Abstract
Matrix metalloproteinases (MMPs) are a family of zinc-dependent proteolytic enzymes that degrade extracellular matrix (ECM) components like collagen, fibronectin, and laminin. While this activity is important for normal development, morphogenesis, and wound healing, deregulation of MMP activity has been implicated in a number of cardiovascular diseases, including congenital heart defects, atherosclerosis, myocardial infarction, and congestive heart failure. MMPs are good potential diagnostic indicators of cardiovascular disease, but current detection methods are time consuming and quite laborious. This review will discuss MMP biology, current methods for detection of MMPs from patient samples, and potential new developments in multiplexed analysis of MMPs.
MMP Biology
Extracellular enzymes play a crucial role in the interaction between cells and their environment. Matrix metalloproteinases (MMPs) are a family of zinc-dependent proteolytic enzymes important in the degradation and turnover of extracellular matrix (ECM) components like collagen (Visse and Nagase, 2003). Since the initial description of collagenolytic activity in tadpole tail resorption (Gross and Lapiere, 1962), MMPs have been shown to function in tissue remodeling, embryonic development, morphogenesis, reproduction, cell-cell adhesion, migration and invasion, and growth factor and cytokine signaling (Nagase and Woessner, 1999). There are five major types of MMPs that were initially categorized according to substrates, including collagenases, gelatinases, stromelysins, matrilysins, and membrane-type MMPs (Folgueras et al. 2004). It is now known that many MMPs have multiple substrates, and the current nomenclature uses a numerical system in which these enzymes are classified by domain structure rather than by substrate. To date, twenty-four different MMPs have been identified, based largely on sequence homology with collagenase-1 (MMP-1). Table 1 contains a list of MMPs, including the current name, the old substrate-based name, information about the molecular features of the enzyme, and possible roles in human physiology for each enzyme. Figure 1 shows a schematic diagram of the MMP structure showing the arrangement of the various domains.
Comprehensive list of human matrix metalloproteinases.

Name as given in Woessner and Nagase, 2002.
Calculated using proteomic tools (pl/MW) at www.expasy.ch/pi_tool.html
dVacant MMP designations (Woessner and Nagase, 2002); numbers were assigned to these MMPs but they proved to be one of the known MMPs.
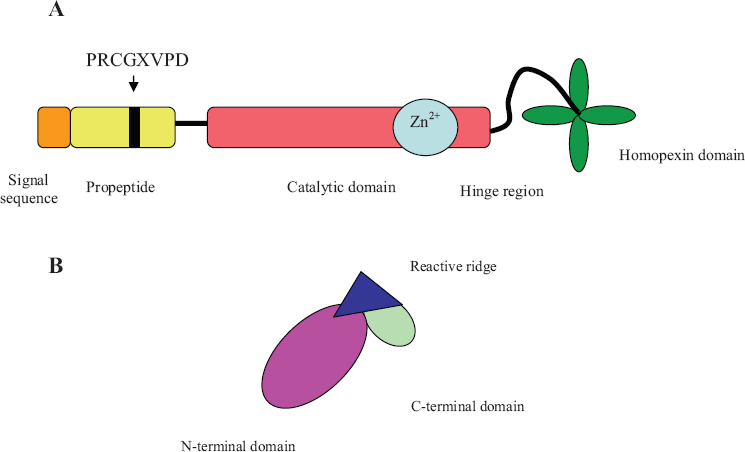
Schematic diagram of MMP domain structure (
Each MMP consists of a specific domain sequence which includes the signal peptide, the pro-peptide domain, the catalytic domain, and the C-terminal hemopexin-like domain, which is present in almost all MMPs. Several MMPs (e.g. MMP-14, -15, -16, and -24) have additional domains such as a trans-membrane and a cytoplasmic tail (Nagase and Woessner, 1999). MMPs are typically found in an inactive pro-enzyme form known as a zymogen, and cleavage of the pro-peptide domain yields the active proteinase. The pro-peptide domain is part of a cysteine switch motif that contains the conserved amino acid sequence PRCGXPD; this maintains the enzyme in an inactive form by interacting with the zinc ion at the active site and preventing binding of the substrate (Van Wart and Birkedal-Hansen, 1990; Becker et al. 1995). The catalytic domain contains the catalytic zinc ion bound by three histidine residues in conserved sequence
MMPs are generally secreted as pro-enzymes requiring cleavage in order to be activated. In vivo, activation commonly occurs through interaction with plasmin or via interaction with other proteinases (Lijnen, 2001). MMP activity can also be controlled by a series of endogenous inhibitors, including ≈2-macroglobulin in the plasma (Cawston and Mercer, 1986) and TIMPs (tissue inhibitors of metalloproteinases) in the tissues. Four TIMPs are currently known to form complexes with MMPs and inhibit enzyme activity (Visse and Nagase, 2003). The overall ratio of TIMPs levels to MMPs levels are thought to be a major determinant of MMP activity. TIMPs generally consist of an N-terminal domain and smaller C-terminal domain connected by a ridge-like loop structure (Fig. 1B). Both the N-terminal and C-terminal domains contain several disulfide bridges, and the entire molecule forms a wedge structure that inserts into the active site of the MMP, inhibiting enzyme activity. In addition, MMP activity may be inhibited by several other molecules, including the NC1 domain of type IV collagen (Petitclerc et al. 2000), tissue factor pathway inhibitor-2 (Herman et al. 2001), and RECK (reversion-inducing cysteine-rich protein with kazal motifs) (Oh et al. 2001; Liu et al. 2003). The extensive and complex regulation of MMPs highlights their important role in normal human physiology.
Deregulation of MMPs has been implicated in a wide variety of human diseases, most notably in tumor progression and metastasis for several forms of cancer, particularly breast (Duffy et al. 2000; Yoon et al. 2003; Folgueras et al. 2004; Stuelten et al. 2005; Jinga et al. 2006), ovarian (Kim et al. 2006), and gastric carcinomas (Torii et al. 1997) Other pathologies associated with inappropriate MMP activity include rheumatoid arthritis (Maeda et al. 1995; Brennan et al. 1997; Yamanaka et al. 2000), hepatitis (Lichtinghagen et al. 2000; Koulentaki et al. 2002), and osteoarthritis of the hip (Masuhara et al. 2002). In addition, MMPs have been recognized as having significant roles in the development of a wide range of cardiovascular diseases, including congenital heart defects (Brauer, 2006), atherosclerosis (Galis et al. 1994; Kalela et al. 2002; Watanabe and Ikeda, 2004), and myocardial infarction (Kai et al. 1998; Zhao et al. 2005). The concentrations of MMPs found in patient samples in some of these diseases are shown in Table 2. Whether MMPs can be used as reliable indicators of disease state or not remains to be seen and will likely depend on advances in analytical detection methods.
Concentrations of MMPs in selected human pathologies.
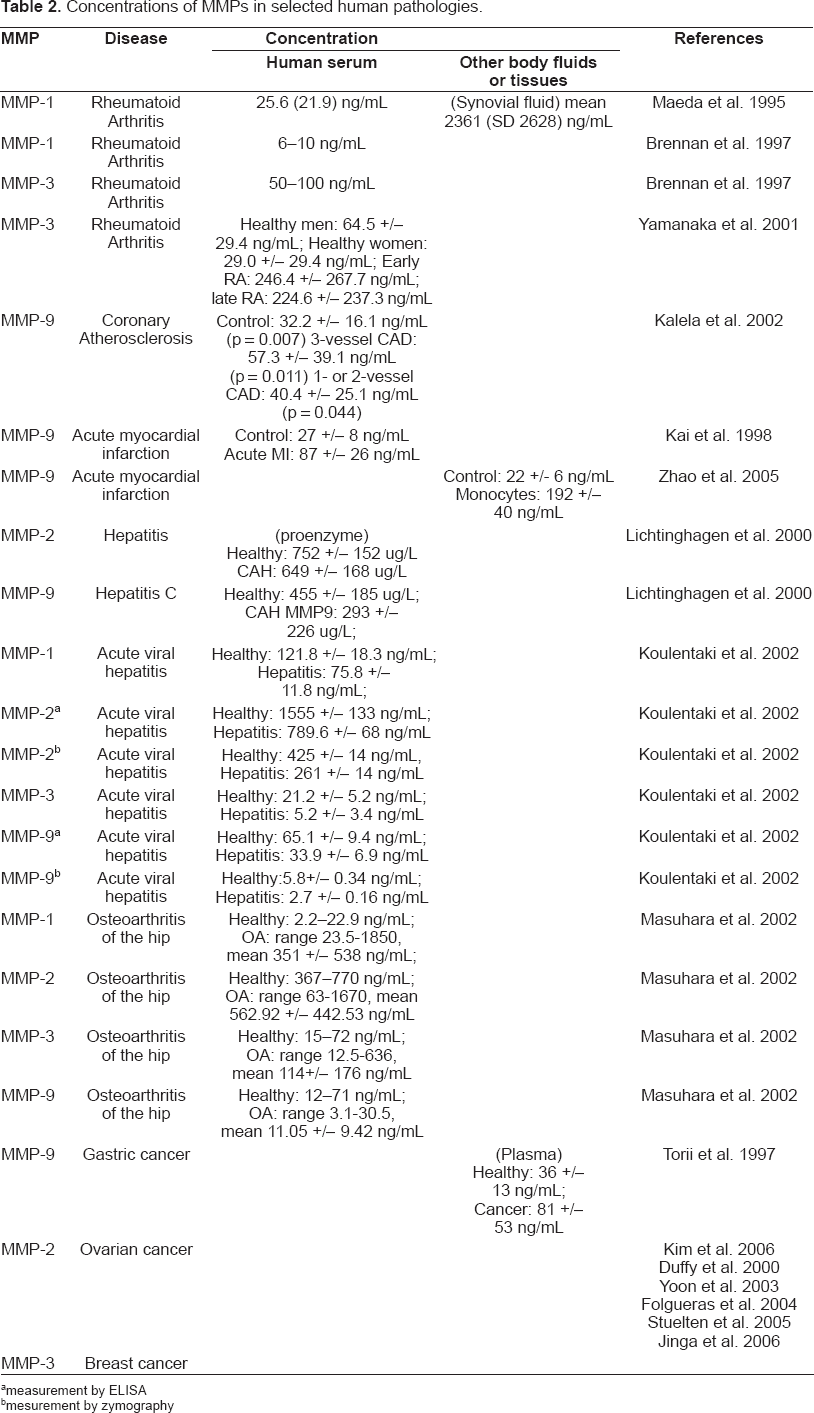
measurement by ELISA
mesurement by zymography
MMPs in Cardiovascular Disease
Each year more than 25,000 babies in the United States are born in with congenital heart defects. During embryogenesis, the formation of the heart and vasculature requires extensive cell migration, proliferation, and organization into three-dimensional networks, and MMPs are key regulators of these processes. MMP-2 is one of the earliest MMPs known to be expressed during heart development (Cai et al. 2000). Neutralizing antibodies for MMP-2 or the use of broad spectrum MMP inhibitors led to severe heart tube defects in avian embryos (Linask et al. 2005). Midline fusion was prevented and cardiac bifida occurred because the MMP-2 inhibition, suggesting a crucial role for this enzyme in normal cardiac development. Additional studies are required to understand the extent of MMP involvement in embryonic cardiovascular development and the degree to which they contribute to congenital defects.
A growing body of evidence indicates that increased MMP activity plays a role in the development of adult cardiovascular disease. Atherosclerosis occurs when injury to the endothelial lining leads to accumulation of lipids and infiltration of activated monocytes, leading to formation of a plaque. MMP activity, mainly from activated vascular macrophages, may contribute to either plaque formation or the rupturing of plaques leading to coronary occlusion, or both (Watanabe and Ikeda, 2004). Atherosclerotic plaques were found contain significantly higher quantities of activated MMP-2 than non-atherosclerotic tissue extracts (Galis et al. 1994), and subjects with acute coronary syndrome have increased plasma levels of MMP-1, -2, and -9 (Palazzuoli et al. 2006). MMP-8 concentrations were reported to be higher in patients with coronary artery disease (CAD) compared to those without CAD, and MMP-8 levels were also increased with the number of stenotic vessels (Kai et al. 1998). Another study involving patients with coronary atherosclerosis indicated similar findings for MMP-9 (Kalela et al. 2002); serum concentrations were highest in patients with 3-vessel CAD (see Table 2 for 1- and 2-vessel CAD), and the difference remained statistically significant after adjustment for age, diabetes, and sex (p = 0.025). MMP-9 was reported to be a novel predictor of CAD mortality based on a study involving 1127 patients with documented CAD (Blankenberg et al. 2003); the results showed that 97 patients who died from cardiovascular causes had a median concentration of serum MMP-9 of 62.2 ng/mL as compared to 47.8 ng/mL for those who did not have a fatal cardiovascular event (p = 0.0001). MMP-9 and TIMP-1 concentrations were found to decrease and increase, respectively, across the coronary sinus (Inokubo et al. 2001).
In in-vivo studies using apolipoprotein E-deficient mice, over-expression of activated MMP-9 by tissue macrophages enhanced the incidence of plaque rupture (Gough et al. 2006). In direct contrast, increased plaque rupturing was observed in apoE/MMP-9 double knockout mice, suggesting that MMP-9 might normally play a protective role in plaque stabilization (Johnson et al. 2005). The same study demonstrated that MMP-12 might contribute to atherosclerotic lesion expansion and destabilization, as lesion size was reduced in apoE/MMP-12double knockout mice compared to controls. Given the size of the MMP family, their overlapping substrates and numerous inhibitors, it is challenging to clarify the role of individual MMPs in the disease process.
Critical issues in the analysis of MMPs
The critical issues when developing analytical tools for biomarker analysis include sample integrity, and dealing with different dimensions of complexity (i.e. complexity determined by the tissue origin, complexity due to the number of biomarkers that are being monitored to determine the progression of disease). It is especially important to take note of the methods and conditions under which the sample is collected. As shown in Table 2, there was a large variability in the MMP concentrations reported in the literature. Most of the researchers took samples either from blood plasma or serum. Some recommended heparin plasma to study MMPs (Jung et al. 1996; Yamanaka et al. 2000; Jung et al. 2001; Mannello, 2003; Meisser et al. 2005) because clot activator appears to release MMPs from cells. Others have found that buffered citrate is the anticoagulant of choice because it slows down the release of MMPs by blood cells, and thus is not as dependent upon time as other anticoagulants (Coussens et al. 2002; Snoek-van Beurden, 2005; Thrailkill et al. 2005). The effect of blood collection methods on the concentration of MMPs was investigated by Manello who reported 2- to 10-fold higher concentrations of MMP-9 in serum than in heparin- and EDTA-plasma, and found that plastic tubes with a silica gel coated surface, as clot activator, gave even a 2-fold higher concentrations for serum samples (Manello, 2003). Nonetheless, he recommended heparin-plasma because platelet activation and neutrophil mobilization during clotting could produce such differences. Furthermore, he hypothesized that some of the disparity resulted from different storage temperatures when the blood was allowed to clot and the time at which the samples were centrifuged (Mannello et al. 2003). Regarding the storage temperature of human samples, it was reported that after 2 years storage at −80°C the MMP-9 concentration in citrate-plasma dropped by 65%, and it was estimated that only 1% would remain after 43 months (Rouy et al. 2005). In summary, the concentration of MMPs detected depends greatly on pre-analytical conditions of the blood samples, so great care should be exercised in keeping conditions constant for sample preparation. Additionally, it is also essential to note whether it is desired to measure latent, active, MMP-TIMP complexes, or both latent and active MMPs, because different techniques account for different forms of MMP. Other analytical challenges include the fact that most MMPs are regulated by post-translational mechanisms that make it difficult to identify them with conventional genomic and proteomic tools, and that MMPs are inhibited by endogenous binding proteins such as TIMPs.
Multiplexed profiling of MMPs in biological samples
Although MMP gene profiling conducted to date is most extensive for cancer, several studies have identified genetic variants associated with cardiovascular disease (Ye, 2006). Flex et al. (2007) found that MMP-1 and MMP-3 gene polymorphisms were significantly associated with peripheral arterial occlusive disease (Flex et al. 2007), and a specific MMP-3 polymorphism was strongly associated with acute myocardial infarction (Abilleira et al. 2006). While MMP genotype profiling may not be the best indicator of disease state, it is possible that polymorphism analysis can help identify patients at high risk or those who are more likely to respond to various therapeutics.
A much better indication of current disease state may be ascertained through detection of specific molecules and protein expression profiles of patient samples. However, these data are typically obtained by enzyme immunoassays and mass spectrometry, respectively, which are very tedious processes, despite the latest developments in both technologies. The feasibility of using these methods to monitor MMPs as biomarkers is further complicated by the requirement for specific antibodies-one for capture and one for detection in enzyme immunoassays, and mass spectrometry is not sensitive enough to detect individual proteins that span over fourteen orders of magnitude in concentration in the human serum or plasma. Furthermore, cardiovascular disease likely results when multiple molecules or systems have gone awry, thus such analytical tools that can provide that information reliably and fast would need to be developed.
Currently, analytical methods for the determination of MMPs in biological samples include ELISA (enzyme-linked immunosorbent assays, including multiplexed ELISAs in the form of antibody arrays); zymography; optical methods such as near-IR (infrared) optical imaging, fluorescence, and surface plasmon resonance spectroscopy; the use of active-site probes followed by enzymatic digestion of the captured MMPs, and LC-MS/MS (liquid chromatography-mass spectroscopy) analysis of the digested MMPs.
Enzyme Immunoassays
Of the currently available methods for MMP detection, there are several commercially available ELISA kits (e.g. Biotrack, Fluorokine Multianalyte Profiling and Quantikine HS from R&D Systems), as well as a Luminex assay (Thrailkill et al. 2005), but these assays yield imprecise and potentially misleading results because they cannot distinguish between the active and latent forms of the enzyme or between specific MMPs and their TIMP complexes (Zucker et al. 1994; Maeda et al. 1995; Brennan et al. 1997; Torii et al. 1997; Yamanaka et al. 2000; Koulentaki et al. 2002; Masuhara et al. 2002; Dandona et al. 2003; Watanabe et al. 2005, Nilsson et al. 2006). Sandwich ELISAs do have greater sensitivity, but they require two different antibodies for each individual MMP, and a separate assay plate must be used for the measurement of each MMP, making for a very time consuming assay. The detection limits for Quantikine HS from R&D Systems (Nilsson et al. 2006) vary greatly for each MMP (i.e. MMP-2 is 0.37 ng/mL; MMP-3 is 2.35 ng/mL and MMP-7 is 0.09 ng/mL). Multiplexed ELISAs have been reported, but these tend to exhibit high cross-reactivity among various MMPs because these proteins share common domains. A 10 × 10 bead matrix (Luminex Corporation) in which each bead contains a specific proportion of a red/orange fluorescent dye was reported for analysis of MMP-1, -2, -3, -8, and -9 in plasma (Thrailkill et al. 2005). This assay allowed for simultaneous determination of each of these MMPs, thus eliminating any possible operator error in sampling and also reducing sample volume to only 10 μL for all five analytes. When compared to single analyte ELISA, this represented a five-fold reduction in sample volume and makes the assay more amenable to pediatric population (Thrailkill et al. 2005).
A prototype antibody array (Agilent Technologies) consisting of forty capture monoclonal antibodies to human serum proteins, including MMPs, TIMPs, and MMP-TIMP complexes was used to assess the differential protein levels between control (7 healthy subjects chosen randomly) and patient (11 subjects with a history of cardiovascular disease) plasma samples taken one hour after the treadmill experiment. Based on the resulting data (Fig. 2), proteins were then arranged in descending order starting with those having higher average levels in the patient group compared to control (above the red horizontal line in the heatmap). Proteins with relatively high average levels in both the control and patient group were placed near the bottom of this listing. Finally, proteins for which the control group had higher average levels than the patient group were ranked at the very bottom (below the red line). In each group, the proteins were further ordered by Gaussian error score so that the proteins with the most different distributions of protein levels between the two sample group are on the top. In the heatmap (Fig. 2A), each row represents color-coded protein levels for each of the thirty three target proteins, and each column represents all protein levels for one sample. Blue indicates the lowest protein level in a given row, and yellow indicates the highest level. Plots in Figure 2B show actual distributions of specific protein levels among the two groups for three proteins with most dramatic difference between the two groups. Below the plots, green x's show protein levels in control group, red x's show protein levels in Treadmill patient groups. Green and red graphs show corresponding normal fits. Among the most differentiating proteins are MMP-2/TIMP-2 complex, MMP-10, MMP-9, TIMP-1 and TIMP-2. Classification analysis using leave one out cross-validation algorithm showed that one can predict normal or disease status of the sample based on protein levels of top five proteins mentioned above. The data generated with this prototype array is unique because this allows simultaneous determination of thirty nine analytes, including several MMPs and MMP-TIMP complexes, requires only 10 μL sample volume, and the amount of antibody present on each feature of the array is 35 pg. This was possible because the arrays were printed using an inkjet deposition tool that delivers 35 pL on each spot with a deviation of only 1-2 μm for a spot of 50 μm in diameter. Although 60 monoclonal antibodies were deposited onto the array, the multiplexed assay targeted only thirty nine analytes because only thirty nine detection antibodies were working properly in the assay.

Zymography
This method is also widely used and identifies MMPs by the degradation of their preferential substrate and by their molecular weight (Snoek-van Beurden, 2005). All types of zymography are similar to gelatin zymography, the substrate simply differs depending on the MMP being analyzed. In zymography, the proteins are separated by electro-phoresis under denaturing (in the presence of SDS) and in nonreducing conditions; the separation is done in a polyacrylamide gel that contains the specific substrate. After electrophoresis, the gel is washed to remove SDS, following which the MMPs partially renature and recover their activity. The gel is then stained blue, and the MMPs are visible as clear bands against the blue substrate in the background and can be measured by densitom-etry (Snoek-van Beurden, 2005). Proenzymes and active forms of MMPs can be distinguished by molecular weight. An added benefit of this method is that it also results in separation of MMP-TIMP complexes (Snoek-van Beurden, 2005). Gelatin zymography is extremely sensitive for gelatinases like MMP-2 and MMP-9 (i.e. 10 pg of MMP-2 can be easily detected) while casein zymography is more suitable for stromelysins MMP-1, -7, -11, -12, and -13. Collagen zymography is also used for detection of MMP-1, -2, -9, and -13. In situ zymography, which can detect only active MMPs, allows the localization of MMPs in tissue sections and uses a substrate that is deposited on or under a frozen section of an unfixed tissue sample. In these assays it is important to use control slides with appropriate MMP inhibitors, to discriminate between the different classes of MMPs (Snoek-van Beurden, 2005).
Optical Methods
Near-IR optical imaging has been used primarily for detection of MMPs in tumor tissue (Bremer et al. 2001). An optical contrast agent was developed that was highly activatable by MMP-2-induced conversion. Signal characteristics of the probe were measured ex vivo with a recombinant enzyme. Animal tumor models were established with MMP-2-positive (human fibrosarcoma cell line, n = 4) and MMP-2-negative (well-differentiated mammary adenocarcinoma, n = 4) tumor cell lines. Both tumors were implanted into nude mice and were optically imaged after intravenous administration of the MMP-2-sensitive probe (Bremer et al. 2001).
Another optical method, which measures only the MMPs activity and has been used to evaluate MMP inhibitors, is a collagenase assay using capillary gel electrophoresis (CGE) and laser-induced fluorescence detection or LIF (Sano et al. 2004). This collagenase assay employs dynamic fluorescent labeling with NanoOrange dye, and measures fragments of type I and II collagen, which are produced from the native collagen by cleavage with a particular MMP. The CGE-LIF with a capillary array system is ideal as a high throughput system. An assay for detection of both the proform and the active form of MMP-2 with a surface plasmon resonance (SPR) spectrometer, was reported by Pieper-Furst et al. 2004. This assay involves mixing of the MMP-2 with anti-MMP-2 antibody adsorbed to colloidal gold and injection into the flowcell of the SPR where TIMP-2 is immobilized. TIMP-2 has at least two binding sites for MMP-2; when the antibody binds the MMP-2 first, one site in MMP-2 is occupied by the antibody, leaving a second site for TIMP. The detection limit for the SPR assay corresponds to 36 pg/mL and takes only 30 min including the regeneration of the sensor (Pieper-Furst et al. 2004). The SPR assay is not applicable to the detection of the MMP-TIMP complex since TIMP is one of the ligands.
Mass Spectrometry
Identification of MMPs and other metalloprote-ases with a set of alkyne-tagged hydroxamate-benzophenone (HxBPyne) probes was reported recently (Sieber et al. 2006), and it involves enzymatic digestion after capture of the MMPs using biotin-azide tags and avidin chromatography followed by analysis of the MMP-digests by LC-MS/ MS. The LC-MS/MS method is very lengthy, and it is still not fully reliable when it comes to identifying proteins in the digested form, because the identification often relies on the presence of only a few peptides from the digested protein. Nonetheless, analysis of cell and tissue proteomes treated with an optimal set of probes (i.e. HxBPyne probes) resulted in the identification of more than twenty different metalloproteases (i.e. MMP-1, MMP-3, MMP-7, MMP-12 at detection limits of 0.5, 2.5, 0.25, and 1 μg/mL of proteome, respectively).
Is there a need for new tools to profile MMPs?
Although TIMP-1 and TIMP-2 were considered initially as good inhibitors of MMPs in cancer progression, it turned out that small molecule inhibitors containing both hydroxamate and non-hydroxamate binding sites were more attractive and five different inhibitors (i.e. Marimastat, Prinomastat, Tanomastat, BMS 275291, and Neo-vastat) made it into advanced stages of the clinical development with disappointing results. The rapidity with which these inhibitors moved into clinical trials raised a very important question: “Do any MMPs play an important role in advanced lung and pancreatic cancer, and if so, which ones?” (Coussens et al. 2002). Because there is no analytical tool that can answer this question for all twenty-four MMPs measured at the same time, what is really needed is a better tool for multiplexed profiling of this important class of proteins. The data included here for the prototype antibody array demonstrate that MMPs can be reliably measured at ng/mL concentrations in human serum, but this array is not yet available commercially and the authors are not aware of any commercial multiplexed ELISA kits that can measure all twenty-four MMPs simultaneously.
One potentially very promising analytical method for profiling all MMPs in a sample under development in the author's laboratory uses two separation steps: immunoaffinity chromatography using an anti-MMP catalytic domain antibody to “fish out” the MMPs from the tissue (i.e. human serum or plasma) and high- pressure liquid chromatography (HPLC) to separate the MMPs as individual proteins (see Fig. 3). These separations are followed by a detection step in which the metal ion (i.e. zinc at mass-to-charge of 66 amu identified here as 66Zn) is measured by inductively-coupled plasma mass spectrometry (ICPMS). The 66Zn scan is specific only to MMPs because they are isolated from the sample (i.e. human serum) by immunoaffinity chromatography with an anti-MMP antibody. This technique is very powerful in the sense that it can profile all MMPs in one single analysis that can be accomplished in thirty minutes or less. In addition, it is both very sensitive and selective (i.e. compare the UV scan and the ICPMS scans in Figure 3) because it will focus specifically on the metal ion that is at the heart of the protein functionality. For details on the use of HPLC-ICPMS for the identification of metalloproteins, the reader may refer to a paper on determination of ceruloplasmin (Cp), a copper protein, in human serum published by Lopez-Avila et al. 2006 in which the identification of Cp is based on the retention time match of the unknown in the serum sample with the Cp external standard and the presence of 63Cu and 65Cu at a ratio of 2.2 ± 0.1.
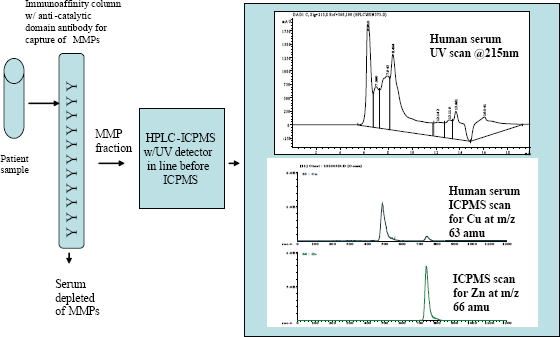
Multiplexed analysis of MMPs in biological samples by immunoaffinity chromatography followed by HPLC-ICPMS is a new concept and has not been reported in the literature for this class of proteins. In this case, MMPs are identified by retention time match with specific MMP standards and by the zinc scan, which is done using the ICPMS system. Because MMPs are “fished-out” from the sample with a reagent that is quite specific for MMPs (i.e. antibodies that recognize the catalytic domain, which is common in all MMPs), this method will generate an MMP fingerprint that can help identify the disease, track disease progression, and help evaluate how therapeutic agents can alter the MMP fingerprint. An MMP fingerprint analyzing levels of all MMPs is crucial, especially because it has been reported that MMP-7, -8, -9, and -13 can be inactivated by MMP-3 (Samnegard et al. 2006).
There are difficulties in implementing such approach because many of the MMPs are not available commercially as intact proteins but as recom-binant fragments, and they will need to be expressed and purified for such developmental work to be accomplished.
Summary
MMPs are an important class of enzymes that play roles in both normal physiology and several disease states. Efforts to clarify the importance of particular MMPs in specific pathologies have been hampered by the complex biology of this large family of related enzymes and their TIMP binding partners. In addition, accurate quantitation of multiple of MMPs in patient samples is confounded by limitations of the currently available detection methods. With the further development and commercialization of the HPLC-ICPMS technique, it seems likely that multiple MMPs levels will be able to be more easily monitored and correlated with various aspects of human disease.
Footnotes
Acknowledgements
The authors acknowledge the contributions of Amber Hess (summer intern) who helped with some of the literature search and Anya Tsalenko of Agilent Technologies who did the computational analysis of the protein array data for selected MMPs.
