Abstract
Degenerative retinal diseases, such as glaucoma, age-related macular degeneration, and diabetic retinopathy, have complex etiologies with environmental, genetic, and epigenetic contributions to disease pathology. Much effort has gone into elucidating both the genetic and the environmental risk factors for these retinal diseases. However, little is known about how these genetic and environmental risk factors bring about molecular changes that lead to pathology. Epigenetic mechanisms have received extensive attention of late for their promise of bridging the gap between environmental exposures and disease development via their influence on gene expression. Recent studies have identified epigenetic changes that associate with the incidence and/or progression of each of these retinal diseases. Therefore, these epigenetic modifications may be involved in the underlying pathological mechanisms leading to blindness. Further genome-wide epigenetic studies that incorporate well-characterized tissue samples, consider challenges similar to those relevant to gene expression studies, and combine the genome-wide epigenetic data with genome-wide genetic and expression data to identify additional potentially causative agents of disease are needed. Such studies will allow researchers to create much-needed therapeutics to prevent and/or intervene in disease progression. Improved therapeutics will greatly enhance the quality of life and reduce the burden of disease management for millions of patients living with these potentially blinding conditions.
Keywords
Background
Definitions of Key Terminology.
In eukaryotic cells, various proteins play important roles in establishing and maintaining the structure of DNA within a cell. Histone proteins are the primary protein components of chromatin and are responsible for condensing the DNA into nucleosomes that form a Chromatin Modifications Affect Gene Expression. Changes in Epigenetic Marks, Such as DNA Methylation and Histone Modifications, Each Contribute to the Regulation of Gene Expression. DNA Methylation in the Promoter Regions of Genes is Generally Associated with Decreased Gene Expression. Histone Modifications can be Either Activating or Repressive. Histone Acetylation and Phosphorylation are Generally Associated with Active Genes; Histone Methylation and Ubiquitination Arrangements are Associated with Either Active or Repressed Genes.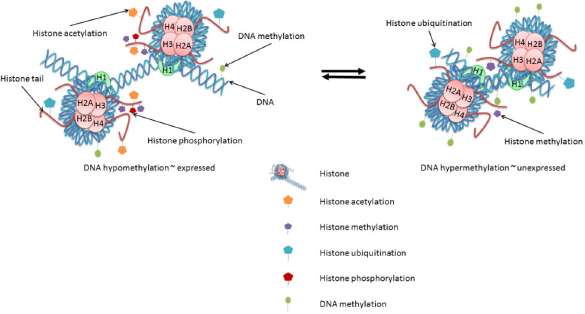
Additionally, the N-terminal ends, or
Histone modifications also serve as markers recognized by specific TFs involved in activation or repression of eukaryotic gene expression. Much effort of late has gone into deciphering the so-called
A number of
In addition to histone modifications, modifications to the DNA itself have profound effects on gene expression. DNA methylation-the addition of a methyl group to the C5 position of cytosine bases that are followed by guanosine bases within the DNA sequence (5′-CpG-3′)-is strongly associated with the repression of gene expression. CpG-rich regions of the genome, referred to as CpG islands, are often found in the 5′ regulatory regions of genes.
10
Actively expressed genes generally have unmethylated CpG islands near their transcription start sites, whereas unexpressed genes generally have methylated CpG islands near their transcription start sites. Methylation patterns can be heritable across both meiotic and mitotic cell divisions. In genomic imprinting, for example, methylation is used to ensure that single copies of particular genes are repressed during egg or sperm cell generation in a maternal-or paternal-specific pattern, which persists into the adult.11, 12 DNA methylation is also used during development to program cell differentiation by specifying the particular subset of genes to be expressed by each cell type.
13
Of particular relevance to the development of age-related disease is that environmental factors induce DNA methylation changes throughout an organism's lifespan.
14
Figure 2 illustrates a number of environmental factors associated with retinal degenerations such as age-related macular degeneration (AMD) that have also been shown to associate with changes in DNA methylation, including tobacco smoke, alcohol, diet (particularly omega-3 fatty acids and antioxidants), and obesity.15–27 The cumulative effects of each of these factors on the epigenetic landscape of an individual's genome may gradually perturb the expression profile of a tissue sufficiently to transform it from an otherwise healthy state to a diseased state.
Epigenetics in Aging and Age-Related Disease. Stable Epigenetic Marks may be the Link Between Genetic and Environmental Processes Involved in the Development of Age-Related Diseases, Such as AMD. Environmental Influences Contribute to Epigenetic Changes that Accumulate with Age. Risk Factors, Such as Diet, Obesity, Smoking, Sun Exposure, and Age, may Elicit Epigenetic Changes that Accumulate Over a Lifetime, Eventually Resulting in Altered Expression of Genes Involved in the Disease Process. These Environmental Influences Contribute to Epigenetic Modifications, such as DNA Methylation (Green Ovals), Histone Methylation (Purple Pentagons), Histone Acetylation (Orange Pentagons), Histone Ubiquitination (Blue Pentagons), and Histone Phosphorylation (Red Pentagons). The Epigenetic Changes that Accumulate Throughout the Genome May Associate with Transcriptional Changes at the Affected Genomic LOCI. Such Expression Changes at Disease-Relevant Loci May Promote Either Protection Against or Progression of Age-Related Disease. The Sum of these Effects Over Time May then Perturb the Normal, Healthy Homeostasis Enough to Result in the Development and/or Progression of Diseases Such as AMD.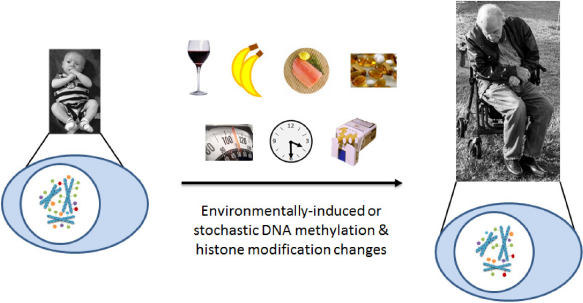
Such gradual epigenetic modifications may play significant roles in the development of age-related retinal diseases. The retina is a region at the back of the eye onto which the lens focuses an image and which contains the photoreceptor cells that transmit the image as a signal to the brain. The macula is the region of the retina responsible for central vision, with the fovea at its center. Figure 3A shows a fundus image (photograph of the back of the eye) and an optical coherence tomography (OCT) image (which shows a cross-section of the macula) of a healthy retina. In the OCT image, the retina is the upper cellular layer, with the retinal pigment epithelium (RPE) and choroid below (posterior to) the retina. The choroid comprises blood vessels that support the retina, with the RPE serving as the inner blood-retinal barrier through which nutrients and waste must pass to transfer between the retina and the choroidal blood supply.
28
Progression of Retinal Disease. (
Definition of Epigenetics
As stated earlier, changes to DNA structure that do not involve changes to the DNA sequence are considered
The epigenetic landscape of a particular gene or genome can be affected by various factors, including genetic and environmental influences, as noted earlier. The activation and repression of gene expression is a complex interplay of numerous factors that must be tightly regulated to ensure proper response to environmental signals. Upon detection of an environmental signal, sequence-specific DNA-binding transcription activators or repressors (TFs) are activated alternatively by various methods, such as: (1) Directly binding to the signal, e.g., the nuclear receptor family of TFs in metazoans, including the estrogen receptor alpha and the retinoic acid receptor. 31, 32 (2) Sensing a disruption to the cellular homeostasis caused by the environmental signal, e.g., protein kinase R-like endoplasmic reticulum kinase (PERK) sensing misfolded proteins in the endoplasmic reticulum of eukaryotes, leading to activation of TFs involved in apoptosis.
33
(3) Through a signaling cascade propagated from the extracellular signal-binding receptor, e.g., vascular endothelial growth factor (VEGF)binding to VEGF receptors to induce downstream activation of several TFs in metazoans.
34
Once activated, the TFs bind to target sequences within regulatory regions of the genome to modulate the transcription of target genes. Transcriptional activators recruit RNA polymerase II (RNA pol II) and other transcriptional machinery, including chromatin remodelers, to the gene to initiate transcription. The so-called
Alternatively, transcriptional repressors can bind to their target sequences and recruit chromatin remodeling proteins to convert the local chromatin to, or maintain it in, a heterochromatic state. For example, the Krüppel-associated box (KRAB) zinc finger proteins are sequence-specific DNA-binding transcriptional repressors that induce the formation of heterochromatin through the recruitment of KRAB-associated protein 1 (KAP1), which then recruits histone methyltransferases (ie, SETDB1) and deacetylases and promotes the binding of heterochromatin protein 1 (HP1), which maintains the heterochromatic state.36–38
In addition to the function of gene-specific TFs, the process of transcription itself is generally associated with the exchange of histones in chromatin. This histone exchange is mediated by adenosine triphosphate (ATP)-dependent chromatin remodelers, such as SWI/SNF, FACT, and Spt6, and facilitates the progression of RNA pol II through the chromatin. 39 In addition to allowing the progression of RNA pol II along the DNA template, the dynamic deposition and removal of histones from the DNA allows for dynamic control over nucleosome composition. The replication-independent, transcription-dependent deposition of noncanonical histones, such as H3.3 and H2A.Z, provides one example of changing histone composition.40, 41 Additionally, the continued removal and deposition of nucleosomes requires the continual proximity of protein complexes involved in that process, therefore providing a mechanism for rapid changes in response to environmental changes, either continued induction through continual recruitment of nucleosome remodelers or rapid shutoff upon loss of chromatin remodeler recruitment. The turnover of the nucleosomes also provides a potential mechanism for conservation of a histone modification pattern by copying a pattern from a redeposited H3/H4 heterotetramer or H2A/ H2B dimer to a newly deposited counterpart. The turnover also allows for the dilution of histone marks in the absence of the continued activity of histone-modifying enzymes by incorporation of unmodified histones.
The expression of canonical histones (H3, H4, H2A, and H2B) is limited to the S-phase of the cell cycle, when the histone proteins are needed for incorporation into the daughter chromosomes.42, 43 The noncanonical histones, such as H3.3 and H2A.Z, are expressed throughout the cell cycle and are deposited in the wake of RNA pol II during transcription-dependent histone exchange. 44 This likely contributes to the observation that, in postmitotic cells, the noncanonical histones accumulate, while the canonical histones are gradually lost from the cell with age. 39
Genome-wide epigenetic studies of retinal disease, risk factors, and related pathologies.

Epigenetic Resource Databases.

Aging
Throughout development, cells acquire chromatin changes that regulate the transcriptional output of each cell. In gametogenesis and embryogenesis, DNA methylation is erased and reset at specific reprograming stages to ensure appropriate gene expression and totipotency of embryonic cells. 52 This reprograming is used by the cell for processes such as X-chromosome inactivation for dosage compensation, repression of invasive DNA such as retrotransposons, imprinting to ensure parental allele-specific expression, and to otherwise control the expression of genes relevant to the developmental process. Additionally, the process of cellular differentiation, in which individual cells acquire specialized gene expression profiles specific to their individual tissue types and functions, requires further changes to the chromatin landscape (reviewed for various tissues such as skeletal muscle, 53 myeloid cells, 54 and the lens and retina 55 ). Much effort of late has focused on defining the tissue-specific differential methylation patterns in differentiated tissues in relation to differential gene expression patterns in an attempt to understand the underlying mechanisms of tissue-specific processes.56–60 Even after a cell has reached its terminally differentiated state, further changes that associate with the aging process can continue to accrue throughout the lifespan of the cell. Methylation levels generally decrease with aging, although individual genes may have increased methylation levels with age.61–70 Replication-independent histone exchange results in the accumulation of histone variants, such as H3.3 and H2A.Z, in older cells. 39 Cells from aged human beings have been shown to have reduced levels of both the heterochromatin maintenance protein HP1 and the H3K9me3 histone modification known to recruit HP1 to chromatin. 71 Experiments in model organisms showing that the reversal of age-dependent chromatin changes results in extension of lifespan suggest that at least some of the age-related chromatin changes are indeed causative of aging, rather than just a result of aging.72–74 Much work is still needed to fully understand the role of acquired chromatin changes in the aging process.
Sirtuins are a class of HDACs implicated in lifespan regulation and the promotion of healthy aging through various epigenetic and nonepigenetic cellular roles, including telomere maintenance, DNA repair, metabolism, stress tolerance, cellular differentiation, apoptosis, and inflammation.75, 76 Sirtuins have also been implicated in age-related diseases, such as diabetes, cardiovascular disease (CVD), neurodegenerative diseases, and some cancers. 9 Roles for sirtuins have been observed in ocular aging. The involvement of sirtuin 1 (SIRT1) in ocular aging, including retinal degeneration, has been the most well characterized of the seven mammalian sirtuins. SIRT1 has been associated with retinal stem cell self-renewal, and decreased expression of SIRT1 with age has been suggested to contribute to the aging process. 77 SIRT1 also exhibits neuroprotective effects in the retina at least partially through its antioxidant, energy balancing, and antiapoptotic functions. 78 Abnormal SIRT1 localization is also thought to promote the apoptosis of photoreceptor cells and precocious aging in the rd10 mouse model of retinal degeneration. 79 SIRT6, which is regulated by SIRT1, has also been implicated in retinal aging, and SIRT6 deficiency in mice results in increased levels of retinal cell apoptosis.80, 81 Altered TF functions have also been observed in the aging mammalian retina. For example, the function of the oxidative stress response master regulator nuclear factor (erythroid-derived 2)-like 2 (Nrf2) was shown to be altered in aged C57Bl6/J mice compared with young C57Bl6/J mice. 82 The aged mice in this study exhibited higher uninduced (basal) expression levels of Nrf2 antioxidant targets compared with the young mice. Upon induction with an oxidative stressor, the Nrf2-dependent antioxidant genes were robustly induced in young mice, but not in aged mice. Nrf2 was also induced by the oxidative stressor in young mice, but not in aged mice. Expression of Nrf2 is regulated by DNA methylation and modulated by dietary factors such as sulforaphane; differential methylation of the Nrf2 promoter is also associated with various diseases. 83
The question of whether complex age-related diseases are the inevitable eventual outcome of the aging process or whether they are, in fact, the result of aberrant aging is yet to be elucidated and is of great interest in the field today. Along with the extension of lifespan achieved to date by modern medical advances has come an increase in the prevalence of age-related conditions that severely impact the quality of life of older adults. Degenerative diseases of the retina, such as glaucoma, AMD, and DR, affect >15 million people >40 years old in the US and are the leading causes of blindness in older adults.84–86 As of 2013, 11 million individuals were affected with AMD, per year, in the United States alone. Because aging is the greatest risk factor, this number is anticipated to increase to 22 million by 2050 as a result of the population aging (http://www.brightfocus.org/).
We present here what is known of epigenetic effects in major retinal diseases associated with aging: glaucoma, AMD, and DR. Further work in this field has promise to revolutionize the way we approach therapeutic interventions by revealing points of intervention with potential long-term influences on the progression of disease.
Glaucoma
Glaucoma is the leading cause of irreversible blindness in the US and worldwide, according to the World Health Organization (2015); it ranks second overall after cataract, which is reversible.87–89 Primary open-angle glaucoma is considered the
Studies of gene expression in donor eye samples affected with glaucoma have shown upregulation of profibrotic factors in the TM and AH, including transforming growth factor beta (TGF-β), thrombospondin-1 (TSP1), and connective tissue growth factor, compared with unaffected controls.102–104 Extensive investigations of epigenetic mechanisms have yet to be performed in human tissues affected by glaucoma. However, epigenetic processes involved in fibrosis in other cell types/disease processes give insight into likely processes involved in the glaucoma-associated fibrosis. Hypermethylation of TSP1 in colorectal cancer suppresses TSP1-mediated activation of TGF-β. 105 TGF-β-mediated regulation, in turn, is sensitive to the chromatin modifications, including DNA methylation and histone acetylation, of its downstream target genes.106–108
Hypoxia is thought to contribute to the degeneration of retinal ganglion cells in glaucoma. 109 The expression of hypoxia-induced factor 1 alpha (HIF1α), which induces the adaptive transcriptional response to hypoxia,110, 111 is greater in the retina and optic nerve head of glaucomatous eyes compared with controls. 110 HIF1α is stabilized under hypoxic conditions by the loss of a specific oxygen-dependent degradation mechanism involving proline hydroxylation of HIF1α, 112, 113 which allows HIF1α to dimerize with HIF1β to form an active HIF1 heterodimer. 114 Hypoxia induces the translocation of HIF1α from the cytoplasm to the nucleus, where it recruits the histone acetyltransferase CBP/p300 coactivator and regulates transcription of target genes. 115 HIF1α activity is regulated epigenetically in a few ways. The HIF1α promoter contains a hypoxia response element with a CpG that is methylated under hypoxic conditions; a DNA methyltransferase (DNMT) inhibitor increased hypoxia-induced gene expression; methylation of the hypoxia response element influences HIFα binding; and various histone modifications at hypoxia responsive genes are specific to hypoxic conditions. 116
Further evidence for epigenetic effects in hypoxia has been provided by Shahrzad et al who showed an inverse relationship between DNA methylation and hypoxia in primary cancer cells and in normal human dermal fibroblasts. 117 Also, under chronic hypoxia in prostate epithelial cells, the induced HIF1α stability is lost, and both H3K9 hyperacetylation and DNA hypermethylation accumulate, which are proposed to maintain the hypoxia-conditioned state in the absence of HIF1α.m
Genome-wide epigenetic studies utilizing well-characterized/phenotyped glaucoma-affected tissues and matched normal controls will be important for elucidating the epigenetic changes that occur within the eye throughout the progression of the disease, from normal to the onset of increased IOP, to the damaging effects of that pressure on the optic nerve. Additionally, the use of monozygotic twin pairs discordant for glaucoma for genome-wide epigenetic analysis will further highlight epigenetic processes associated with the development of glaucoma. The information gained by combining rigorous genome-wide epigenetic, genetic, and expression data will lead to improved understanding of the disease and, therefore, better means to treat it.
Age-related Macular Degeneration
Similar to glaucoma, AMD is also a multifactorial, progressive, neurodegenerative disease. AMD is characterized by the sub-RPE accumulation of yellowish lipid-rich, protein-containing
Genome-wide association studies (GWAS) for AMD risk alleles have identified 19 genetic variants that associate with AMD at genome-wide significance levels, including genes involved in the complement pathway, lipid metabolism, angiogenesis, and atherosclerosis, among other cellular processes.
126
A recent exome chip study identified 34 loci representing both common and rare variants independently associated with AMD.
127
This study found evidence for causal roles for three very rare coding variants in three genes (
Twin studies estimate the genetic component of AMD risk at ~40%-70% of the total risk.
130
Known environmental risks include smoking, obesity, and diet.
25
Environmental contributions to disease potentially involve epigenetic changes that affect the expression of genes involved in the generation of disease and may also convert environmental exposures into heritable phenotypes.
131
This relationship between epigenetics, environment, and heritability may also lead to increased measures of heritability in monozygotic twin studies-due to the shared zygotic epigenetic history of monozygotic twins-than would arise from genetic heritability alone.
131
Therefore, epigenetic factors may contribute to both the heritable and nonheritable components of risk. This is consistent with observations that some chromatin changes are conserved through gametogenesis, whereas others are erased in the process.
52
The idea that environmental forces introduce small epigenetic effects that accumulate with time (illustrated in Fig. 2) is also consistent with both the observations that age is the primary risk factor for AMD and that siblings with identical AMD risk profiles from major AMD genetic loci, smoking, body mass index (BMI), and CVD can still be extremely discordant for disease development (Fig. 4).
Siblings Extremely Discordant for AMD. These Fundus Images Illustrate the Retinas of Discordant Siblings with Identical Risk Profiles for Major AMD Genetic Loci, Smoking, BMI, and CVD. Note the Severe Neovascular AMD in One Sibling and Normal Macula in the Other at a Comparable Age. Environmentally Induced Epigenetic Changes May Explain the Occurrence of Such Discordance for AMD or Other Age-Related Disease. OD, Oculus Dexter (Right Eye); Os, Oculus Sinister (Left Eye). Photographs Taken from Deangelis Laboratory Patient Cohorts. The Study Protocol was Reviewed and Approved by the Institutional Review Board at the University of Utah and Conforms to the Tenets of the Declaration of Helsinki.
Evidence for the involvement of epigenetic processes associated with macular degeneration has been accumulating in the field. Early work looking at proteins involved in the modification of chromatin showed requirements for these functions in relation to AMD. Suuronen et al demonstrated that HDAC inhibitors increased the expression and secretion of clusterin, the major protein component of drusen, in a human RPE cell line, 132 suggesting that histone deacetylation activity is important to limit the amount of clusterin produced in RPE cells to prevent its accumulation in drusen. Chen and Cepko provided further evidence for the role of histone deacetylation in the maintenance of a healthy retina when they showed that HDAC4 promotes the survival of retinal neurons in mouse. 133 (p4) Given that neovascularization is thought to be a response to hypoxia caused by drusen deposits interrupting the flow of oxygen from the choroid through the RPE and into the retinal cells, it is also relevant that knockout of the noncanonical H2A.X histone in mice reduced the neovascular response to hypoxia. 134
In addition to functional studies looking at the effects of histone-modifying activities, studies have looked at the association of chromatin modifications at genes relevant to the AMD disease process in human tissue samples. Using a DNA methylation microarray and bisulfite pyrosequencing of frozen human donor RPE/choroid samples with a postmortem interval of up to nine hours, Hunter et al identified the
Another early example of differential methylation in AMD correlating inversely with gene expression came from Wei et al's report of hypomethylation in the promoter of
As a step in that direction, Oliver et al investigated DNA methylation levels in peripheral blood samples and frozen sucrose gradient-treated peripheral retinas (postmortem interval of donor eye tissue was not reported) of AMD patients with either GA or neovascularization compared with unaffected control patients.
138
In the only genome-wide epigenetic study of AMD to date, they observed hypomethylation at the
The genome-wide methylation study by Oliver et al in AMD-affected RPE/choroid and retina might be a step toward understanding the epigenetic mechanisms underlying gene expression of AMD. Building on their study, researchers will need to go on to evaluate the epigenetic changes that occur throughout the progression from normal to severe disease. This will require well-characterized tissue samples with short postmortem intervals to tease out epigenetic changes at each stage of the disease: preclinical through advanced stages. This will allow for the identification of early changes that may influence the progression of AMD and, therefore, present early therapeutic interventions to prevent the progression to severe forms of the disease.
Further methodological improvements will also need to be incorporated to accomplish this goal. Increased quality of well-characterized donor eyes for each stage of the disease, along with well-characterized phenotypically normal controls, will increase the power of the study to identify changes associated with each step of the disease process. Also, the postmortem interval between death and preservation of the tissue must be carefully selected to preserve the integrity of the sample and minimize the noise in the genome-wide data. 48 It is also important that these studies be carried out in a fresh tissue that is directly involved and relevant for the specific disease process being studied. Similar to gene expression profiles, DNA methylation profiles differ among tissues.13, 14 Therefore, it cannot be taken for granted that an easily accessible tissue, such as peripheral blood, will adequately substitute or replicate for the relevant, disease-involved tissue.
Diabetic Retinopathy
In 2012, ~10% of Americans had diabetes, according to the American Diabetes Association.
139
Every diabetic patient is at risk of developing DR, which is the most common cause of blindness among working-age adults in developed countries.
140
The risk for DR increases with increasing time since diagnosis of diabetes.141, 142 Similar to the age-related risk for AMD and glaucoma, the time component of risk for DR is also consistent with the idea that environmental influences introduce small epigenetic effects that accumulate with time (Fig. 5).
Why do Some Patients Progress Beyond and Others Stop at an Intermediate Stage of Disease? The Complex Functional Networks at Play Within a Cell Or Tissue are Likely Susceptible to Small Perturbations Exerted by Environmental Influences Over Long Periods of Time. The Gradual Accumulation of Altered Expression or Function of Key Factors Within a System May then Lead to Dysfunction and Disease. Disease-Promoting Factors (Such as Smoking, Obesity, and Poor Diet) Work to Alter the Networks in Ways that Lead to Pathological States, Whereas Disease-Preventing Factors (Such as Antioxidant Consumption and Healthy Bmi) Oppose the Progression of Disease by Keeping the System Within the Bounds of Healthy Function. The Ratio of Disease-Promoting and -Preventing Influences May Balance at Different Equilibrium Points, Leading to Individual Cases that Progress Only so Far Along the Pathological Pathway Before Reaching that Balance Point. By Discovering the Various Promoting and Preventing Factors, Along with their Respective Significances at Each Point Along the Disease Progression Timeline, Researchers (And Eventually Clinicians and Patients) Will be Able to Balance the Networks to Prevent or Reverse the Progression of Disease. Photographs Taken from the Deangelis Laboratory Cohorts. The Study Protocol was Reviewed and Approved by the Institutional Review Board at the University of Utah and Conforms to the Tenets of the Declaration of Helsinki.
Type 2 diabetes is characterized by hyperglycemia due to impaired insulin secretion by pancreatic β-cells and/or to insulin resistance in peripheral tissues, including skeletal muscle, adipose, and liver. Aging is a major risk factor for type 2 diabetes, along with obesity, low physical activity levels, poor diet, and genetics. DNA methylation at the type 2 diabetes risk genes,
Many studies have shown links between epigenetics and the development of type 2 diabetes. An early study linked nutrition to heritable, potentially epigenetic changes that increased or decreased the risk for type 2 diabetes. 146 Exposure to famine during the paternal grandfather's childhood is protective against, whereas access to abundant food supplies associates with increased risk for diabetes as a cause of death for his grandchild. Also, studies have shown that diabetes during pregnancy increases the risk for that child to be diagnosed with diabetes as an adult. 147
In addition to the multigenerational heritability of epigenetic changes involved in the development of type 2 diabetes, many studies of late have shown evidence for long-term epigenetic changes within an individual with hyperglycemia. The observation of a
A genome-wide methylation study looking at differential DNA methylation in skeletal muscle biopsies from type 2 diabetic patients and age-matched controls identified 838 promoter regions that were differentially methylated between the two groups, including the promoter for
A more recent genome-wide methylation study identified 1,649 genes with differential DNA methylation in pancreatic islets between subjects with type 2 diabetes and normal controls, most of which involved modest changes in percent methylation in the range of 5%-10%. 155 Of these 1,649 differentially methylated genes, 102 also showed differential gene expression in type 2 diabetics compared with controls. Pathway clustering of these genes identified pathways enriched within these gene sets, providing insight into the possible pathological mechanisms. Genome-wide studies such as this provide targets for further functional analysis that will lead both to a better understanding of the disease process and to novel therapeutic targets for improved management of the disease.
Numerous PTMs to histone proteins have also been associated with the development of type 2 diabetes and accompanying complications.148, 149 One example is that the
DR is a common complication of diabetes, affecting greater than half of patients having type 2 diabetes for ≥10 years and is the leading cause of blindness among working-age adults.
142
Although glaucoma and AMD each have associations with processes, including immune function, which suggest that they may have systemic contributions to the disease processes, DR is clearly the result of systemic disease. DR develops via hyperglycemic induction of vasoactive and inflammatory factors that leads to vascular damage and neuronal injury, further leading to ischemia and the subsequent formation of new vessels, which are prone to bleeding and lead to retinal detachment and ultimately vision loss (Fig. 3D).157, 158 Specific epigenetic modifications associated with the development of DR include decreased H3K4me2 in the promoter and enhancer regions of the
These studies have provided insights into the epigenetic mechanisms that may underlie gene expression in type 2 diabetes and DR. Genome-wide epigenetic studies are needed to further elucidate the epigenetic processes involved in the progression of DR. In particular, investigations comparing the epigenetic profiles of monozygotic twins discordant for DR will potentially highlight differentially modified loci with significant influences on disease development. Such studies will improve the understanding of disease progression and identify potential opportunities for therapeutic intervention.
Future Directions: Epigenetics as Therapeutics
Studies seeking to understand the various epigenetic changes-and their relationship to gene expression-involved in retinal diseases will inform researchers regarding the intricate molecular mechanisms and processes that lead to the development and progression of complex blinding diseases. With an understanding of the disease mechanisms, researchers will develop treatments to effectively manage the full spectrum from preclinical to advanced stages of disease, including both interventions that prevent clinical manifestations in asymptomatic patients and those that prevent the progression to blinding advanced stages and/or promote the regression of existing disease phenotypes.
Data indicating which genes undergo epigenetic modifications associated with the development of retinal disease will provide potential therapeutic targets for interventions that alternatively fix the aberrant chromatin modification(s), target the particular gene/gene product whose expression is pathologically altered as a result of the aberrant chromatin modification(s), or target proteins or RNAs that interact with or facilitate the effects of the affected gene/gene product. Therapeutics that target chromatin-modifying enzymes, such as HDAC inhibitors and DNMT inhibitors, were initially developed as treatments for cancers, and some are currently in clinical use or clinical trials (see the NIH website for current clinical trial information: ClinicalTrials.gov).162–164 More recently, epigenetic therapeutic agents are being considered as potential treatments for noncancerous complex diseases, such as CVD and diabetes, which share common risk factors with age-related retinal diseases.165–168 Additionally, preclinical studies are investigating the potential use of epigenetic therapeutics for the treatment of AMD and DR. 169 However, at the time of writing this report, clinical studies of therapeutic agents that directly modulate epigenetic processes have yet to be executed.
A potential epigenetic intervention was observed in a recent clinical trial looking at specific B-vitamin supplementation. The study looked at the association of folic acid, vitamin B6, and vitamin B12 supplementation on the development of AMD in women with either preexisting CVD or at least three risk factors for CVD, which are also risk factors for AMD. 170 The authors found an association between folic acid/vitamin B6/vitamin B12 supplementation and protection against AMD development. An ongoing clinical study is also investigating the effects of folic acid, vitamin B6, and vitamin B12 supplementation on the progression of DR. 171 Folic acid is an upstream methyl donor for DNA methylation; vitamins B6 and B12 are involved as cofactors in the process of transferring the methyl group indirectly from folic acid to the DNA. Recent data suggest an association between dietary folic acid intake and global methylation levels. 172 Therefore, the association of folic acid and vitamins B6 and B12 supplementation with decreased incidence of AMD may indicate an epigenetic influence on disease progression and, therefore, an epigenetic modulation of disease risk via supplementation. However, the effect of folic acid and vitamins B6 and B12 supplementation on disease risk may result from alternative molecular mechanisms, such as direct antioxidant effects. 170 Therefore, additional studies are needed to elucidate the molecular mechanism of folic acid and vitamins B6 and B12 supplementation in the progress of AMD and DR.
Further studies into the epigenetics of blinding retinal diseases will provide therapeutic targets for improved treatment strategies. Current treatments for neovascular AMD have limited results and require frequent intraocular injections, which are not ideal for the patient. The development of therapies that reduce or eliminate the need for injections will greatly reduce the burden of care on patients and the healthcare system at large. For example, administration of pharmacological agents orally or via eye drops will not only reduce the pain and trouble associated with current therapies but could also improve vision over the long term.
Perspectives
The progression of complex disease is a multifactorial process influenced by various inputs, including genetic, epigenetic, and environmental elements. Extensive genome-wide analyses are insufficient to explain the development and progression of the complex disease from phenotypically normal through advanced disease. Studies utilizing well-characterized collections of tissue, which take into account tissue integrity and represent each stage of the disease, may help us to understand the molecular changes taking place as the disease progresses. Genome-wide epigenetic and expression data for each stage of the disease will provide candidates for involvement in the progression of disease. However, whether these factors are causes or effects of disease progression will have to be teased out with additional studies designed to determine the mechanistic role of each of these factors.
An ideal experimental setup for genome-wide epigenetic experiments will need to incorporate various features to ensure sufficient power and noise reduction in the resulting data (Fig. 6). Each of the nine characteristics illustrated in Figure 6 contributes to the production of meaningful epigenetic data that can be used to identify genes that are differentially methylated/modified in specific disease states. The generation of high-quality genome-wide epigenetic data pertinent to the development or progression of disease requires that relevant tissue(s) be sampled from both thoroughly phenotyped diseased subjects and well-characterized control subjects who are appropriately age and gender matched. Sampling from both affected and unaffected tissues for both the diseased and the control subjects will also allow for determination of changes that are isolated to the relevant tissue(s). Furthermore, the postmortem interval (time from death to preservation of the tissue sample) is critical for the extraction of accurate information. Both the number and quality of the samples influence the power to identify statistically significant differences associated with disease.
Ideal Experimental Setup for Genome-Wide Epigenetic Experiments. Each of These Nine Characteristics Contributes to the Production of Meaningful Epigenetic Data that can be Used to Identify Genes that are Differentially Methylated/Modified in Specific Disease States. The Combination of Robust Epigenetic Data with RNAseq Expression Data Provides Additional Information Pertaining to the Mechanism of Disease by Correlating Expression Changes with Underlying Chromatin Modifying Processes and Providing Initial Insights into the Mechanisms of Disease. These Studies Identify Genes and/or Regulatory Regions that Potentially have Causative Roles in the Development or Progression of Disease. Further Mechanistic Studies Focusing on the Roles of Each Correlated Gene/Regulatory Region Will then Provide Additional Understanding of the Disease and Targets for Therapeutic Interventions. Incorporation of Genetic and Lifestyle Data Will Also Improve Analysis by Integrating the Full Spectrum of Risk into Studies of Disease Mechanisms.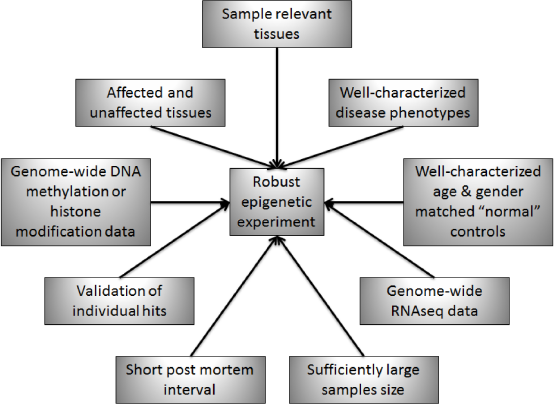
The combination of robust high-resolution epigenetic data with RNAseq expression data provides additional information pertaining to the mechanism of disease by correlating expression changes with underlying chromatin modifying processes and providing initial insights into the mechanisms of disease. These studies identify genes and/or regulatory regions that potentially have causative roles in the development or progression of disease. Further mechanistic studies focusing on the roles of each correlated gene/regulatory region will then provide additional understanding of the disease pathways, networks, and points for therapeutic interventions. Incorporation of genetic, lifestyle, and other environmental data will also improve analysis by integrating the full spectrum of risk into studies of disease mechanisms.
Why do some patients’ diseases progress and others’ stop at an intermediate stage? The complex functional networks at play within a cell or tissue are likely susceptible to small perturbations exerted by environmental influences over long periods of time (Fig. 5). The gradual accumulation of altered expression or function of key factors within a system may then lead to dysfunction and disease. Disease-promoting factors (such as smoking, obesity, and poor nutrition) work to alter the networks/pathways in ways that lead to pathological states, whereas disease-preventing factors (such as antioxidant consumption and healthy BMI) oppose the progression of disease by keeping the system within the bounds of healthy function. The ratio of disease-promoting and -preventing influences may balance at different equilibrium points, leading to individual cases that progress only so far along the pathological pathway before reaching that equilibrium point. By discovering the various promoting and preventing factors, along with their respective significances at each point along the disease progression timeline, researchers (and eventually clinicians and patients) will be able to balance the networks and pathways to prevent or reverse the progression of disease. Such interventions are desperately needed to address the growing costs of complex diseases on the quality and duration of life of millions of people. Additionally, therapeutics capable of preventing progression to advanced, debilitating stages of disease will also decrease the overall financial cost of managing these patients’ diseases.
Summary
Scientists in the field need to tease out the various relationships of cause and effect among genetic, environmental, and epigenetic contributions on both gene expression changes and protein interactions which lead to or result from pathological processes, including potentially circular effects of self-propagating cycles of decline.
With an understanding of the complex relationships between genetic, epigenetic, and environmental factors involved in the development of complex diseases, such as degenerative retinal diseases, preventative and therapeutic interventions will follow that will both decrease incidence of debilitating diseases and also reduce the severity of and complications arising from such diseases, thereby improving the quality of life for older patients.
Author Contributions
Analyzed the data: KLP and MMD. Wrote the first draft of the manuscript: KLP. Contributed to the writing of the manuscript: KLP and MMD. Agree with manuscript results and conclusions: KLP and MMD. Jointly developed the structure and arguments for the paper: KLP and MMD. Made critical revisions and approved final version: KLP and MMD. All authors reviewed and approved of the final manuscript.
