Abstract
This mini-review will schematically update the progress of the expanding Mi-2/Nucleosome Remodeling Deacetylase (NuRD) complexes in cancer and in normal development such as stemness, with a focus on mammals and the increasingly popular and powerful model organism Caenorhabditis elegans. The Mi-2/NuRD complexes control gene activity during the development of complex organisms. Every Mi-2/NuRD complex contains many different core polypeptides, which form distinct multifunctional complexes with specific context-dependent regulators. The Mi-2/NuRD complexes have unique ATP-dependent chromatin remodeling, histone deacetylase, demethylase activities and higher order chromatin organization. They can regulate the accessibility of transcription factors or repair proteins to DNA. In this review, we summarize our current knowleges in the composition, interaction and function of the subunits within the Mi-2/NuRD complex, the methodology used for the identification of Mi-2/NuRD complexes, as well as the clinical and therapeutic implications targeting the Mi-2/NuRD subunits.
Overview
To gain an insight into the functions of the Mi-2/ NuRD complex, we first need to understand the composition of this complexes. The Mi-2/NuRD complex is a expanding family of protein complexes, which control gene activity spatiotemorally during the development of complex organisms. In general, it has been regarded as one biochemical entity containing a number of core polypeptides but forming distinct multifunctional complexes with specific context-dependent regulators.1,2 Some of these complexes are active during normal development such as stemness3–7 and in cancer8,9 (Table 1). These complexes are characterized by their unique ATP-dependent chromatin remodeling, histone deacetylase, demethylase activities and higher-order chromatin organization.2,24 Nucleosomes form the building blocks of chromatin and, in general, these nucleosomes inhibit processes that require access to the DNA template, such as transcription and DNA repair. Mi-2/NuRDs use ATP hydrolysis to change the nucleosomes in DNA and they can regulate the accessibility of transcription factors or repair proteins to DNA. 25 The two highly homologous proteins Mi-2α/CHD3 and Mi-2β/CHD4 represent the catalytic ATP hydrolyzing subunits in the complex. The histone lysine-specific demethylase (LSD1/KDM1) was recently shown to be recruited to the NuRD complex via interaction between the Tower domain and metastasis-associated proteins MTA1-3 in breast cancer cells. 8 This newly identified subunit demethylates both di- and mono-methylated K4 (H3K4me2/1). 26 In addition, the core of the Mi-2/NuRD complex contains:
at least two histone deacetylases, HDAC1 and HDAC2;
the retinoblastoma-associated tumor suppressors RbAp48 (RBBP4) and/or RbAp46 (RBBP7);
MTA1 and/or 2 and/or 3;
GATAD2b (p66α) and/or GATAD2a (p66β);
mCpG-binding domain proteins MBD2 and/or MBD3. 27
The neoplasms associated with core subunits of the Mi-2/NuRD compelx.
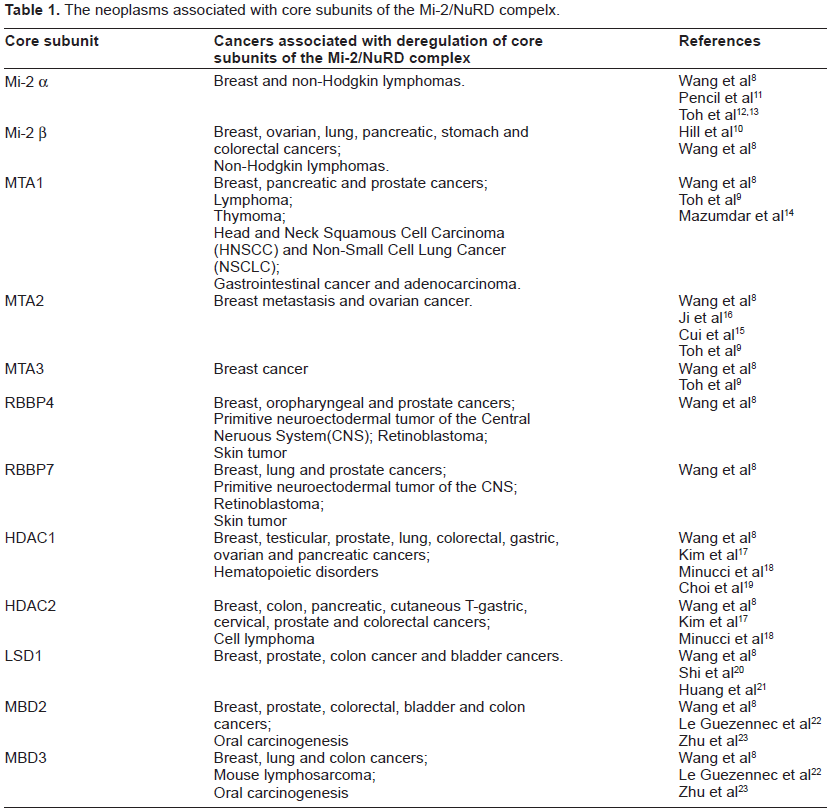
Importantly, the stemmed Mi-2/NuRD complex emerges from these core components. The gene targeting mechanism of MBD2-recruitting Mi-2/NuRD (ie, the MeCP1 complex) 28 and the gene targeting mechanism of the transcriptional factor-interacting M-2/NuRD complex (eg, the lymphoid transcription factor Ikaros, 29 the transcriptional corepressor KAP-1, 30 the tumor suppressor p5331 and Bcl-6)32,33 have been determined as the interactions that are mediated by Mi-2α (Ikaros and KAP-1), MTA-2 (p53) or MTA-3 (Bcl-6), and therefore different transcriptional regulators use distinct NuRD subunits for recruitment. However, the details of the mechanisms of many other co-regulators (Supplementary Table 1) interacting with Mi-2/NuRD complexes remain largely unknown. The distinct tissue- and cell-specific Mi-2/NuRD complexes can execute their multifunctional, fine and spatiotemporal regulation in a variety of fundamental biological processes. In principle, this diversity could challenge the capture of all or most interactors from one purification experiment using core subunits of the Mi-2/NuRD complex tagged with a single condition; on the other hand, for cancer, cell-specific transcriptional factors and co-regulators of the Mi-2/NuRD complexes offer unexpected therapeutical benefits for designing both systemic and local site treatments (eg, RNAi therapeutics) in the future and may enable different therapeutic strategies tailored to patients with different profiles. These studies have benefitted from the comparative advantages of interspecies Mi-2/NuRD complexes.34,35 Finally, increasing our knowledge of the Mi-2/NuRD complex could significantly advance cancer therapy.
The Interacting Proteins and Functions of the Core Subunits within the Mi-2/NuRD Complex
The Mi-2α, β/CHD3, 4 proteins
The evolutionarily conserved chromatin remodeling factor Mi-2 protein was first identified as an autoantigen in patients with dermatomyositis36,37 and belongs to the chromodomian CHD3-CHD4 ATPase subfamiliy (CHDs) II. Mi-2 has been isolated from human cell lines, mice, plant, fly, worm and Xenopus egg extracts27,36–43 (Fig. 1). The human genome contains two genes encoding isoforms of Mi-2: Mi-2α/CHD-3 and Mi-2β/CHD-4. 44
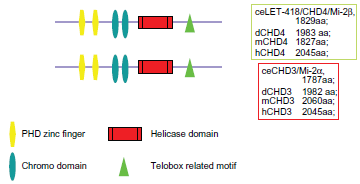
The structural domains of Mi-2α,β/CHD3,4 proteins.
Mi-2 proteins contain several domains: two PHD zinc fingers, two chromodomains (chromatin organization modifier), a SWI2/SNF-like helicase domain and a telobox-related motif 43 (Fig. 1a). PHD zinc finger domains are thought to be involved in protein-protein interactions. In fact, the Mi-2 PHD domains were shown to be required for Mi-2's interaction with HDAC1. 43 At least part of the nucleosomal recognition is contributed by Mi-2 itself, since the ATPase activity of the recombinant enzyme is strongly stimulated by nucleosomes, but does not react to the presence of free histones or DNA. 45 Mi-2 and its complex regulate gene expression by modifying chromatin accessibility during development. The Mi-2 subunits are implicated in a wide variety of cancers (Table 1). Mi-2β acts as a transcriptional repressor and as an activator, which is required for hematopoietic stem cell self-renewal and differentiation.46,47 The Mi-2/NuRD complex has significant roles in DNA damage and repair.25,35,48–50 TWIST1 interacts with several components of the Mi-2/NuRD complex, Mi-2β, RbAp46, MTA2 and HDAC2, and recruits them to the proximal regions of the E-cadherin promoter for transcriptional repression. Depletion of these TWIST1 complex components from cancer cell lines, which depend on TWIST1 for metastasis, efficiently suppresses cell migration and invasion in culture and lung metastasis in mice. 51
Previous reports showed that Mi-2α and Mi-2β can coexist within the same complex.39,42 So far, it remains unknown whether these Mi-2α and Mi-2β are forming heterodimers or whether they are assembled into distinct complexes.
But it is likely that all 3 types of complexes exist like ie, Mi-2α/NuRD, 30 Mi-2αβ/NuRD and Mi-2αβ/ NuRD. Furthermore, though the increasing number of published interactors of Mi-2 proteins show up (Fig. 2; supplementary Fig. 1), we are in await for a systematic screening of interactors of Mi-2 proteins via current proteomic assay, which has evolved and matured to level where it is able to assess the complexity of the human proteome. 52
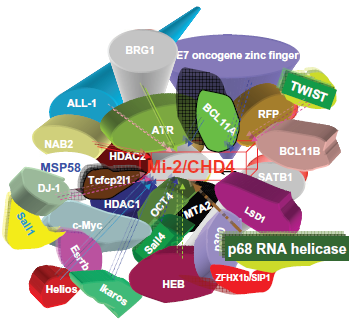
The interactors of mammalian Mi-2β/CHD4.
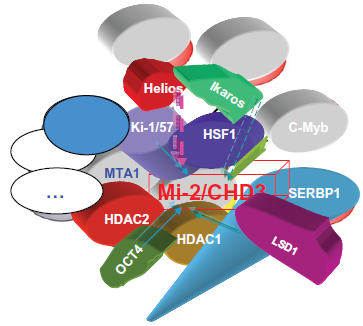
The interactors of mammalian Mi-2α/CHD-3.
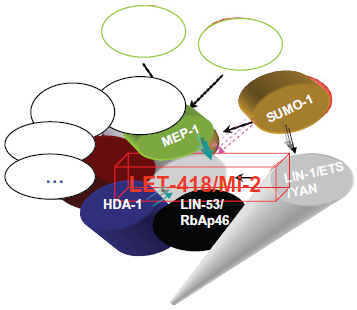
The interactors of C. elegans Mi-2β/LET-418.
Two Caenorhabditis elegans Mi-2 orthologs, LET-418 and CHD-3, have been cloned and characterized. 53 let-418 has proven to be an essential gene expressed in most, if not all, nuclei of the worm. Mutations in let-418 have pleiotropic phenotypes, including vulval defects and sterility and, without maternal contribution, result in L1 larval arrest. 53
Furthermore, let-418 is required for the maintenance of somatic differentiation in C. elegans. 54 By contrast, mutations in chd-3 do not display any obvious phenotype under standard culture. However, a requirement for chd-3 becomes apparent in let-418(lf); chd-3(lf) double mutants, which display strong vulval defects and L4 arrest or, without any let-418 maternal contribution, embryonic arrest. These results suggest that let-418 and chd-3 have essential and partially redundant functions during development. 53 Previously, we clarified the functions of LET-418/Mi-2β using genetic and biochemical techniques including Chromatin Immuno-Precipitation (ChIP), quantitative real time reverse transcriptase polymerase chain reaction (RT-PCR), co-immunopreciptation, reporter assays and suppression subtractive hybridization (SSH); we demonstrated that LET-418/Mi-2β along with LIN-1/ ETS directly binds the promoter of the lin-39/Hox gene to repress its expression 55 and LET-418/Mi-2β directly regulates LAG-2/Delta in the LIN-12/Notch pathway. 56 LIN-39/Hox itself performs an auto-regulatory function as well as directly controlling EGL-17/ FGF5 and LAG-2/Delta55,57,58 (Zhang and Mueller, unpublished; Fig. 3). The role of regulation of Mi-2/ NuRD in the LIN-12/Notch pathway has been demonstrated in different labs.53,59,60 Furthermore, spr-5 encodes the C. elegans LSD1 homolog and is implicate in Notch signaling.61–64 Mittal et al recently observed high-level expression of Notch receptors and ligands, and its increased activation in several human breast cancers and early precursors. This places Notch signaling as a key player in breast cancer pathogenesis. Notch signaling cooperates with the Ras/MAPK pathway in transformation and it offers combined inhibition of the two pathways as a new modality for breast cancer treatment. 65 The cross-talk and precise molecular cooperation between EGF/RAS and LIN-12/Notch cell signalling pathways have been clearly shown in C. elegans vulva development. 66
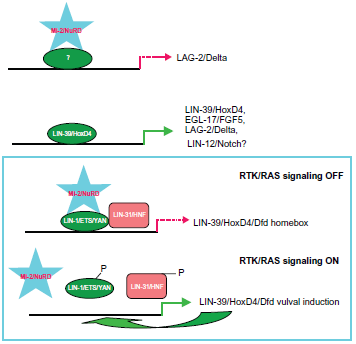
The downstream target genes of the Mi-2/NuRD complex in C. elegans.
The mCpG-binding domain (MBD) proteins
MBD2/Mi-2/NuRD and MBD3/Mi-2/NuRD
As Figure 4 shows, MBD proteins have the mCpG-binding domain. MBD3's MBD domain has two point mutations that abolish the binding. MBD1, MBD2 and the methyl CpG binding protein 2 (MeCP2) have the Transcriptional Repressive Domain (TRD). MBD2 and MBD3 can form both homo- and heterodimers because both have the Coil-Coil (CC) domain. MBD4 is involved in DNA repair. Nearly all cases of Rett Syndrome are caused by a mutation in the MeCP2. The MBDs bind sites of hypermethylation in human cancer cell lines. MBDs function in transcriptional repression and long-range interactions in chromatin, and also appear to play a role in genomic stability, neural signaling 80 and transcriptional activation. 81 MBD3 was shown to be an integral component of the Mi-2/NuRD complex, 82 whereas murine MBD2, which can bind methylated DNA, 83 targets the Mi-2/NuRD to methylated CpG dinucleotides.82,84 In general, MBD2 and MBD3 have been reported in a common Mi-2/NuRD complex, but different sets of inertactive proteins of MBD2 or MBD3 have been reported (Fig. 5). MBD2/Mi-2/NuRD complex (ie, the MeCP1 complex) was shown to be present at very early stages of development and to inactivate methylated promoters in the mouse embryo prior to implantation. 85 However, in MBD2/MBD3 tagged complex purification, in oral carcinogenesis and colon cancer, the MBD2 eluate but not that of MBD3 contained the arginine methyltransferase PRMT5, MEP50 and importin α nuclear transport proteins. The association between MBD2 and importins may indicate that MBD2 shuttles between the cytoplasm and the nucleus. The over-expression of DOC-1 identified in purification results in a G1 arrest and significant growth retardation compared to wild-type cells, which is consistent with loss of the protein in tumors. A proposed model is shown in Figure 6c,d: the MBD2/NuRD complex deacetylates the nucleosomes surrounding the targeting site;the addition of transcriptional repressive arginine methyl marks the H4 tail by its associated PRMT5. The hypoacetylated and arginine methylated nucleosomes surrounding the MBD2/ PRMT5 targeting site in turn may provide a binding scaffold for the MBD3/NuRD complex. The MBD2/ NuRD and MBD3/NuRD complexes co-occur on some CpG islands. The MBD3/NuRD complex can further deacetylate the nucleosomes and then facilitate the spreading of deacetylation and maintenance of transcriptional repression. 22 Unlike the other MBD members, MBD3 is unable to bind methylated DNA Human MBD3 and HDAC1 are localized at Aurora-A-positive centrosomes in the M phase. 86 MBD3 is transiently phosphorylated during the late G2 and early M phase, and Aurora-A is the candidate kinase for its phosphorylation.
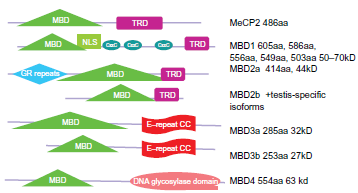
The structural domains of the mCpG-binding domain (MBD) proteins.
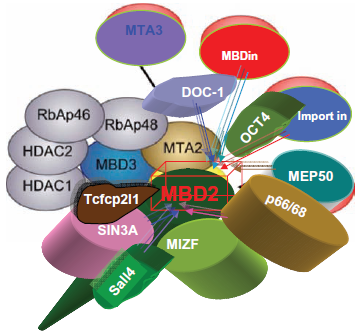
The interactors of mammalian MBD2.
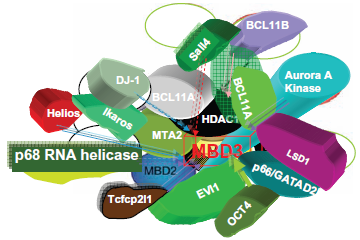
The interactors of mammalian MBD3.
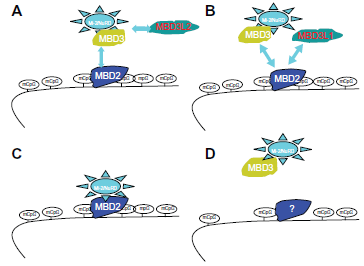
Summary of MBD/Mi-2/NuRD complex models.
In mice and humans, two proteins with high homology to MBD3 were identified: MBD3L1 and MBD3L2 (methyl-CpG-binding protein 3-like 1 and 2), which lack the MBD domain. 87 MBD3L1 can enhance MBD2-mediated repression in CpG-methylated promoters. MBD3L1 and MBD3 were shown to bind to overlapping regions of MBD2. MBD2 may interact with MBD3 or with MBD3L1 to form two different complexes with redundant function. 87 The Mi-2/NuRD complex may interact either with MBD2 or MBD3L2, via MBD3 (Fig. 3). MBD3L2 was shown to oppose MBD2/NuRD-mediated methylation silencing 88 by recruiting the complex away from methylated DNA and reactivating transcription. The hetero-oligomeric complexes can be formed between the four related proteins MBD2, MBD3, MBD3L1 and MBD3L2 and therefore different subforms of the Mi-2/NuRD complex (Fig. 6a,b).
Interestingly, the C. elegans genomic DNA is unmethylated, so C. elegans mbd-2 89 may exist in vestigial mammalian MBD3. The general role of MBD proteins in epigenetic regulation may be independent of methyl-DNA binding. Interestingly, while searching for the C. elegans homologs of components of the Mi-2/NuRD complex, we identified that CID-1, FLT-1 (now the C. elegans ACF1 homolog) adn R05D3.11 (now Met-2) have about 25% or better amino acid indentityl to the mammalian MBD domain, and knock-down (KD) animals with RNA interference against these genes had a low percentage of weak multivulvae, a phenotype of LET-418/Mi-2 deficiency. For the checkpoint protein CID-1, it remains tempting to know if it could transiently recruit Mi-2/NuRD complex in order to execute its function in longevity 90 (Zhang, unpublished).
The metastasis-associated (MTA) proteins
A decade ago, the function of the MTA (metastasis-associated gene) subunits of Mi-2/NuRD complex were largely unknown. At that time, MTA protein was a newly discovered family of cancer progression-related genes and their encoded products. The MTA gene family is clearly involved in oncogenesis and it is also used as an important marker of prognosis in cancer. The vertebrate MTA family of proteins is encoded by three genes that code for five isoforms: MTA1 and a splice variant known as MTA1s, MTA2, MTA3 and a longer isoform MTA3L, and the ZG29p (Fig. 7), an N-terminal truncated form of MTA1 that is present in the zymogen granules of the pancreas (Fig. 4a). MTA1 is the original member of a small family of metastasis-associated genes which also includes MTA2 and MTA3. The longer MTA1 isoform contains a bromo-adjacent homology (BAH) domain, an EGL-27 and MTA1 homology (ELM2) domain, a GATA-type zinc finger and a MYB domain. It contains four phosphorylation sites two serine and two threonine and two SRC homology-3 (SH3)-binding motifs. 93 MTA1s (short isoform) is missing the 285 AA C-terminal sequence and has a substituted 33 AA C-terminal compared to the long form. MTA1s contains an estrogen receptor binding motif (LRILL) not found in the long form. The distinct Mi-2/NuRD complexes are defined by different MTA variants with different composition and distinct functions that presumably target different sets of gene promoters.27,93 Presumably, MTA2 is expressed constitutively and may be involved in housekeeping functions of the Mi-2/NuRD complex, whereas MTA1 and MTA3 could be involved in cell type-specific transcription, since they are expressed in a tissue-specific manner. 27
MTA1/Mi-2/NuRD
Through differential cDNA screening, MTA1 was initially identified as being abundantly overexpressed in highly metastatic rat mammary adenocarcinomas.11–13 Elevated expression of MTA proteins has been associated with many types of metastatic cancers such as gastrointestinal and esophageal carcinomas, and mammary adenocarcinomas, along with increased tumor aggressiveness, invasiveness, metastasis and poor prognosis27,91–94 (Tables 1 and 2; Fig. 7).
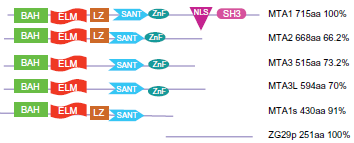
The structural domains of the metastasis-associated (MTA) family members. All major members can be a subunit of the Mi-2/NuRD complexes. The different role of each member of the family is still not clear but there is high similarity among them. The BAH domain is involved in protein-protein interaction, whereas the SANT domain binds to histone tails.
A short list of target genes of the core subunits of the Mi-2/NuRD complex.

Several studies have shown that these MTAs splicing variants form distinct protein complexes with Mi-2/NuRD components.42,43,82,93 As mentioned above, the MTA1 gene yields two isoforms (MTA1 and MTA1s (Fig. 9a, b)) via alternative mRNA splicing and one novel zymogen granule protein (ZG29p) via alternative transcription initiation.95,96 MTA1 is associated with the Mi-2/NuRD complex. 42 The splice variant MTA1s is cytoplasmic because it lacks a nuclear localization sequence. 27 ZG29p mediates an interaction with amylase and is involved in condensation-sorting in the exocrine rat pancreas, and it could possibly be linked to the deadly pancreatic cancer9,97 (Fig. 4a). MTA1s plays an important role in breast cancer malignancy by sequestering estrogen receptors in the cytoplasm. MTA1 could affect the transcription of disease-related genes via chromatin remodeling. However, Yao and Yang reported that MTA1 associates with specific NuRD complexes that contain HDAC1/2, RbAp46/48, and MBD3, but neither Sin3 nor Mi2. 93
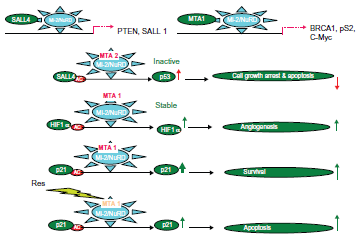
The downstream paths of the Mi-2/NuRD complex in carcinogenesis and cancer progression.
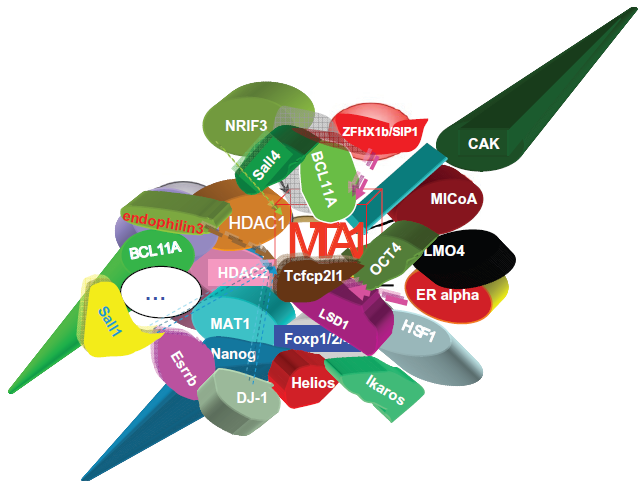

The interactors of mammalian MTA1.
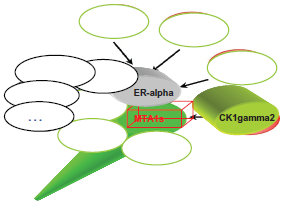
The interactors of mammalian MTA1s.
The NuRD-dependent mechanisms of tumorigenicity induced by MTA1 have been intensively studied by researchers such as Toh and Kumar. 9 MTA1 is involved in regulation of genes such as the estrogen receptor (ER-α), c-Myc, BRCA1, p53 and many more (Fig. 8, Table 2 and Supplementary Table 1). Indeed, MTA1 converts breast cancer cells to a more aggressive phenotype by repression of the estrogen receptor (ER-α) trans-activation function through deacetylation of the chromatin in the ER-responsive element of ER-responsive genes. Furthermore, MTA1 plays an essential role in c-MYC-mediated cell transformation. Overexpressed MTA1 (the long isoform) in specific NuRD complexes may modulate tumor aggressiveness via interference with estrogen signaling. Activation of the heregulin-beta1/HER2 pathway 14 of breast cancer cells induces MTA1-dependent repression of estrogen receptor transactivation, leading to enhanced anchorage-independent growth and hormone independence.96–101 Since MICoA and NRIF3 are MTA1-binding partners, their coactivators and corepressors may coexist in a single super complex. c-Myc upregulates MTA1 expression (Fig. 8) and activation of NuRD complexes containing MTA1. 102 Importantly, c-Myc is one of four induced pluripotent stem (iPS) cell factors and has been linked to the dedifferentiation of somatic cells to the iPS cell phenotype.103,104 MTA1/Mi-2/NuRD complexes have been shown to repress the BRCA1 tumor suppressor gene, and this resulted in an abnormal centrosome number, and chromosomal instability and inhibition of p53-induced apoptosis.105,106
MTA1s is upregulated by FGF-2 in breast cancer cells, and it sequesters ER-α in the cytoplasm and represses ER-α transcriptional activity, estrogen-induced proliferation and anchorage-independent growth of the human breast cancer cell line MCF-7. 107 In an animal model, MTA1s peptides block the tumor progression of MCF7 overexpressing ER-α, which promotes the malignant phenotypes. 108 MTA1s associates with casein kinase I-γ2, an estrogen-responsive kinase that phosphorylates and modulates the functions of MTA1s. The acetylated MTA1- histone deacetylase (HDAC) interaction facilitates the recruitment of the MTA1-HDAC complex to the Gαi2 regulatory element, and consequently for the repression of Gαi2 transcription and expression, which leads to activation of the Ras-Raf pathway. 78 MTA1 and MTA1s stimulate Wnt1 transcription through promoting its derepression from the Six3 corepressor. 109 Moreover, a hypoxia-inducible factor-1α (HIF-1α) is deacetylated and stabilized by MTA1, resulting in angiogenesis (Fig. 8). The HSF1, one interactor of MTA1 in the MCF7 cell line, has interactions with HIF-1α after stress (Li et al, unpublished). Thus, MTA1, and probably other MTA proteins, belong to the master co-regulatory molecules involved in carcinogenesis and the progression of various malignant tumors.
The ELM2 domain in vertebrate and invertebrate MTA1, -2 and -327,110 (Fig. 7) was first described in a C. elegans protein, EGL-27, which is a nuclear protein involved in embryonic patterning, cell polarity, cell migration and vulval development. 111 Based on sequence, EGL-27 is likely to be an ortholog of atrophin in the nematode C. elegans. Therefore, EGL-27 may be involved in nuclear receptor signaling in C. elegans. C. elegans has MTA1 homologs, egl-27 and egr-1, which are related to embryonic patterning. The Mi-2/NuRD complex including egr-1 antagonizes vulval development induced by the Ras signal transduction pathway, at least partly by promoting cell fusion between the vulval precursor cells and the hypodermal syncytium at an early larval stage. This inhibitory function of lin-40 might be carried out by downregulating lin-39 Hox expression112,113 (Fig. 3). Thus, understanding their physiological functions will be absolutely necessary to understand the pathological functions of MTA proteins in human cancers. However, one RNAi/ egr-1 study 60 reported a failure to detect ectopic expression of lag-2::gfp, but this could be caused by a lower efficiency of RNAi, especially with the genomic RNAi construct; it would be interesting to examine the ectopic expression of the lag-2::gfp phenotype in egr-1 mutant and egl-27 mutant backgrounds (Fig. 3).
MTA2/Mi-2/NuRD
MTA2, along with MTA1 and MTA3 and their isoforms, is a member of the MTA family of novel nuclear receptor coregulators. MTA2 (Fig. 9c) appears in Mi-2/NuRD complexes composed of HDAC1/2, RbAp46/48, MBD3, Sin3 and Mi-2, and is associated with maintaining homeostasis of the cell. MTA2 is involved in the regulation of NuRD complexes, where it modulates the activity of the core histone deacetylase (HDAC) complex. 82
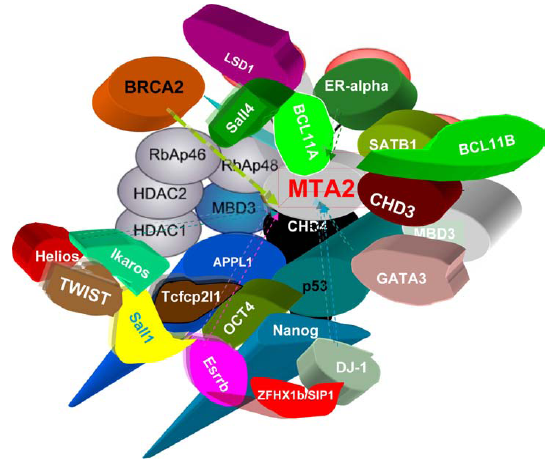
The interactors of mammalian MTA2.
MTA2 is in many cancers including breast cancer and epithelial ovarian cancer 15 but it is not correlated as strongly with metastasis as MTA1. MTA1 and MTA2 both exert histone deacetylase and gene repressor activity. MTA2 (but not MTA1) forms complexes with the transcription factors YY1 and FKBP25. 93 YY1 is required for development. The MBD domain of MBD3 is necessary and sufficient for binding to HDAC1 and MTA2. 114
MTA2 has normal physiological roles. In experiments with MTA2 null mice, MTA2 was shown to be important for embryonic survival and to be involved in modulating IL-4 and IFN-gamma expression in T-cell immune responses. These null mice develop lupus-like autoimmune symptoms. 115 MTA1 and MTA2/PID expressiones repress p53-dependent transcriptional activity such as p53-mediated cell growth arrest and apoptosis via deacetylation of p5331 (Fig. 8). TWIST1 interacts with MTA2/NuRD to repress the E-cadherin promoter for transcription of E-cadherin and has essential roles as a component of the Mi2/NuRD complex in cancer metastasis; TWIST1's role is independent of another master regulator in “epithelial to mesenchymal transitions” (EMT), Snail, which is regulated by MTA3/NuRD, as discussed below. 51 In addition, BMI1/PRC1and EZH2/PRC2 also interact with TWIST1, and they bind to promoters of E-cadherin and p16/INK4a. It would be interesting to know how these activities are coordinated. MTA2/NuRD complex has a role in mouse genomic imprinting, and is involved in proper imprinted expression of H19 and Peg3 during mouse preimplantation development. 116
MTA3/Mi-2/NuRD
Both isoforms of MTA3 (Fig. 9d) are associated with the vertebrate Mi-2/NuRD complex. 117 MTA3 has been shown to be part of an estrogen-dependent pathway that regulates growth and differentiation of mammary epithelial cells. 117 MTA3 is induced by estrogen and represses the expression of the transcriptional repressor Snail, a master regulator of EMT, resulting in the expression of the cell adhesion molecule E-cadherin and maintenance of a differentiated normal epithelial phenotype in breast cells (Fig. 10).
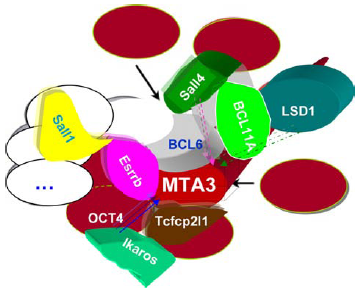
The interactors of mammalian MTA3.
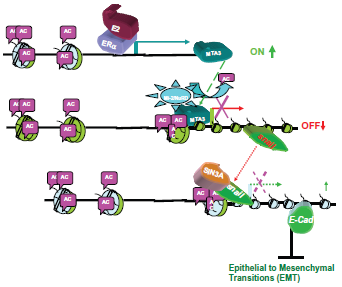
MTA3, a potent suppressor of “epithelial to mesenchymal transitions” (EMT). Through the promoter deacetylation of the Mi-2/NuRD complex, MTA3, an estrogen responsive gene, represses the expression of the transcriptional repressor Snail, thus enhancing the expression of the cell adhesion molecule E-cadherin and maintaining a differentiated normal epithelial status in breast cells.
Depletion of MTA3 leads to aberrant upregulation of the transcriptional repressor protein Snail. An increased expression of Snail is correlated with loss of differentiation and metastasis in breast tumors. MTA3 is also required to regulate B lymphocyte differentiation32,33 by acting as a co-repressor for the transcriptional repressor BCL-6.
The systematic screening of the target genes of the MTA/Mi-2/NuRD complex is being developed. First, LSD1, MTA1/2/3 ChIP-DSL gives a list of genes, including components of the TGFβ signaling pathway, which is a key player in metastatic tumor invasion. Many of these targets are in the main cell signaling pathways, which are involved in apoptosis, Wnt pathways, MAPK, cell communication, cell cycle, focal adhesion, the actin cytoskeleton, etc.
The Histone Binding Proteins/ Retinoblastoma a Tumor Suppressor RbAp46/P48
The Mi-2/NuRD complex contains RbAp46 and RbAp48 (Fig. 11), two proteins that bind to the retinoblastoma A tumor suppressor.39–42,118 They contain a number of WD repeats, a sequence motif that forms the basis for the beta-propeller structure of the beta-subunit of the G protein transducin. 119 These two proteins are presumed to be structural subunits of the NuRD complex, with the potential for different blades of the propeller structure to act as protein interaction surfaces. Interestingly, RbAp46 and RbAp48 have been shown to be components of several multiprotein chromatin modification complexes, 120 in which they interact directly with core histones H3 and H4121 as an adaptor that attaches to tetramers.122,123
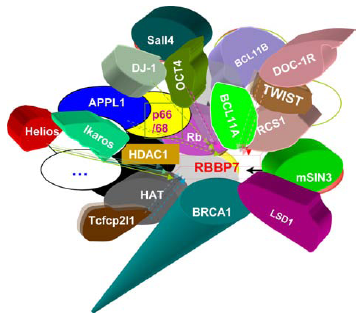
The interactors of mammalian RBBP7/RbAp46/LIN-53.
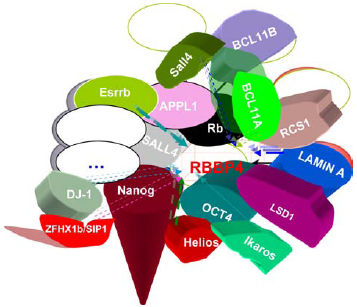
The interactors of mammalian RBBP4/RbAp48.
The Rb complex component LIN-53 (mammalian RbAp48/46)124,125 antagonizes a C. elegans Ras pathway and a synthetic multivulva B (synMuv B) pathway, which includes the homologs of the core subunits of Mi-2/NuRD complex, such as LET-418/ Mi-2β, HDA-1/HDAC1, dcp66, etc. The mutations in the Rb pathway components enhance RNA interference (RNAi) and cause somatic cells to express genes limited to germline-specific P granules. Furthermore, gene inactivations that disrupt RNAi reverse the cell lineage transformations of Rb pathway mutants. These findings suggest that mutations in Rb pathway components cause cells to revert to patterns of gene expression normally restricted to germ cells. Rb may act via a similar mechanism to transform mammalian cells. 126 RBBP7 interacts with DOC1R, a MAP kinase substrate that controls microtubule organization of metaphase II mouse oocytes.127,128
The histone deacetylases
Human histone deacetylases (HDACs) are targets for cancer therapy, and they are classified into three types. 129 The Class I histone deacetylases HDAC1 and HDAC2 are ubiquitously expressed but restricted to the nucleus. Histone deacetylases sometimes combine with histone acetyltransferases (HAT) to form a super HAT HDAC complex. 130
HDAC1 and HDAC2 have no preference for specific DNA sequences; therefore they associate with coactivators and co-repressors which bind to DNA in a specific manner. They both are part of the Mi-2/NuRD complex and bind to the histone binding proteins RbAp46/48 to form a HDAC core complex. HDAC1 and HDAC2 are also associated with other different co-regulators (Fig. 12; Fig. 13a, b Supplementary Table 1). Rb was shown to depend on HDAC1 for the transcriptional repression of E2F target genes. 131 They promote nonhomologous DNA end-joining in the DNA damage response. 50

The interactors of mammalian HDAC1.
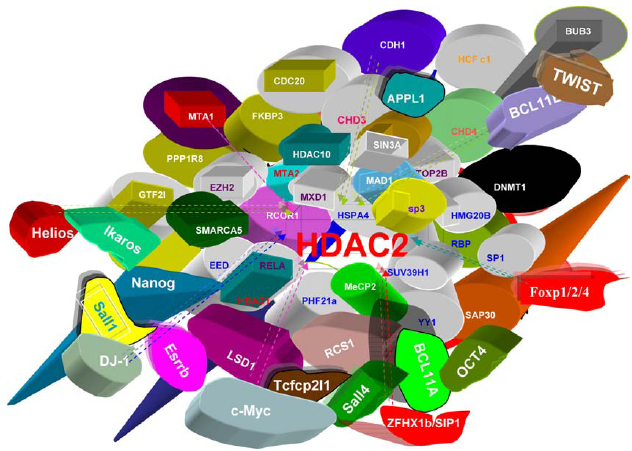

The interactors of mammalian HDAC2.
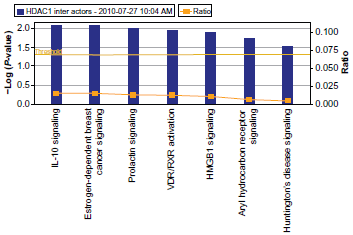
The pathways of interactors of mammalian HDAC1. Reprinted with permission from ingenuity® systems, www.ingenuity.com
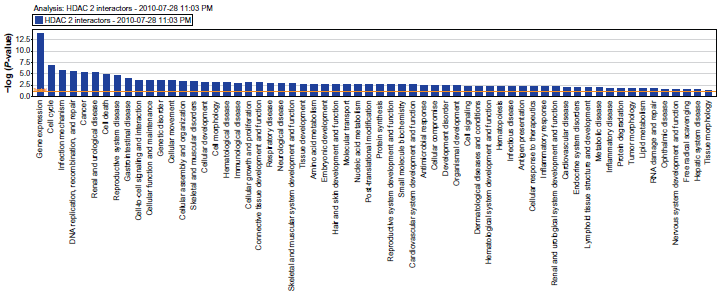
The pathways of interactors of mammalian HDAC2. Reprinted with permission from ingenuity® systems, www.ingenuity.com.
The microarray analysis using C. elegans embryos identified tissue-specific and extracellular matrix (ECM)-related genes as major HDA-1 targets. Ectopic expression of HDA-1 or C. elegans cystatin, an HDA-1 target identified from the profiling, significantly affected mammalian cell invasion. Similarly, RNAi depletion or overexpression of human HDAC1 also affected cell migration. Therefore HDA-1/HDAC1 may play a critical evolutionarily conserved role in regulating the extracellular microenvironment.34,59 The sumoylation of Smo-1 on HDA-1 inhibits LIN-12/ Notch signalling in the vulva. 60 HDA1 contributes to huntingin polyglutamine toxicity. 132 HDA1 is required for cell migration and axon pathfinding. 133
GATAD2a/b (p66/p68)
Some versions of the NuRD complex are also thought to incorporate a second structural and/or regulatory subunit, p66a or p66b,45,134 also known as Gatad2a (p66) and Gatad2b (p68) (Fig. 14). As previously mentioned, the p66 subunits interact with the MBD2 subunit and may be involved in interactions between the Mi-2/ NuRD complex and methylated DNA. 28 Like the RbAp46/48 subunits, both p66 isoforms have the capacity to interact directly with core histones. 135 Small ubiquitin-like modifier (SUMO)-modified forms of p66 efficiently interact with HDAC1, whereas RbAp46 binds to SUMO-p66. 136 The C. elegans p66 negatively regulates the expression of the Delta homolog lag-2 and RNAi knock-down has low penetrance of multiple Muv phenotypes 60 (Zhang and Mueller, unpublished).
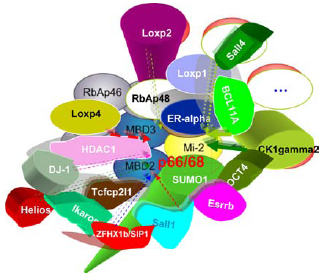
The interactors of mammalian p66/68 (GATAD2a/b).
The Histone 3 Lysien-Specific Demethylase LSD1 independent, font big as the equivalents/other components of Mi-2/NuRD. Since its identification in 2004, LSD1 has been shown to be essential for many cellular processes. Significantly, Wang et al identified LSD1 as an integral component of the human Mi-2/ MTA1-3/NuRD complexes that had profound implications for oncogenesis. 8 In breast cancer cells, LSD1 interacts directly with all MTA1-3 proteins and these different MTA/Mi-2/NuRD complexes have different genomic DNA landing signatures. MTA2 is required for nucleosome LSD1 demethylation. Furthermore, BRCA2-LSD1/Mi-2/NuRD complexes are associated with cancer susceptibility. LSD1/Mi-2/ NuRD complexes suppressed breast cancer metastasis. Overexpression of LSD1 has been correlated with prostate cancer and tumor recurrence during therapy. The association of LSD1 with the androgen receptor switches its substrate specificity from H3K4me/me2 to H3K9me/me2. Polyamine analogues inhibit LSD1 and result in re-expression of aberrantly silenced genes in colon carcinoma.20,137–139 A 3’ domain of the first identified long intergenic noncoding RNAs (LincRNA), HOTAIR, binds the LSD1/CoREST/REST complex, and, consequently, CoREST prevents LSD1 degradation. This coordinated tethering of PRC2 and LSD1 to chromatin couples histone H3 lysine 27 methylation and lysine 4 demethylation of target genes. 140
LSD1 function is implicated in the DNA damage response by demethylating p53. LSD1 maintains DNA methyltransferase, Dnmt1. Loss of LSD1 demethylase activity results in reduced levels of Dnmt1 and global DNA methylation. Genetic ablation of LSD1 causes early embryonic lethality. Loss of LSD1 in embryonic stem (ES) cells reveals a reduction in CoREST levels and the aberrant transcription of 588 genes 7 (Table 2). Like MTAs, LSD1 has SANT and ELM domains. It is unclear whether LSD1 and MTA proteins could have any redundant functions with such domains in a subset of target genes.
As mentioned earlier, spr-5 encodes the C. elegans ortholog of the human histone demethylase LSD1; loss of spr-5 activity can phenotypically suppress mutations in sel-12/presenilin and derepresses expression of hop-1, the second C. elegans presenilin in the LIN-12/Notch pathway. SPR-5 interacts with SPR-1, the C. elegans ortholog of CoREST.61–63
Lamin a and the Mi-2/NuRD Complex
Hutchinson-Gilford Progeria Syndrome (HGPS) is caused by mutations in the LMNA gene, which encodes nuclear lamins A and C. The in vivo physical interaction of lamin A with RBBP4, RBBP7 and HDAC1 points to lamin A being a co-regulator for the nuclear lamina and the Mi-2/NuRD complex in normal cells. Multiple Mi-2/NuRD components are lost in HGPS. Similar to RBBP4/7KD, silencing of any subunit increased the percentage of cells lacking H3K9me3 and HP1γ heterochromatin foci. Furthermore, the KD of HDAC1, MTA3, CHD3 or CHD4 in primary human fibroblasts increased the percentage of cells containing phosphor-H2AX positive foci. Progerin causes the loss of RBB4/7, which is an early event in ageing-associated chromatin defects. Loss of any Mi-2/ NuRD component and reduction of HDAC1 activity is sufficient to recapitulate several ageing-associated chromatin defects. Induction of progerin or the KD of RBBP4 and RBBP7 results in changes within the heterochromatin structure, followed by accumulation of DNA damage. Loss of RBBP4/7 compromises the histone modifications and higher order chromatin structure, possibly making chromatin more susceptible to DNA damage. Thus the Mi-2/NuRD complex is a mediator of ageing-associated chromatin defects. 190
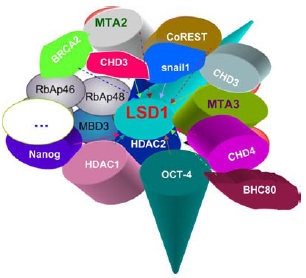
The interactors of mammalian LSD1.
Post-transcriptional modifications and upstream cell signaling pathways of Mi-2/NuRD
A large number of different posttranslational modifications were identified in all major Mi-2/NuRD subunits. These include phosphorylation sites in Mi-2α, Mi-2β, p66α, p66β, HDAC1 and HDAC2. The role of p66 in the NuRD complex arises from its sumoylation.
SUMO-modified forms of p66 efficiently interact with HDAC1, whereas RbAp46 binds to SUMO-p66. 136 MTA proteins display tissue-specific differential expression giving rise to distinct Mi-2/NuRD complexes. Sumolyation of HDA1 in C. elegans is evident. The Mi-2/NuRD complex has many phosphorylation sites, especially on sites in MBD2, MTA1, MTA2 and MTA3. Several post-translational modifications have been conserved, and these might therefore have a role in regulating protein -protein or protein DNA interactions, or in fine-tuning of enzymatic activities. More potential amino acid modifications in chromatin remodeling are determined by DNA methylation and by multiple histone modifications including methylation, ubiquitination, sumoylation, phosphorylation and acetylation.
From Table 1, we analyze the interactors of HDAC1 and HDAC2 or pooled interactors of the subunits of the Mi-2/NuRD complex, except for HDAC2, by using the Ingenuity Pathways Analysis (IPA) program for functional pathway analysis (Fig. 13a, b; Fig. 17).
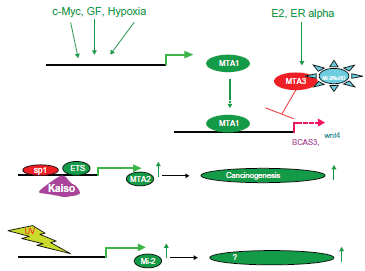
Model of upstream effects on the Mi-2/NuRD complexes.
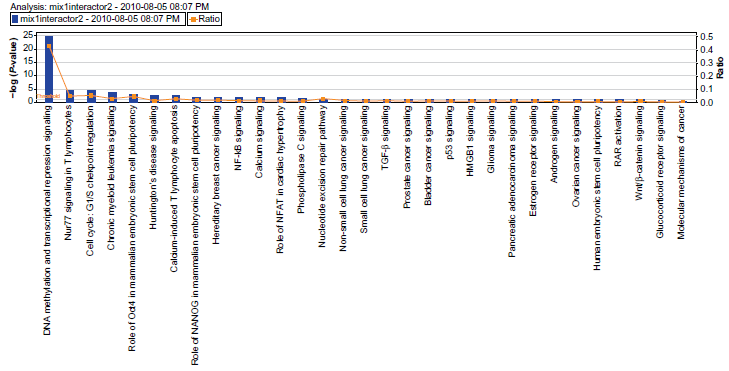
The pathways of pooled interactors of the core subunits of Mi-2/NuRD (excluding those of HDAC2). Reprinted with permission from ingenuity® systems, www.ingenuity.com.
Finally, the expression of MTA2 is partly regulated by SP1 and ETS elements in its promoter 141 and by Kaiso, a component of the human N-CoR complex. 142 Kaiso recruits the NCoR complex to the MTA2 promoter in a DNA methylation-dependent manner, resulting in hypoacetylation and methylation at H3K9 at the promoter region. 142 Therefore, this establishes that Kaiso is a DNA methylation-dependent transcriptional repressor of the MTA2 gene (Fig. 16, middle).
The Methodology for Identifying of Mi-2/NuRD Complex Composition
Classical biology has become “big biology”, ie, high-tech, high throughput and high computation. Identifying protein protein interaction is becoming increasingly important in gaining a molecular understanding of protein function and regulation in cells, in organisms under normal development and in diseased states. However, conventional chromatography has led to the discovery of the Mi-2/NuRD complex.28,82,83 Moreover, single tagging studies like GST contributed significant insights for interacting proteins of the cell or tissue-specific Mi-2/NuRD67,143 (Table 3). The yeast two-hybrid system further identified fruitful co-regulators of the Mi-2/NuRD complex9,67,73,101,144 (Table 3).
Advantages and disadvantages of the methods used for the identification of Mi-2/NuRD complexes.

Mass spectrometry-based proteomics, combined with tandem affinity (TAP)-tag-based protein purification, 145 is one of the most effective strategies for isolating and identifying protein complexes and has revolutionized proteomic experiments (Table 3). Generally speaking, mass spectrometry can identify medium or high abundance proteins. Therefore, previously infrequently expressed proteins could need a crosslink fixation 146 (Zhang, unpublished). Now it has become routine to allow protein identification with high sensitivity and accuracy, and it recently produced several protein interaction network reports. The classic TAP using Protein A and CBP tags has proven successful in yeast to allow rapid purification of protein complexes and minimize the background. However, this classic TAP is not suitable for the purification and identification of proteins from tissues and cell lines. The large tags on either the N-terminus or the C-terminus destabilize the protein (Table 3). Some alternative other tags (eg, FLAG and His (FH-TAP), FLAG and CBP (FC-TAP), protein G and the streptavidin-binding peptide (GS-TAP), and FLAG and STREP (SF-TAP)) provide protein yields from tissues and mammals that are about an order of magnitude higher than the classic one147–149 (Table 3; Zhang et al in preparation). Semi-quantitative specificity filters that are based on peptide spectral count measurements could derive an accurate complex composition. 150 Furthermore, to be extended to targeted therapy, such proteomics-based strategies could identify novel biomarkers that indicate a response of cancer cells to PI3K pathway inhibitors. 151 For Mi-2/ NuRD, Le Guezennee et al. succeeded in identifying MBD2/NuRD and MBD3/NuRD complex compositions; to purify Ikaros/NuRD complexes under reduced stringency conditions, a two-step TAP procedure was successfully used. 152
However, some studies in mammals and C. elegans rely on cDNA overexpression driven by exogenous promoters or transgenic random integration approaches. These transgenes can lack alternate promoters, enhancer elements, and 3 untranslated region (UTR) elements, which play critical roles in the control of gene expression in vivo. In C. elegans, for example, both the daf-12 and fah-1 genes have important enhancer elements that lie outside of the proximal promoter which were missed in promoter-only constructs. Furthermore, many transgene constructs use the unc-54 3'UTR, which prevents regulation by the appropriate microRNA genes. Consequently, generating transgenes with large segments of mammalian and worm genomic DNA would be ideal for capturing all of promoters, splice variants, and 3'UTR control elements. Along with mammalian Bacterial Artificial Chromosome (BAC) or fosmid library, a C. elegans fosmid library, which consists of ~40 kb regions of genomic DNA and covers almost all of the genome, has been recently constructed. Previously, we developed fosmid DNA recombineering technology to carry out profiling studies of the DAF-12 targets by powerful genome-wide screening techniques including ChIP-tiling arrays and assay of interactors by proteomic analysis on transgenic C. elegans lines153,154 (Hochbaun et al in preparation). Importantly, we could capture most, if not all, previously identified targets, and thus their related mechanisms, as well as thousands of novel targets. For the proteins expressed with low abundance, like DAF-12, the integration of a few copies rather than a single copy in transgenic animals significantly increased the capture of bona fide target genes (unpublished).
Recently, an epitope-tagging strategy for the purification with recombination of a full-length BAC that contains Oct4 has pulled down the stemned Mi-2/ NuRD complexes in mouse ESCs. As previously mentioned, fosmid recombinant cloning has the advantage of maintaining the endogenous promoter and therefore natural transcriptional regulation. The technology is amenable to high-throughput delivery, as demonstrated by random integration of tagged BAC transgenes, 155 and should considerably facilitate systematic tagging of genes and analysis of protein complexes with roles in development in different contexts, be it in stem cells, differentiated cell types or even mouse tissue. 156 Previous proteomic studies of Oct4 protein complexes needed lengthy single or tandem purifications from nuclear extracts with streptavidin capture 157 or anti-Oct4 antibodies, 3 and yielded small datasets; in contrast, this novel approach has produced by far the most extensive analysis of Oct4-associated proteins to date. The advantage of using whole extracts and nuclear extracts is that the analysis is not restricted to the nuclear environment, and the dataset encompasses diverse aspects of the life of Oct4, both nuclear and non-nuclear. With such a broad dataset, Oct4 obviously is involved in diverse cellular processes that can have an impact on many aspects of stem cell biology.
For the dissection and assay of different tissue/ organ cancers, going small is the new big, ie, it is able to achieve comprehensive integrated molecular views of such defined cell populations. For the downstream sequences of target genes, low-quantity digital gene expression (LQ-DGE) 158 allows mRNA profiles of as few as 250 cells to be characterized without an amplification step. The new application of chromatin immunoprecipitation combined with high-throughput sequencing (ChIP-seq) from a limited number of cells (10,000) hematopoietic mouse progenitors.159,160 The mass spectrometry-based proteomics assays have become ready for human proteomes. 161 Such a strategy is certainly a hypothesis generator, 161 with some of the most interesting connections revealed being completely unanticipated.
As high-sensitivity and high throughput mass spectrometry, microarray transcriptional profiling and DNA sequencing become more common, assays that scan an entire genome, proteome, ribonome or metabolome will generate a huge amount of data. Bioinformatics tools such as Cytoscape, DAVID, Metacore and Ingenuity Pathway Analysis (IPA) could certainly provide us with functional modules and/or genetic cause-and-effect regulatory networking.162,163
The emerging stemnessed Mi-2/NuRD complexes
Oct-4 expression must be tightly regulated; too much or too little will cause differentiation of the cell. Oct4 can both activate and repress transcriptional targets in mouse and human ESCs.164,165 To date, Oct4 has been shown to be associated mainly with members of repressor Mi-2/NuRD complexes.3,5,6,157 MTA1/Mi-2/ NuRD may have an essential role in normal development as a mediator of pluripotency. MTA1 appears to be the preferred core component of NuRD complexes for Nanog and OCT4 interaction 3 as a potential regulator of pluripotency. Recently, Sall4, a well-known Oct4 partner, and other members of the Spalt-like family of transcriptional cofactors have been shown to associate with NuRD. 166 One of the stemness factors for iPS, c-Myc, is also associated with the Mi-2/ NuRD complex.
Clinical and therapeutic implications
Human cancer is heterogeneous, and this is driven by progressive genetic and epigenetic abnormalities. The Mi-2/NuRD complex is the master regulator of epigenetic alterations in changed patterns of histone modification. In contrast to genetic changes, the epigenetic alterations in cancer cells are reversible by the epi-drugs, for example, the inhibitors of histone deacetylases, DNA methyltransferases and histone demethylases.
MTAs: targets for drug development
MTA1 protein, as a master co-regulator, or its gene, could be an excellent molecular target for cancer therapy, as well as being useful in cancer diagnosis or prognosis. The antisense silencing against MTA1 mRNA had a growth-inhibitory effect on human metastatic breast cancer cell lines.167,168 RNAi knock-down of MTA1 in a human esophageal squamous cell carcinoma cell line resulted in significant inhibition of in vitro invasion and migration properties of the cancer cells. In malignant melanoma cells, knock-down of MTA1 by RNAi successfully suppressed the growth in vitro and experimental metastasis of mouse melanoma cells in vivo. MTA1s may also be a useful target for fighting breast cancer. In an animal model, the effect of the MTA1s peptide blocks the tumor progression of MCF-7, which overexpresses ER-α. MTA1 is a promising antigen for tumor rejection in that it is overexpressed in many different tumors and is expressed at lower levels in normal tissues. An initial study demonstrated the presence of immunogenic MHC class I-restricted peptides of MTA1. 169
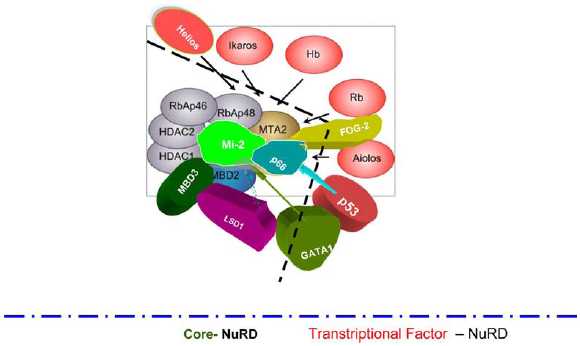
The core-NuRD and transcriptional factor-NuRD.

The core-NuRD, transcriptional factor-NuRD and stemmed Nanog-NuRD.

The core-NuRD, transcriptional factor-NuRD, Stemmed Nanog-NuRD and stemmedOrchestra Oct4-NuRD.

The core-NuRD, transcriptional factor-NuRD, stemmed Nanog-NuRD stemmed Oct4-NuRD and stemmed SALL4-NuRD.

The core-NuRD, transcriptional factor-NuRD, stemmed Nanog-NuRD, stemmed c-Myc-NuRD and stemmed Oct4-NuRD.
MBDs: targets for drug development
MBD proteins may be important modulators of tumorigenesis through the gene silencing mechanism of DNA hypermethylation and may be excellent novel therapeutic targets. 170 Some intensively-investigated DNA methylation inhibitors include the pyrimidine nucleoside analogs decitabine (Dacogen, SuperGen, Inc.) and azacitidine (Vidaza, Celgene), and the nonnucleoside inhibitor hydralazine. Azacitidine and decitabine are both US Food and Drug Administration (FDA) approved for the treatment anemia and chronic myelogenous leukemia (CML). Hydralazine was originally approved for use as an antihypertensive and recently used to reactivate the expression of tumor suppressor genes in cancer. Pharmacologic inhibition of DNA methylation blocks DNMTs and their targeted degradation, and desilences genes that have been aberrantly silenced by hypermethylation. Finally, it causes inhibition of clonal expansion and tumor cell growth, induction of cell differentiation, and cancer cell death. Only rapidly dividing cells such as tumor cells will be targeted as these agents function by being incorporated into newly replicated DNAs.
HDACs: targets for drug development
Epigenetic therapy tries to reverse the aberrations with natural compounds and/or synthetic molecules that are active on specific epi-targets. Vorinostat (Zolinza, Merck) is the first HDAC inhibitor that has been licensed for clinical use. HDAC inhibitors have been tested as promising agents in treating blood-borne cancers, and also in treating polyglutamine diseases.171–174 Two HDACs, hydralazine and magnesium valproate, can sensitize tumor cells to chemotherapy in patients with advanced and solid refractory tumors. 175 Inhibition of histone deacetylation enhances tumor radiosensitization. 176 So far, more than 11 HDAC inhibitors are in clinical development for epigenetic cancer therapy.
G9a: a target for drug development
The MTA proteins are associated with HDAC1/2 and methyltransferase G9a, and specific G9a inhibitors have been recently reported. 177 This raises the possibility that G9a inhibitors such as BIX-01294 might also be used, alone or together with the current FDA-approved HDAC inhibitors, for treating MTA-involved cancers and neurological diseases. 178
LSD1: a target for drug development
The development of LSD1 inhibitors may represent a significant weapon against cancer. To date, only two drugs, pargyline 179 and tranylcypromine,20,180 have been described as LSD1 inhibitors, but their action is not as specific as the well-known anti-Monoamine Oxidase (MAO) agents. Pargyline blocks demethylation by LSD1, and consequently it blocks androgen-receptor-dependent transcription. Thus, modulation of LSD1 activity offers a new strategy to regulate androgen receptor functions, which may turn out to be important in prostate cancer models. 179
RNAi and potential combined therapy
Hypoxia inducible factor (HIF)-1 accumulation favors tumor angiogenesis. 181 5-AZA-CdR represses the hypoxia response pathway by downregulation of HIF-1 targets vascular endothelial growth factor (VEGF). 182 HDACi also plays an important role in inhibiting tumor angiogenesis by concomitantly upregulating antiangiogenetic factors such as VHL and by downregulating pro-angiogenic factors such as HIF-1 and VEGF in cancer cells. 183 MTA1/NuRD has a role in tissue maintenance, via HIF-1a/VEGF expression against hypoxic condition.
Recently, ALN-VSP02 (KSP/VEGF siRNAs), a lipid nanoparticle containing two small interfering RNAs (siRNAs) for kinesin spindle protein (KSP) and vascular endothelial growth factor (VEGF), has demonstrated its potential antitumor activity. Upon intravenous administration, the siRNAs in KSP/ VEGF siRNAs ALN-VSP02ALN bind to both KSP and VEGF messenger RNAs (mRNAs), preventing translation of KSP and VEGF proteins; this may result in growth inhibition of tumor cells that overexpress KSP and VEGF. 184
Interestingly, as already mentioned, some components of the Mi-2/NuRD complex (eg, LIN-53/ RbAP48) in C. elegans could enhance RNAi. Furthermore, the long-term RNAi of Transcriptional Gene Silencing (TGS) with a chromatin remodeling-related mechanism might potentially be used in the future for therapeutic applications of RNAi for prolonged, epigenetic gene silencing. 185 Thus, it may be useful to apply such RNAi against components of the Mi-2/NuRD complex combined with thermo- or radiation- therapy in preclinical cancer.
Furthermore, the Mi-2/NuRD complex plays a crucial role in DNA damage and repair.25,35,48–50 Curcumin (diferuloylmethane) targets DNA damage and repair. 186 A combined strategy of the HDAC inhibitors (eg, Vorinostat, ie, suberoylanilide hydroxamic acid (SAHA) or the safe dietary compound resveratrol) and curcumin therapy provides new hope for cancer patients.
Future Mi-2/NuRD-based approaches
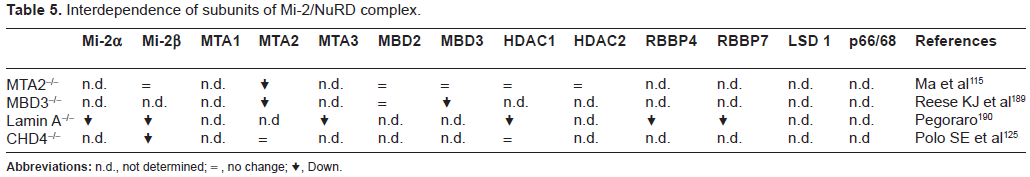
It is highly likely that greater understanding if Mi2/ NuRD subunit structure could contribute to a break-though in anti-cancer drug development since our knowledge in this field and the interdependence among the subunits is in its infancy (Tables 4 and 5). In the future, further understanding of the relationships between the Mi-2/NuRD complex structure and activity, as well as their biological mechanism(s) of action, will hopefully provide novel epigenetic approaches for fighting human cancer.
Crystal structure of subunit of Mi-2/NuRD complex.

Interdependence of subunits of Mi-2/NuRD complex.
Future Mi-2/NuRD stemness-based treatment
A deeper understanding the origin of cancer stem cells (CSC) 190 and their epigenetic characteristics is important for the rational design of cancer treatment. Cancer derived from CSC may be more prone to drug resistance. However, epigenetic changes are mainly reversible through the administration of epi-drugs. Such drugs have been developed for targeting total cell populations and not CSC's specifically. Finally, a stem cell-based vaccine has been shown to successfully address colon cancer. 192 An iPS-based vaccine has also been developed too.192,193 Advancing our knowledge of the Mi-2/NuRD complex could possibly contribute to an improved design to achieve better effects.
The significance of cross-validation with interspecies (Mammals and C. elegans): a personal view
C. elegans has almost the same number of protein-coding genes as a human–-about 20,000 is the latest estimate–-and most of those genes encode similar functions.34,35 So the basic parts set for animal development was established several hundred million years ago. Furthermore, C. elegans has two small evolution-distance nematode sister species, C. briggsae and C. remanei, and similar studies could provide very strong mutual cross-species validation.58, 195 This study of the Mi-2/NuRD complex in C. elegans proved that such cross-research can be a fruitful and probably cost-effective way to study cancer. 196 Besides, we are now able to perform more high-tech, high throughput assays, such as ChIP-seq, quantitative proteomics 197 and TAP on Mi-2/NuRD complexes and their interactors to learn more about them and their clinic implications for beating cancer. Screening all the genes or proteins in an organism is not much more difficult than analyzing a small subset, and robotics and high-throughput screening techniques are now within the reach of most labs. Second, the cost of systems biology scales sub-linearly while the payoffs scale super-linearly. 162 A huge amount of information will need to be analalyzed with current powerful bioinformatic tools 163 so we can obtain information on the behavior of gene groups at a system level, as well as their functional modules and networks. This could be done quickly with genetically tractable organisms like C. elegans, and then projecting its mechanisms to human beings, as has already been demonstrated successfully for C elegans originals such as RNAi, programmed cell death and miRNAs. Put simply, we should be ready to worm our way into elucidating differential regulatory networks between healthy and cancerous tissues.
Discussion
We are towel on the road from “big biology” to “bigger biology” 198 with both “data first” 199 and “hypothesis first” 200 approaches being used together rather than alone; from HGP and NextGen Sequencing to the NextGen mass-spectrometry and HUPO initiatives. Since the international Human Genome Project and Celera Genomics Corporation completing the initial draft of the human genome, the past decade has witnessed remarkable progress in our understanding of the expanding family of Mi-2/NuRD complexes and their powerful master roles in gene regulation during normal development and carcinogenesis across species. This rapid trajectory can continue with the emergence of the human proteome project. The Cancer Genome Atlas, a comprehensive platform to accelerate our understanding of the genetics of cancer using innovative genome analysis technologies, is now available. Weekly, even daily developments are emerging from this interesting Mi-2/NuRD complex family. Finally, all these new findings could be promisingly pave the way for a highly effective genetic and epigenetic therapy in the near future to cure cancer.
Disclosure
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
Footnotes
Acknowledgements
The authors thank the working groups of Dr. Mueller, Dr. Fisher and Dr. Calderwood for their personal communications and insightful discussions. Also thanks to the Swiss Federal Government for their fellowship support. We apologize to our many colleagues whose work could not be cited here owing to space considerations.
