7, 8). Fragile histidine triad (FHIT) is a tumor suppressor gene that spans the FRA3B at chromosome band 3p14.2, the most active of the common fragile sites in the human genome (9). Expression of FHIT mRNA is detectable in most tissues, and the highest levels of expression of FHIT mRNA and protein are detectable in epithelial cells and epithelial tissues (9, 10). A number of studies have demonstrated that FHIT expression is altered in a variety of human cancers and cancer cell lines, including those of lung, head and neck, stomach, cervix, breast and kidney (10–17). Aberrant FHIT transcripts were detected in 80-100% of small cell lung cancer (SCLC) and in 40-80% of NSCLC specimens (18). In addition, FHIT protein expression is lost or reduced in the majority of NSCLCs (12, 19–22). A study in a large series of stage I NSCLCs indicated that alteration of FHIT expression is an independent, early and frequent event in lung carcinogenesis and precancerous lesions (12). Thus analysis of aberrant FHIT expression may be used as a tool in early detection of lung cancer.
The p16 tumor suppressor gene that maps to chromosome band 9p21 is an important component of cell cycle regulation. The p16 gene encodes a cell cycle protein that is an inhibitor of CDK4 and CDK6, and negatively regulates cyclin D-dependent phosphorylation of the Rb gene product, thereby inhibiting cell cycle progression from G1 to S-phase by sequestration of E2F (23, 24). The p16 gene is inactivated in >70% of cell lines derived from all histological types of human NSCLC (23) and in up to 63% of tumor specimens of NSCLC patients (25) through homozygous deletion, promoter methylation of the gene, or, less commonly, point mutation (26–29). Sanchez-Cespedes et al reported that inactivation of the p16 gene is a common event in all NSCLCs, but the mechanism of gene alteration differs between smokers and nonsmokers (29). Some mechanisms of p16 gene alteration, such as promoter methylation, have been found to be an early event in lung cancer and may constitute a potential biomarker for early detection and monitoring of prevention trials (30). In the present study, we examined the deletions of FHIT and p16 mRNA in specimens obtained by bronchoscopic biopsy from patients with lung cancer and benign lung disease. Our aim was to investigate their potential diagnostic value as molecular markers of lung cancer.
Patients and Methods
Patients and Tumor Specimens
We prospectively analyzed the deletion status of FHIT and p16 mRNA in biopsies of 52 patients with lung cancer who underwent diagnostic fiberoptic bronchoscopy between January 2010 and July 2011 at the Affiliated Hospital of Jiangsu University. All patients were newly diagnosed and had not received any form of therapy. The diagnosis of lung cancer was confirmed by one or more of the following approaches: cytology or histology from biopsy or surgery specimens. Tumors were classified as central or peripheral on the basis of CT and bronchoscopy findings. Peripheral lung cancer was defined as a tumor that had no endoscopic findings visible at bronchoscopy. The routine diagnostic procedure with bronchoscopy consisted of bronchial brushing (BB) and bronchial alveolar lavage (BAL) for cytological analysis, and bronchial biopsy or transbronchial pulmonary biopsy (TBPB) for histological examination. TBPB was performed under the guidance of x-ray television fluoroscopy. Bronchial biopsy and TBPB specimens from 19 patients with benign lung disease were included in this study as controls. All subjects gave written informed consent before the investigation. The study was approved by our hospital ethics committee.
At bronchoscopy, the normal lung was examined first, and then the bronchoscope was wedged into a segmental or subsegmental bronchus showing the mass, nodule, or infiltrative shadows on the chest CT in patients with peripheral lung cancer, or into the middle lobe in patients with diffuse lung disease. BAL fluid was obtained by instilling isotonic saline solution (three 40-mL aliquots) into the segmental or subsegmental bronchus and withdrawing fluid by medical suction. The fluid was centrifuged, and sediment was smeared on several slides and fixed with 95% ethanol. Then TBPBs were performed for these patients with peripheral lesions. When an endoscopically visible tumor was identified during bronchoscopy, forceps biopsy and BB were directed toward the endobronchial abnormality to obtain tissue and cytology samples. At least 5 biopsy samples were obtained from each patient by TBPB or endobronchial biopsy. Two or 3 biopsies were fixed in formalin solution. Another 3 biopsies were snap-frozen and stored at −80°C for molecular analysis. BBs were smeared onto slides and fixed in 95% alcohol and transported to the department of pathology with biopsies for cytological and histological assessment. The fact that the gene product was not detectable could be related to the absence of cells in the biopsies. Therefore, snap-frozen biopsies, 0.5-1 mm in diameter and stored at −80°C, were cut in a cryostat chilled to −30°C. A 5-μm slide was processed with toluidine blue stain to assess the tumor cells and tissue elements. If the slide contained tumor cells or bronchial and/or alveolar epithelial cells, the biopsy sample was considered to be qualified for the study.
RT-PCR analysis of FHIT and p16 mRNA deletion
RT-PCR was used to detect the deletion of FHIT and p16 mRNA. Total RNA was extracted from 50 mg frozen specimen using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. RNA was purified by precipitation with 500 μL of isopropanol. The RNA pellet was washed once in 70% ethanol, dried, and resuspended in 50 μL DEPC-treated RNase free ultra-pure water (DEPC H2O) and stored at −80°C. cDNA was synthesized using a RevertAid™ first-strand cDNA synthesis kit (Fermentas Co., Lithuania) according to the manufacturer's protocol. Total 2 μg RNA and 1 μL Oligo (dT)18 primer (Shanghai Invitrogen Biotechnology Co. Ltd, China) in 6 μL volume of DEPC H2O were heated to 70°C for 5 minutes and then chilled on ice for 5 minutes. The remainders of the reaction components were added to a final volume of 20 μL at a concentration of 10 mM dNTP (Promega Co., USA) and 1 μL (200 U/μL) of RevertAid™ M-MuLV reverse transcriptase in the provided buffer concentrate. The samples were incubated at 42°C for 60 minutes and 72°C for 10 minutes, then chilled on ice and stored at −20°C.
The cDNA was amplified using the MyCycler PCR system (Bio-Rad Co., USA), with the addition of 2.5 μL buffer, 1μL cDNA samples, 2.5 μL (2 mM) dNTP, 1.0 μL forward and reverse primers, respectively (10 μmol/L), 0.2 μL Taq polymerase (5 U/μL) and 15 μL dd H2O to each 25 μL reaction. Cycling conditions were as follows: initial denaturation at 95°C for 5 minutes, then 95°C for 30 seconds, annealing at 56-95°C for 1 minute, and finally extension at 72°C for 10 minutes. The number of amplification cycles was 35. As an internal reference, amplification of β-actin was performed with similar cycle optimization. The primer sequences and expected product sizes are listed in Table I. The PCR products were separated by polyacrylamide gel electrophoresis, stained with ethidium bromide, and visualized under UV illumination. The deletions of target gene transcripts in the study were FHIT exon 1-4 and exon 5-9, and p16 exon 1 and exon 2. Gene deletion was considered positive if there was at least 1 site transcript loss for every gene.
Sequences of Primers and Product Size of Amplification

Statistical analysis
The SPSS 13.0 software was used for statistical analysis. Wilcoxon's rank-sum test and chi-square test or Fisher's exact test were used for continuous and categorical variables in univariate analysis, respectively. Results with a p value <0.05 were considered statistically significant. Test sensitivity is equal to TP/TP+FN (TP is true positive, FN is false negative). Test specificity is equal to TN/TN+FP (TN is true negative, FP is false positive). Positive predictive value (PPV) is equal to TP/TP+FP. Negative predictive value (NPV) is equal to TN/TN+FN. Test accuracy is equal to TP+TN/TP+FP+TN+FN. When analyzing the diagnostic yield of a combination of molecular markers, a case was considered positive if at least 1 molecular marker was positive, and negative if 2 molecular markers were negative.
Results
Patient information
The clinicopathological characteristics of the 52 patients with lung cancer are shown in Table II. Tumor histology included 27 adenocarcinomas, 20 squamous cell carcinomas, 3 SCLCs, and 2 NSCLCs of unspecified histological type. The TNM stages of the tumors were as follows: stage IB, 3 patients; stage IIA, 8 patients; stage IIB, 7 patients; stage IIIA, 8 patients; stage IIIB, 14 patients; stage IV, 12 patients. In 30 (57.7%) lung cancer patients the diagnosis was made on the basis of the first diagnostic bronchoscopy. In the remaining 22 patients, the diagnosis of lung cancer was confirmed in 6 patients by transthoracic needle aspiration biopsy, in 2 patients with pleural effusion by pleural biopsy, and in 14 patients by thoracotomy. The group of patients with benign lung diseases comprised 8 cases of pulmonary tuberculosis, 5 cases of pneumonia, 5 cases of diffuse interstitial lung disease, and 1 case of sarcoidosis in the thorax. The mean ages of patients with lung cancer and patients with benign lung disease were 64.4±12.1 and 59.8±14.8 years, respectively. The male/female gender distributions in the 2 groups were 3.73:1 and 2.17:1.
Association Between Clinicopathological Characteristics of Patients and Gene Loss
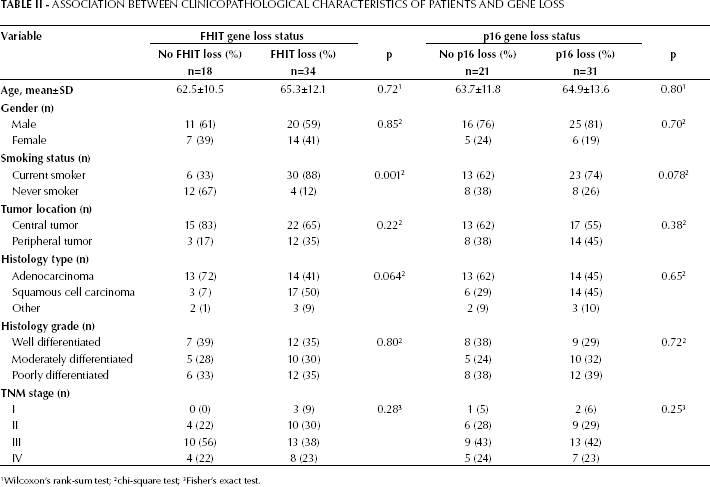
Wilcoxon's rank-sum test;
chi-square test;
Fisher's exact test.
Deletion rate of FHIT and p16 mRNA
Biopsies obtained by bronchoscopy from 52 lung cancer patients and 19 patients with benign lung disease were analyzed using RT-PCR. Figure 1 shows examples of samples investigated for detection of FHIT and p16 mRNA. Deletion of FHIT gene transcripts (exon 1-4 and/or exon 5-9) was detected in 34 biopsies from 52 lung cancer patients (65.4%) and in 2 biopsies from 19 patients with nonmalignant disease (10.5%). The nonmalignant lesions with FHIT gene deletion included 1 case of tuberculosis and 1 case of pneumonia. The detection rate of FHIT gene deletion was significantly higher in lung cancer than in nonmalignant lung disease (p=0.001). RT-PCR analysis of gene deletion from biopsies of lung cancer patients showed 32 losses (61.5%) of FHIT on exon 1-4, and 24 losses (46.2%) of FHIT on exon 5-9. In 22 specimens we detected simultaneous loss on both exon 1-4 and exon 5-9. No significant difference in deletion rate was observed between FHIT exon 1-4 and exon 5-9. In 30 patients with positive cytology and/or histology after the first bronchoscopy, 25 patients (83.3%) showed deletion of the FHIT gene. It is noteworthy that FHIT deletion was seen in 9 of 22 patients (40.9%) with a negative cytology and histology result. We found that FHIT deletion was more frequent in smokers than in nonsmokers (p=0.001). FHIT gene loss tended to be more frequently observed in squamous cell carcinoma than in adenocarcinoma, but the difference was not statistically significant (p=0.064). In addition, no association was found between FHIT gene loss and age, gender, tumor location, differentiation grade, or TNM stage (Tab. II).
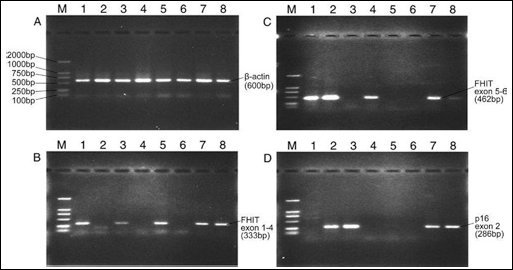
RT-PCR analysis for FHIT and p16 mRNA loss in biopsies obtained by bronchoscopy. Lane M, molecular weight marker. Lanes 1-7, specimens from the patients with lung cancer. Lane 8, specimen from a patient with benign lung disease. A) β-actin, an endogenous reference gene. B) Expression of FHIT exon 1-4 mRNA is observed in lanes 1, 3, 5, 7, 8. The absence of products is seen in lanes 2, 4, 6. C) Expression of FHIT exon 5-9 mRNA is observed in lanes 1, 2, 4, 7. The absence of products is seen in lanes 3, 5, 6, 8. D) Expression of p16 exon 2 mRNA is observed in lanes 2, 3, 7, 8. Note the absence of product in lanes 1, 4, 5, 6.
Deletions of p16 gene transcripts (exon 1 and/or 2) were detected in 31 biopsies from 52 patients with lung cancer (59.6%). p16 gene loss was found in a single biopsy sample from the 19 patients with benign lung disease (5.3%) and concerned a case of tuberculosis. There was a statistically significant difference between lung cancer patients and benign lung disease patients in the frequency of p16 gene loss (p<0001). In 31 samples with p16 gene loss, 30 samples had only exon 2 loss and 1 sample had concomitant p16 loss in both exon 2 and exon 1. p16 gene loss was identified in 23 (76.7%) of 30 lung cancer patients with positive cytology and/or histology and in 8 (36.4%) of 22 lung cancer patients with negative cytology and/or histology. No statistically significant difference in age, gender, smoking status, histological type and grade, or TNM stage was found between lung cancer patients with and those without p16 gene loss (Tab. II), although there was a tendency for p16 gene loss to be more frequent in smokers than in nonsmokers (p=0.078).
Diagnostic yields of detection of FHIT and p16 mRNA loss
Decision diagnostic criteria showed that molecular detection by RT-PCR analysis of FHIT gene loss from biopsies had 65.4% sensitivity, 89.5% specificity, 94.9% PPV, 48.6% NPV, and 71.8% test accuracy. For p16 gene loss detection of biopsy samples, we obtained 59.6% sensitivity, 94.7% specificity, 96.9% PPV, 46.2% NPV, and 69% accuracy. Combination of FHIT and p16 mRNA loss identified 80.8% (42/52) of patients diagnosed with lung cancer and 15.8% (3/19) of patients with benign lung disease. Therefore, a sensitivity of 80.8% with a specificity of 84.2% was achieved (Tab. III). These values compare favorably with those of cytology and histology combined, which demonstrated a sensitivity and specificity of 57.7% and 100%, respectively. Although the specificity was reduced to 84.2% by combining FHIT with p16 mRNA loss detection, test accuracy was raised to 83.1%.
Diagnostic Yields of Cytology, Histology and Detection of FHIT and P16 Loss
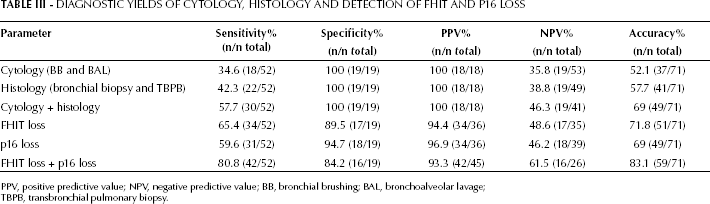
PPV, positive predictive value; NPV, negative predictive value; BB, bronchial brushing; BAL, bronchoalveolar lavage; TBPB, transbronchial pulmonary biopsy.
Our results showed that the sensitivities of cytology and histology by bronchoscopy were lower in patients with stage I-II than in those with stage III-IV lung cancer. By contrast, the diagnostic yield of FHIT and p16 loss detection was not influenced by TNM stage. The sensitivities of FHIT and p16 loss detection in patients with stage I-II lung cancer were similar to those in patients with stage III-IV lung cancer (Tab. IV).
Diagnostic Yields of Cytology, Histology, FHIT and P16 Gene Loss in Patients with Stage I-IV Lung Cancer
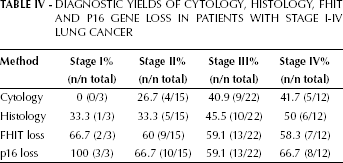
Discussion
A major obstacle to curative treatment of lung cancer is early-onset extrapulmonary dissemination. Only about 30% of patients with NSCLC are treated surgically, implying that the biological sample most frequently available for routine management at the time of diagnosis is biopsy. Biopsies by bronchoscopy are a suitable material because they are routinely performed in most patients suspected to have lung cancer. However, establishing a final diagnosis after the first bronchoscopy fails in about half of these patients (3). Thus there is a need to develop molecular biomarkers for early detection of lung cancer at a stage at which the disease is potentially reversible.
The objective of this study was to evaluate whether FHIT and p16 gene loss in biopsies are useful tumor markers to aid the diagnostic workup for suspected lung cancer. A number of studies have indicated that alterations of the FHIT and p16 genes play an important role in lung carcinogenesis. It has been suggested that alterations of FHIT transcripts are frequent events in lung cancer, and FHIT promoter methylation and loss of FHIT protein function are early events in the pathogenesis of lung cancer (10, 12, 18, 22). Aberrant FHIT mRNA transcripts have been detected in 40-80% of tumor samples (18), whereas mutations are rare. Loss of FHIT protein expression was observed in almost 70% of primary tumors, mainly those arising in smokers, and in 90% of preinvasive lesions (12).
The study by Wali et al showed that 73% of lung biopsies obtained by bronchoscopy from 30 patients with stage III lung cancer exhibited loss of FHIT mRNA expression, and loss of FHIT mRNA expression correlated with FHIT promoter methylation (31). In addition, there was a significant correlation between FHIT transcript and protein expression (31).
Our results show that loss of FHIT transcripts is frequently detected in biopsies by bronchoscopy from lung cancer patients but not in benign lung disease. The sensitivity and specificity of FHIT gene loss for the diagnosis of lung cancer were 65.4% and 89.5%, respectively. No significant difference in the rate of FHIT transcript loss between exon 1-4 (noncoding region) and exon 5-9 (coding region) was found. By contrast, Burke et al demonstrated that exon 5-8 was the most frequently deleted region of FHIT in lung cancer (32). The discrepancy may be attributable to primer design, but it should be noted that the sample sizes were different in both studies.
We also found that loss of FHIT mRNA expression appeared to be strongly associated with smoking status, suggesting that the FHIT gene may be the target of carcinogens in cigarette smoke. This finding is consistent with previous observations of a higher rate of FHIT expression loss in the lung cancers of smokers versus those of nonsmokers (12, 20). Sozzi et al reported that FRA3B is a preferential target of tobacco smoke damage at a molecular level (33). In agreement with previous studies (12, 19, 22), losses of FHIT expression were found in biopsies from all lung cancer stages. This suggests that the inactivation of the FHIT gene is involved in the early phase of lung carcinogenesis and hence is more likely related to the initiation of the neoplastic process than to the progression to invasive tumor and distant metastasis. Tseng et al demonstrated that FHIT expression is markedly reduced in a substantial number of early-stage NSCLC and preneoplastic lesions in chronic smokers, while they did not find a significant association between the lack of FHIT expression and survival in stage I NSCLC (34). Furthermore, Sozzi et al concluded that inactivation of the FHIT gene may happen at an early stage in lung carcinogenesis and could therefore be used as a biomarker. They identified loss of FHIT expression in 85% of dysplastic lesions and 100% of carcinoma in situ lesions at the margins of tumor specimens from patients with lung cancer (12).
Previous reports have described an increased frequency of FHIT expression loss in squamous cell carcinoma with respect to adenocarcinoma (12, 19, 22). In contrast, we did not observe a significant difference in loss of FHIT gene expression between squamous cell carcinoma and adenocarcinoma. Although the cause of this discrepancy is not clear, it may be related to the difference of the sample size and tumor stage between our findings and the results of other studies.
It was reported that the p16 gene is inactivated in up to 63% of NSCLC tumor specimens but rarely in SCLC (25). The inactivation of p16 is mainly caused by homozygous deletion, promoter hypermethylation, or mutation of the gene. Homozygous deletion of p16 can be found in 9-48% of NSCLCs (26, 35–38), whereas the p16 mutation rate is relatively low in lung cancer (38, 39). Studies have shown that aberrant methylation of p16 is an early event in lung carcinogenesis, occurring frequently in all stages of NSCLC, and could be a potential biomarker for early detection of this disease (27, 30). In this study, like FHIT gene loss, deletion of the p16 gene occurred significantly more often in biopsies from lung cancer patients than in biopsies from patients with benign lung disease. We demonstrated 59.6% sensitivity and 94.7% specificity by using the detection of p16 gene loss. It is also noted that loss of p16 mRNA expression appears to be associated with smoking status in lung cancer patients, although the difference in loss of p16 mRNA expression between smokers and nonsmokers did not reach statistical significance (p=0.078). Similar findings were reported by others (27, 28). It is not surprising to find that the changes in the FHIT and p16 genes are associated with smoking status, because lung cancer is characterized by an accumulation of molecular genetic abnormalities resulting from repeated exposure of the respiratory tract of chronic smokers to tobacco-related carcinogens (40). In addition, this study showed that loss of p16 did not correlate with age, gender, tumor location, histology type and grade, or tumor stage. Thus, our results are in line with the role of abnormalities of the p16 gene at a relatively early stage of pulmonary neoplasia.
Although the sensitivity of the 2 biomarkers investigated in this study was moderate, with frequencies of 59.6% and 65.4% in lung cancer patients, combining the 2 markers increased the detection rate to 80.8%. This is superior to cytology and histology from the initial bronchoscopy. Another important observation was that in 22 patients with negative cytology and/or histology, loss of the FHIT and p16 gene was detected in 44.9% (9/22) and 36.4% (8/22), respectively. The results may be explained by the fact that absence of these cancer suppressor genes was detectable in bronchial dysplasia, a precancerous condition, and in carcinoma in situ at the margins of tumor specimens from lung cancer patients (12, 34).
A limitation of the present study is the method used, which is time-consuming and relatively expensive. The gold standard for lung cancer diagnosis remains cytological or histological demonstration of pulmonary malignancy, and tumor markers can, at best, suggest the diagnosis.
In conclusion, our study suggests that, in addition to cytological and histological assessment, the detection of FHIT and p16 mRNA deletion in biopsies obtained during bronchoscopy represents a potential complementary tool for the diagnosis of lung cancer. Although the value of the markers would not be used to confirm the diagnosis of malignancy, molecular analysis can serve as an important adjunct to bronchoscopy in lung cancer diagnosis.
