Abstract
Melanoma is one of lethal malignant skin tumors and the sole effective cure for the disease can be achieved by surgical resection of primary tumor and early diagnosis for melanoma is crucial for patients' prognosis. Detection of novel tumor-related antibodies would aid in the diagnosis of early-stage cancer, in the detection of recurrence and in the development of a more effective immunotherapy. In the middle of the exploration of a candidate of biomarker for melanoma by proteomics-base technique, we encountered the coexistence of autoantibodies for α-enolase and γ-enolase in serum derived from a patient with melanoma, who had received the repeated treatments with alkylating agents and interferon β. Although melanoma is known to be a highly antigenic tumor, it is still unclear why these autoantibodies appeared. To evaluate the usefulness in detecting the circulating α-enolase or γ-enolase autoantibodies in serum from melanoma patients for biomarkers for tumor progression, more studies are needed.
Introduction
The incidences of melanoma have steadily increased worldwide, and the mortality rates have increased as well, with a 5-year survival rate of 49% and 20% for stage III and IV, respectively in Japan. 1 Despite intensive clinical research, the sole effective cure for the disease is surgical resection of primary tumor as early as possible. The poor prognosis for melanoma is due to lack of sensitive and specific serum marker for surveillance of melanoma progression in early-stage melanoma.2,3 So, the identification of novel tumor antigens and corresponding autoantibodies in sera from patients with tumor are important to establish the diagnostic system for early-stage malignant tumor and to explore new and effective immunotherapies.
In 2001, Brichory et al4,5 introduced a proteomics-based approach to identify tumor antigens and corresponding autoantibodies and discovered anti-Annexin I and II autoantibodies in sera from patients with lung cancer. Shortly afterwards, several reports on autoantibodies against novel proteins in cancer-bearing subjects have been published with this new developmental method.6–8
By applying a proteomics-based approach as established by Brichory et al, we have been exploring to identify a novel antigen in melanoma and a corresponding autoantibody in sera from patients using cultured melanoma cell lines. In the process of the project, we happened to find the co-existence of an autoantibody for both α-enolase and γ-enolase in serum derived from a patient with melanoma.
Materials and Methods
Cell lines and sera from patients
The three human melanoma cell lines (C32TG, G361 and Mewo) were purchased from the Health Science Research Bank, Osaka, Japan, and maintained as described previously. 9 Sera from 20 Japanese patients with melanoma (stage 0 in two patients, stage I in three patients, stage II in 11 patients, stage III in four patients) and four healthy controls were analyzed in this study. The present project using patients' sera was approved by the Institute Review Board of Osaka Medical College and all of the analyses were performed after getting informed consent from each patient.
Clinical manifestation of patient No.19
Patient No. 19 was a 40-year-old man with a melanoma occurring in his chest (pT3N0N0, stage II). Six years after the surgical resection and five courses of chemotherapy using interferon β in addition to 5-(3,3-Dimethyl-1-triazenyl)imidazole-4- carboxamide (DTIC), Nimustine hydrochloride (ACNU) and vincristine, left axillar lymph node metastasis was found, when the patient's serum was collected. This patient also had a history of ulcerative colitis; one of the inflammatory bowel disease possibly caused by immunological abnormality, for two years.
Sodium Dodecyl Sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and two-dimensional gel electrophoresis (2D-PAGE) and Western blotting (WB)
The cells were harvested by scrapping and suspended in buffer 1% SDS/PBS/complete (Roche Applied Science, Mannheim, Germany). Solubilized proteins extracted from the mixture of three melanoma cell lines (C32TG, G361, Mewo) were separated by sodium dodecyl polyacrylamide gel electrophoresis (SDS-PAGE) and two-dimensional PAGE.
Proteins (10 μg) were applied to each lane of a Multigel (85 mm x 90 mm) and an immobilized pH3–10 non-linear gradient strip gel (GE Healthcare Bio-Science, Uppsala, Sweden) for the isoelectrofocusing (Ettan IPGphor II: GE Healthcare Bio-Science, Uppsala, Sweden).
Separated proteins were transferred onto a polyvinylidene fluoride (PVDF) membrane (Hybond P, GE Healthcare Bio-Science, Uppsala, Sweden) for 1 h at a constant of 20 V, using a TransBlot SD cell system (BioRad, Hercules, CA) and visualized using SimplyBlue SafeStain (Invitrogen, Carlsbad, CA). After transfer on the PVDF membrane, the membrane was incubated with blocking buffer consisting of Tris-buffered saline (TBS)/0.1% Tween 20 and 5% nonfat dry milk overnight at 4 degrees, washed and then incubated with diluted test sera at a 1:500 dilution for 2 hrs, and the membrane was reacted with horseradish peroxidase conjugated anti-human IgG (GE Healthcare Bio-Science, Uppsala, Sweden), at 1:2500 dilution for 1 hr at room temperature and then developed with an enhanced chemiluminescence system (GE Healthcare Bio-Science, Uppsala, Sweden) and then detected with LAS-3000 Luminoimage Analyzer (Fuji Film, Tokyo, Japan).
Simultaneously, the 2D-PAGE gel was stained with coomassie brilliant blue and the pieces of stained gel corresponding to the WB positive spots or bands were excised. Each piece of the gel spots or bands was prepared using the standard method reported by Brichory FM et al. 4 The extracts were desalted in MonoTip C18 micro-columns (GL Sciences Inc., Tokyo, Japan) before MS analysis.
Protein identification by mass spectrometry
The digests were loaded onto the AnchorChip Target plate (Bruker Daltonics, Bremen, Germany) by mixing 1 micro L of each solution with the same volume of an alpha-cyano-4-hydroxycinnamic acid solution (0.3 g/mL of acetone-ethanol (1:1 v/v)). Measurements were performed using an Ultraflex I matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF/TOF) mass spectrometer (Bruker Daltonics, Bremen, Germany). Calibration was accomplished by using external peptide calibration standard II (Bruker Daltonics, Bremen, Germany). The peptide mass fingerprint was used for protein identification using the Mascot Search engine based on the entire NCBIN and SwissProt protein databases.
Detection of autoantibody against α-enolase and γ-enolase by 2D-PAGE and WB
Goat polyclonal anti-α-enolase antibody (sc-31857; Santa Cruz, CA) and monoclonal anti-γ-enolase antibody (sc-21738; Santa Cruz, CA) were used at 1:500 dilution for WB and was processed for incubations with patient sera, horseradish peroxidase conjugated anti-goat IgG (GE Healthcare Bio-Science, Uppsala, Sweden) and anti-mouse IgG, secondary antibody.
Detection of autoantibody against α-enolase and γ-enolase by immunoprecipitation, SDS-PAGE and WB
Serum from a patient with melanoma was added to agarose beads conjugated with protein A and protein G. After incubating for 3 hrs under stirring, α-enolase recombinant protein (097070; Funacoshi Co. Ltd) and neuron specific enolase (RC615; Cosmo Bio Co. Ltd.) were added. After more over-night incubation under stirring and suspension in SDS sample buffer, proteins separated by 2D-PAGE were analyzed by WB with Goat polyclonal anti-α-enolase antibody (sc-7455; Santa Cruz, CA), rabbit polyclonal anti-α-enolase antibody (sc-15343: Santa Cruz, CA) and mouse monoclonal anti-γ-enolase antibody (sc-21738; Santa Cruz, CA).
Results
Figure 1 shows WB pattern of SDS-PAGE of the mixture of proteins solubilized from C32TG, G361 and Mewo using sera from 20 cases of melanoma and four healthy controls.
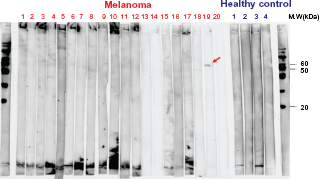
Western blot (WB) of SDS-PAGE of the mixture of proteins purified from C32TG, G361 and Mewo using sera from patients with melanoma and normal subjects. Sera from the patient No.19 showed positive single band.
We found one positive band at ca 55 kDa on PVDF membrane in sera from one melanoma patient (No.19) but not in sera from other melanoma cases and normal controls. This observation suggested that serum from patient No.19 reacted with some specific antigens of melanoma.
Figure 2 represents the WB pattern of 2D-PAGE of mixture of proteins from C32TG, G361 and Mewo with sera from a patient; No.19 (a), with polyclonal anti-α-enolase antibody (b) and with a monoclonal anti-γ-enolase antibody (c). As shown in Figure 2A, we detected a Rosary positive spot in the WB profile from a patient with melanoma (No.19). We confirmed that this spot reacted with a monoclonal anti-γ-enolase antibody (Fig. 2B), but did not react with polyclonal anti-α-enolase antibody (Fig. 2C).
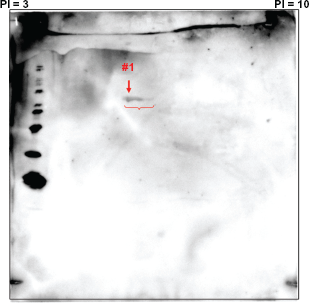
WB of 2D-PAGE of the mixture of proteins derived from C32TG, G361 and Mewo using sera from the patient (No.19). rosary positive spots are seen in the WB profile.
This spot in the CBB stained gel corresponding to the WB profiles was analyzed with the proteomics-based technique using MALDI-TOF/TOFMS. Figure 3A shows that the MS spectra (peptide mass fingerprint) obtained from spot #1 (as shown in Fig. 2 (a)). Peaks numbered in the figure are identified as the m/z of tryptic peptides from α-enolase having high score. As shown in Figure 3B, the MALDI-TOF/TOF mass spectrum of the peak, MH+ = 1908.38 in Figure 3A reveals a part of sequence of α-enolase from 163 to 179 amino acid residues. Figure 4 shows the comparison between α-enolase and γ-enolase sequences, and α-enolase specific sequence was recognized by MALDI-TOF/TOF MS in the peaks; MH+ = 1908.38.
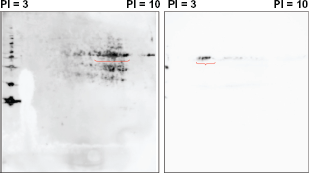
WB with a commercially available polyclonal anti-α-enolase antibody (A;left) and with a monoclonal and anti-γ-enolase antibody (B;right). rosary positive spots in Figure 2. were reacted with a monoclonal human anti-γ-enolase antibody, but not with a polyclonal human anti-α-enolase antibody.
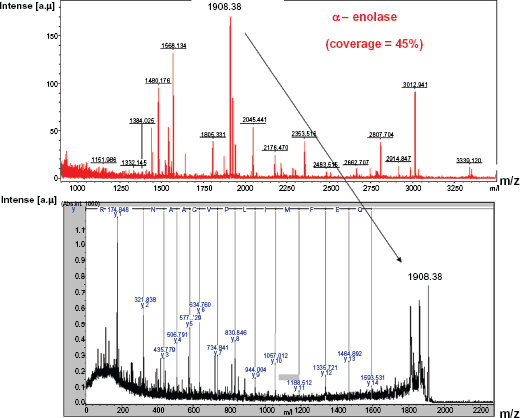
A) (upper) MALDI-TOFMS spectra obtained from the protein in spot #1 shown in Figure 2. B) (lower) MALDI-TOF/TOFMS spectra obtained from a peak, MH+ = 1908.38, which revealed the sequence 163–179 of the α-enolase.
Serum from patient No.19 was immunoprecipitated with commercially available α-enolase recombinant protein (70 KDa) and γ-enolase (55 KDa), and the precipitates were subjected to WB analysis with rabbit polyclonal anti-α-enolase antibody and monoclonal anti-γ-enolase antibody. Precipitation reacted with both mouse monoclonal anti-γ-enolase antibody and rabbit polyclonal anti-α-enolase antibody (Fig. 5), suggesting that serum of patient No.19 included an antibody against α-enolase as well as that against γ-enolase.

Sequence of α-enolase and β-enolase: Matched peptides analyzed by MALDI-TOF/TOFMS are shown in red. The sequence encircled by blue squares were the residues of amino acid or peptides matched only with α-enolase.
Discussions
In this study, we applied a proteomics-based approach to explore novel tumor antigens of melanoma using three commercially available melanoma cell lines (C32TG, G361 and Mewo) and related autoantibodies to these antigens in sera from patients with melanoma. During this research, we encountered autoantibodies against both α-enolase and γ-enolase in serum from one case of melanoma.
We detected rosary positive spots from this patient (No.19) reacted with monoclonal anti-γ-enolase antibody, but did not react with polyclonal anti-α-enolase antibody in the WB profile. However, one of the spots in the CBB stained gel was analyzed with the proteomics-based technique using MALDI-TOF/TOFMS and we identified the sequence of α-enolase in the spot. From the MS data and Swiss-Prot database search, we initially presumed that the epitope of α-enolase in the mixture of proteins from three melanoma cell lines was changed from the normal sequence during the transformation process, possibly resulting in the reaction with monoclonal γ-enolase antibody. Finally, we confirmed that both α-enolase and γ-enolase autoantibodies co-existed in the serum of patient No.19 by immunoprecipitation and WB of SDS-PAGE analyses.
Enolase (2-phospho-D-glycerate hydroxylase) is one of the glycolytic enzymes that consists of three distinct subunits: α (non-neuronal enolase), found in many tissues; β, predominantly existing in muscle; and γ (neuron-specific enolase), found only in neurons and neuroendocrine tissues. 10 Usually each subunit combines to form five different isoenzymes; αα, αβ, αγ, ββ, and γγ. Enolase was also identified in several types of tumor cells or tissues, and its enzymatic activity of enolase has been detected in sera of some cancer patients such as lung cancer and neuroblastoma.11–14
Recently, Lee KH et al have shown that enolase is a multifunctional protein, and its ability to serve as a plasminogen receptor on the surface of variety of hematopoietic, epithelial and endothelial cells suggests that it may play an important role in the intravascular and pericellular fibrinolytic system. 15 In addition, enolase is also recognized as one of the heat-shock proteins, which is also an immune-related molecule and it seems to play a crucial role in the pathogenesis of autoimmune and inflammatory disease. 15 Dot et al 16 clarified changes of the immune response against α-enolase after tumor resection is because of the less acquisition of a cancer-related epitope of α-enolase. Moreover, an α-enolase autoantibody has been detected in sera from various autoimmune disease, ie, systemic lupus erythematosus with renal disease, rheumatoid arthritis, Hashimoto's encephalopathy, inflammatory bowel disease and malignant disorders such as adenocarcinoma of the lung.17–21 In addition, α-enolase was identified in serum from one out of ten melanoma cases using the similar experimental system of proteome approach recently. 22
So far, the existence of an autoantibody for γ-enolase as shown in this study has not been reported either in malignant tumor or in autoimmune disorders.
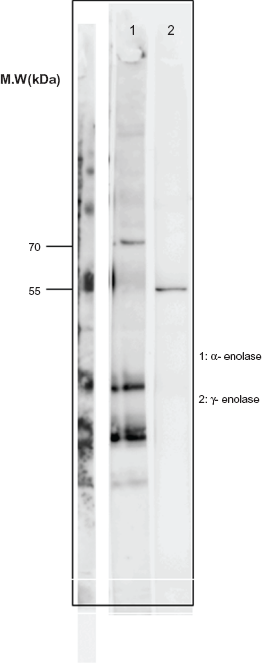
WB of SDS-PAGE of proteins purified from commercial α-enolase recombinant protein(70KDa) and γ-enolase(55KDa) immunoprecipitated with Sera from a patient with melanoma (No.19). Positive band is shown in lane 1 and 2 with an antibody against α–enolase and γ-enolase, respectively.
The biological significance of antigenecity to these two enolases in a melanoma patient remains unproven, while melanoma, a tumor from neuroectodermal cells, is known to be very immunogenic. Chemotherapy to melanoma may have played a role in the generation of autoantibodies. 23 Adjuvant immunotherapy by interferon for advanced melanoma induces serologic and clinical manifestation of autoimmunity.24,25 Autoimmune conditions have been expected to be associated with an improved prognosis in melanoma patients.26,27 Moreover, there may be other factors that may influence the immune response including variability in MHC molecules and in antigen presentation ability among tumors and cancer-prone individuals.
We cannot mention the mechanism for the emergence of autoantibodies to both α-enolase and γ-enolase in the melanoma patient. Also we cannot deny a possibility of susceptibility to produce an autoantibody to α and γ-enolase in relation to the complication of ulcerative colitis in patient No.19. 20 Further examination is needed to elucidate the mechanism of the generation of both α- and γ-enolase antibodies in a melanoma-prone subject.
In summary, we have first identified autoantibodies against both α-enolase and γ-enolase in serum from a patient with melanoma. However, to evaluate the usefulness in detecting the circulating α-enolase or γ-enolase autoantibodies in serum from melanoma patients as biomarkers for melanoma progression, more studies are needed.
Disclosure
This manuscript has been read and approved by all authors. This paper is unique and is not under consideration by any other publication and has not been published elsewhere. The authors and peer reviewers of this paper report no conflicts of interest. The authors confirm that they have permission to reproduce any copyrighted material.
