Abstract
Cyclin D1 plays a key role in cell cycle control, particularly in the transition from G1 to S phase, regulated by cyclin-dependent kinases. The objective of the present study was to screen the cyclin D1 gene (CCND1) for polymorphisms in patients with head and neck cancer (HNC). Genomic DNA was isolated from blood samples of 380 HNC patients and 350 controls. In a hospital-based case-control study using the PCR-SSCP technique we found 3 novel germline mutations: g3578C>A, g3475G>C and g3383delA. The commonly reported guanine to adenine polymorphisms in exon 4 g7656G>A (rs9344) and g10861C>A (rs7177) in 3′UTR of CCND1 were also observed. The calculated frequencies of the g7656G>A (rs9344) polymorphism in GG, GA and AA genotypes were 27.3%, 38.6%, and 33.9% in HNC cases, and 44.2%, 29.4%, and 26.2% in normal healthy controls, respectively. Adjusted by age (in years), sex and smoking status, multivariate logistic regression analysis showed that the AA and GA genotypes were associated with a significantly increased risk (OR 1.34, 95% CI 1.03-1.64, p=0.028) for HNC. The CCND1 AA genotype variant was associated with an increased risk in individuals who were <40 years old (OR 1.45, 95% CI 1.02-2.08, p=0.04). In conclusion, it is suggested that the CCND1 G/A polymorphism is associated with the early onset of HNC and may contribute to HNC susceptibility in a Pakistani population.
Introduction
The CCND1 gene is located on chromosome 11q13, encodes the cyclin D1 protein and is expressed in response to mitogenic signals promoting transition through the restriction point in the G1 phase of the cell cycle (1). Cyclin D1 is a key regulator of the G1 phase of cell cycle progression, which acts by making a complex with the cyclin-dependent kinases CDK4/6 and promotes phosphorylation and inactivation of retinoblastoma protein (2). Aberrations of CCND1, such as chromosome amplification, translocations and inversions, have commonly been observed in different human cancers (3).
Studies suggest that a common single nucleotide polymorphism (SNP) in the CCND1 gene in the exon 4 boundary ([G870A] [rs9344]) creates an alternative splice site, encoding a protein with an altered C-terminal domain that lacks a phosphorylation site in the destruction box critical for ubiquitin-mediated proteolysis (4, 5). The alternatively spliced transcript increases the stability or half-life of the protein. It has been proposed that DNA-damaged cells in individuals with the A allele may bypass the G1/S checkpoint, resulting in the accumulation of an increased proportion of cells with DNA damage and genetic alterations (6).
A number of studies have linked the CCND1 G870A SNP to increased cancer risk (7, 8) but the evidence is not entirely consistent (9) and some controversial results exist regarding the effects on head and neck cancer (HNC) development (10-12). We have evaluated links between the CCND1 G870A polymorphism and susceptibility to HNC in a Pakistani population. The primary objective of the current study was to test the hypothesis that CCND1 germline polymorphisms are associated with an increased susceptibility to HNC.
Materials and Methods
A total of 380 newly-diagnosed patients with HNC (oral cavity, pharynx and larynx) were recruited from the National Oncology and Radiotherapy Institute (NORI), the Pakistan Institute of Medical Sciences (PIMS) and the Military Hospital Rawalpindi (MH). The study was conducted with prior approval from the ethical committees of the hospitals and the COMSATS Institute of Information Technology, Islamabad. A total of 350 healthy individuals matched for age, gender and ethnicity were selected as controls. Their inclusion criterion was absence of a prior history of cancer or precancerous lesions. Patients and controls suffering from any other familial disease (diabetes, hypertension and cardiovascular impairment) were excluded from the study. After giving their informed consent, all individuals were interviewed using a specifically designed questionnaire. Information on age, gender and ethnic group and data regarding smoking status were recorded.
DNA extraction and polymerase chain reaction (PCR)
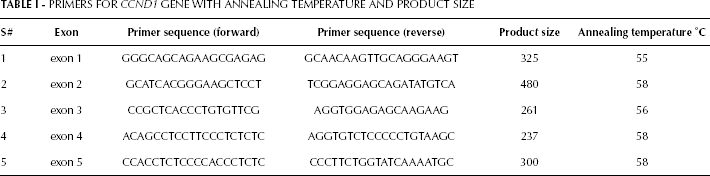
Blood from the subjects was sampled before the start of therapy. Samples were collected in EDTA-containing tubes and stored at 4°C. DNA was extracted from white blood cells using the standard phenolchloroform extraction method (13, 14). 5-ng dilutions were prepared of each isolated DNA and stored at −20°C for further processing. The human CCND1 exon sequence was taken from the Ensembl Genome Browser. Primers were designed using Primer3 software and checked for specific amplification using NCBI Primer-BLAST. The entire coding region and the exonintron boundaries of an approximately 60-bp sequence of CCND1 were selected (Tab. I). Each PCR reaction was performed in a 20-μL reaction mixture containing approximately 20 ng of genomic DNA templates, 1 μL (10 mM) of each primer, 0.24 μL (25 mM) of dNTP, 2 μL (10x) PCR buffer and 0.2 μL (5 U/μL) of Taq polymerase. Amplification was carried out with an initial melting temperature of 94°C for 5 minutes, 35 cycles of 94°C for 45 seconds, annealing temperature for 1 minute and 72°C for 1 minute, and a final extension step of 72°C for 10 minutes and hold at 4°C. PCR products were electrophoresed on a 2% agarose gel along with 100-bp DNA ladder and stained with ethidium bromide.
PRIMERS FOR CCND1 GENE WITH ANNEALING TEMPERATURE AND PRODUCT SIZE
Mutational screening and DNA sequencing
PCR product was analyzed by single-strand conformational polymorphism (SSCP) using the procedure described by Sheen et al (15) with some minor modifications. SSCP results were analyzed for band shifts for the identification of variants, with a gel documentation system (BioDocAnalyze by Biometra). Sequencing was carried out by the MCLab sequencing services.
Statistical analysis
Statistical analysis was done using the chi-square test with Fisher's exact test by means of the GraphPad Prism 5 Demo and SPSS. Odds ratios (OR) and 95% confidence intervals (CI) of genotypes were calculated with a multivariate logistic regression model. Bioinformatic analysis for CCND1 protein structural modeling was performed using the ESyPred3D and UCSF Chimera 1.5.3 programs.
Results
Germline mutation detection
The CCND1 gene coding regions were successfully analyzed for germline mutations. The combination of SSCP and sequence analysis revealed a novel missense and a frameshift mutation in exon 3 (g3475G>C and g3383delA), with a frequency of 0.07% and 0.06% respectively. In exon 3 a novel intronic missense mutation (g3578C>A) was also observed with a frequency of 0.09%. None of these mutations was observed in controls. A silent mutation, G870A (substitution of G A; [g7656G>A] [rs9344]) was observed in 129 patients on the exon 4 splice site, with a frequency of 0.33%. Control samples showed this polymorphism with a frequency of 0.26% (OR 2.39, 95% CI 1.58-3.63; p<0.03). In addition, g10861C>A (rs7177), a C G transition, was observed in the 3' untranslated region (UTR) of CCND1 in 74 patients with a frequency of 0.19%. Control samples harbored this polymorphism with a frequency of 0.07% (OR 1.719, 95% CI 0.903-3.273; p<0.05) (Fig. 1).
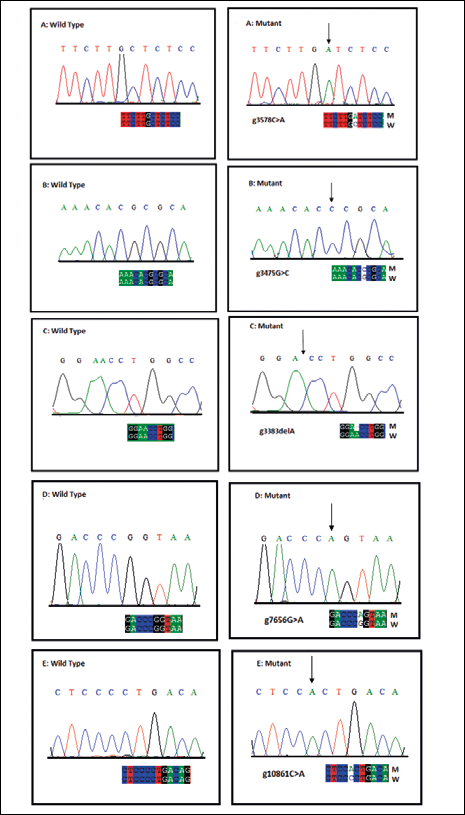
Sequencing electropherogram of mutations in the CCND1gene. The figure shows the wild-type sequence (left) and the mutant sequence (right). A) Missense mutation in exon 3 (g3578C>A). B) Missense mutation in exon 3 (g3475G>C). C) Frameshift mutation in exon 3 (g3383delA). D) Silent mutation in exon 4 (g7656G>A)(rs9344). E) Mutation in 3'UTR (g10861C>A) (rs7177).
Association of germline mutations with other parameters
Regarding the frequency of the missense mutation g3475G>C, no statistically significant difference was observed between men and women (OR 1.72, 95% CI 0.65-4.87; p=0.43). No difference was observed either with regard to smoking status (OR 0.36, 95% CI 0.12-1.07; p=0.11). The G to C substitution was significantly more frequent in patients with cancer of the oral cavity than in patients with cancer of the pharynx and larynx (p=0.05). The frameshift mutation g3383delA was significantly more frequent in patients with no smoking history (OR 4.75, 95% CI 1.24-18.59; p<0.028). The difference in the frequency of this mutation was statistically nonsignificant for gender (OR 2.28, 95% CI 0.79-6.59; p=0.197), age (OR 0.91, 95% CI 0.28-2.95; p=1.00) and tumor site (p=0.36). Regarding the 3′UTR mutation g10861C>A (rs7177), no statistically significant difference in frequency was observed between men and women (OR 0.85, 95% CI 0.44-1.62; p=0.74), between smokers and nonsmokers (p=0.40) and between patients with different tumor sites (p=0.30). A significant difference was observed between patients aged <40 years (OR 2.08, 95% CI 1.02-4.059; p<0.04) and those over 40 (Tab. II).
FREQUENCY OF MUTATIONS OF CCND1 AND RISK OF HEAD AND NECK CANCER
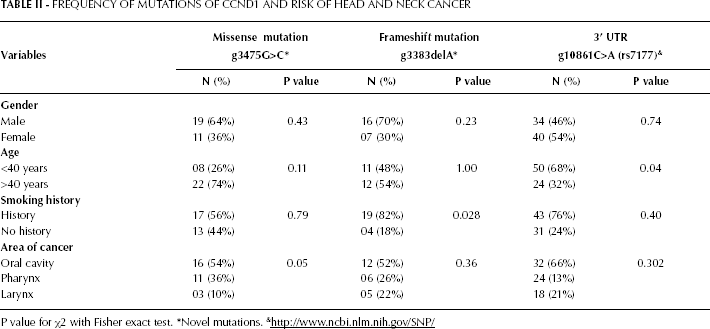
P value for χ2 with Fisher exact test.
Novel mutations.
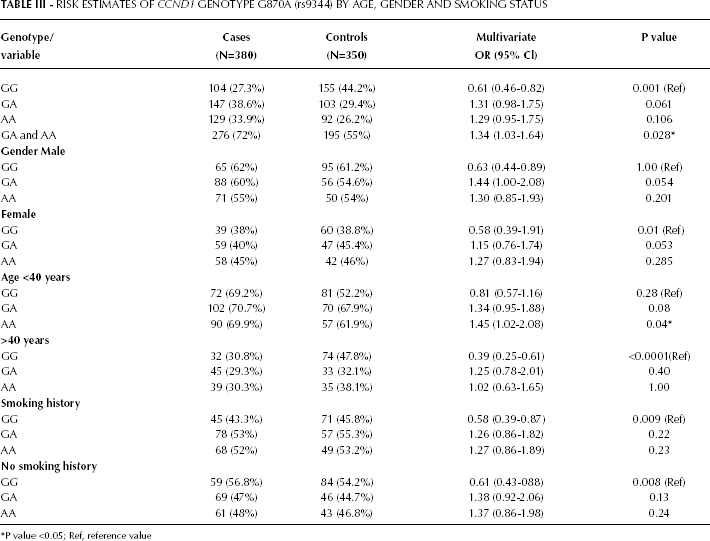
To evaluate the association between the G870A polymorphism and the risk of HNC (Fig. 2), ORs and their 95% CIs were calculated. The GG genotype frequency in cases versus controls was 27.3% and 44.2%, respectively. Both GA and AA were more common in cases than controls (38.6% versus 29.4%; p=0.061; and 33.9% versus 26.2%; p=0.106, respectively). After adjusting for age, gender, and smoking status, compared to individuals with the wild-type GG genotype, individuals with the GA and AA genotypes showed significantly increased cancer risks with adjusted ORs of 1.31 (95% CI, 0.98-1.75) and 1.29 (95% CI, 0.95-1.75), respectively. Combining the heterozygous GA and homozygous variant AA genotypes generated an OR of 1.34 (95% CI 1.03-1.64; p=0.028). When further stratified analyses were performed, the increased risk conferred by the AA genotype was more evident in younger individuals (<40 years) (OR 1.45, 95% CI 1.02-2.08; p=0.04) (Tab. III).
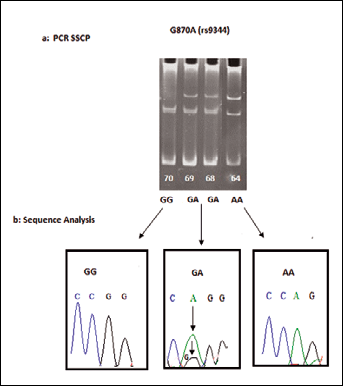
a) PCR-SSCP typing pattern of G870A (rs9344) polymorphism of CCND1 genotypes: GG homozygous wild type; GA heterozygous wild type; AA homozygous mutant. b) Sequence analysis.
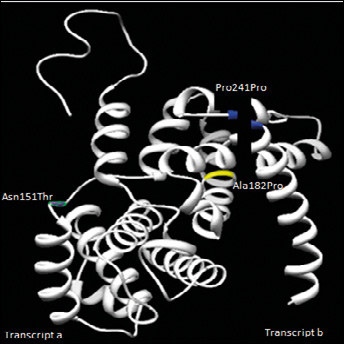
CCND1 protein structure indicating transcript a and transcript b. The mutations Asn151Thr, Ala182Pro and Pro241Pro are represented in the structure, which was obtained by means of the UCSF Chimera 1.5.3 and ESyPred3D programs.
RISK ESTIMATES OF CCND1 GENOTYPE G870A (rs9344) BY AGE, GENDER AND SMOKING STATUS
P value <0.05; Ref, reference value
Discussion
The current study was designed to show germline analysis in the CCND1 gene, supporting the hypothesis that polymorphisms are a susceptibility factor in the molecular progression of HNC. This cancer site is largely associated with environmental factors, especially in individuals with prolonged exposure to tobacco and alcohol (16), and genetic factors, including deregulation of the cell cycle (17). CCND1 has been suggested to play an important role in tumorigenesis and progression of HNC (18).
In this study a novel frameshift and 2 missense mutations were found in exon 3. Frameshift mutations in the coding region change the whole downstream sequence of a gene. Translation of the wrong reading frame is propagated and the resultant premature RNA stability is compromised, so either the mRNA will not be stable or protein degradation will occur (19). Missense mutations in the coding region result in amino acid substitution in the gene, and since the substitute and the original residues differ in chemical properties (polarity, hydrophobicity, etc); mutations might have a substantial effect on the conformation and consequently the functional activity of the gene (20).
The present study also shows that a single base polymorphism in exon 4 (G870A) is associated with a relatively increased risk of HNC in younger individuals. This particular polymorphism, a G to A mutation located at codon 241 on exon 4 in a splice donor site, additionally enhances CCND1's alternate splice capability (21). Several previous studies supported our data and reported associations between the CCND1 G870A SNP and increased risk for a number of solid tumors, including cancers of the colon, lung, skin, head and neck, prostate, bladder, kidney and esophagus (22, 23). Studies observed that the CCND1 G870A variant allele was a risk factor for squamous cell carcinoma of the upper aerodigestive tract in nonalcoholics and nonsmokers (17) and the risk was markedly increased in younger female individuals (24).
Our results are not consistent with a meta-analysis which strongly suggested that the CCND1 G870A polymorphism is not associated with HNC risk. The meta-analysis included 11 studies of different European and Asian populations (25). Although some of the studies found the G870A polymorphism to be associated with an increased risk of HNC (24, 26, 27), after pooling of the results no significant association was found between this polymorphism and HNC risk.
Our study links the G870A polymorphism and others to an increased risk of HNC in a Pakistani population. There could be many possible reasons for the lack of an association between G870A polymorphism and risk of HNC, of which at least 2 need to be addressed, namely study heterogeneity due to population characteristics and the small sample size, as mentioned by Tang et al (25).
Figure 3 shows in silico structural model analysis of the CCND1 protein using the UCSF Chimera 1.5.3 and ESyPred3D programs. The CCND1 870A allele resulted in an alternatively spliced transcript of CCND1 known as transcript b, which lacks the PEST motif containing exon 5. The PEST motif is critical for the degradation of cyclin D1; thus transcript b (A allele) was shown to have a longer half-life than the transcript a (G allele) encoded protein. This suggests that individuals with more copies of CCND1 870A are more likely to bypass the G1-S checkpoint, which contributes to cancer development (21) and might be a useful prognostic factor for oral squamous cell carcinoma (28). Similarly, lower expression of cyclin D1 in neoplastic cells has also been correlated with a higher percentage of disease-free survival, which is indicative of the prognostic value of cyclin D (29).
We also report a known polymorphism in the 3′UTR. Although the mechanism of the C>A (rs7177) polymorphism regulating the CCNDI gene is unknown, mutations in the 3′UTR of CCND1 were associated with increased stability of its mRNA and cyclin D1 overexpression. As suggested by earlier studies, rearrangement of the 3′UTR of CCND1 may contribute to its activation in some lymphoid tumors. Moreover, the finding that the half-life of CCND1 mRNA was >3 hours in cell lines (while it is only 0.5 hour in normal tissues) shows the importance of posttranscription derangement in the activation of CCND1 (30).
In conclusion, our results suggest that the CCND1 G870A allele (rs9344) and 3′UTR (rs7177) SNPs may be independently or jointly involved in the early carcinogenesis of HNC in a Pakistani population.
Footnotes
ACKNOWLEDGMENTS
All authors would like to acknowledge the participants who contributed to this research and the hospital staff at the National Oncology and Radiotherapy Institute (NORI) and the Pakistan Institute of Medical Sciences (PIMS) for their cooperation. The authors would also like to acknowledge unwavering moral and emotional support from family and friends, especially Kashif Ali and Uzma Inayat for helpful discussions and M.W. Hussain (Research Associate, Shaukat Khanum Cancer Hospital and Research Centre) for guidance in learning basic molecular biology techniques. The authors would like to express their gratitude to Muhammad Qasim Khan for performing bioinformatic analysis.
