Abstract
This study examined the contribution of the osteoarthritis (OA) susceptibility genes ASPN, GDF5, DIO2, and the 7q22 region to the development of radiographic knee OA in patients with a mean age of 40.6 ± 7.9 years (standard deviation) and who suffered from nonacute knee complaints a decade earlier. Dose-response associations of four single nucleotide polymorphisms(SNPs) in the susceptibility genes were determined by comparing 36 patients who showed the development of OA on radiographs (Kellgren and Lawrence score ≥1) with 88 patients having normal cartilage with no development of OA on radiographs. Multivariate logistic regression analysis including the variables such as age, gender, body mass index, and reported knee trauma was performed. A dose-response association of DIO2 SNP rs225014: odds ratio (OR) 2.3, 95% confidence interval (CI) 1.1–4.5 (P = 0.019) and GDF5 SNP rs143383: OR 2.0, 95% CI 1.1–3.8 (P = 0.031) was observed with knee OA development. The ASPN and 7q22 SNPs were not associated with OA development.
Introduction
Osteoarthritis (OA) of the knee is a common cause of musculoskeletal disability and is characterized by late-onset degeneration of articular cartilage, which is marked by the recapitulation of OA articular chondrocytes to a growth plate morphology and signaling and breakdown of matrix proteins. 1 This leads to the development of fibrillations, fissures, and ulcerations at the articular cartilage surface, which could be most sensitively detected by magnetic resonance (MR) imaging. 2 Large-scale genetic studies performed to detect the common underlying pathways that affect susceptibility have revealed a considerable number of robust OA susceptibility gene products that appear to be active in a shared pathway involving the developmental process of endochondral ossification.3,4 Several distinct studies on OA development have demonstrated the replication ability of the susceptibility genes ASPN, 5 GDF5, 6 DIO2, 3 and the 7q22 locus, 7 which are considered to be consistent early OA susceptibility signals, and additional functional follow-up data have further provided insight into the underlying disease mechanisms.8–11 Progression of the disease could further result in the reactivation of genes involved in endochondral ossification, leading to the loss and mineralization of articular cartilage, a process known to contribute to OA.3,12
The aim of this study was to examine the contribution of the risk alleles of ASPN, GDF5, DIO2, and the top SNP of the 7q22 locus to the development of radiographic knee OA in a relatively young study population that presented with nonacute knee symptoms a decade earlier by comparing patients with radiographic OA development and patients without signs of OA in the knee on radiographs and MR images.
Methods
Study population
Study design
Case–control, level of evidence: 3. This study was approved by the Leiden University Medical Centre Medical Ethics Review Board, and written informed consent was obtained from each participant. This research complied with the principles of the Declaration of Helsinki. This study is a follow-up of a trial performed 10 years ago. The initial study consisted of 856 patients (mean age 31 ± 8.0 years standard deviation [SD]) with nonacute knee complaints, defined as persistent knee complaints, such as pain, swelling, and instability, lasting for more than four weeks. 13 After 10 ± 0.90 years (SD), all 856 patients of the initial study were contacted and invited for follow-up, 14 which identified 326 eligible participants. All the 326 eligible participants were contacted and a saliva collection container was sent for DNA extraction. Eventually, 217 patients (67%) were included. Of the 109 lost subjects, 21 patients refused to participate and 88 did not respond, despite the contact letter sent by mail, a second letter sent after one month, and three contact attempts by telephone. MR images and radiographs of the initial symptomatic knee were taken at inclusion and after 10 years. The presence of detectable OA features on radiographs was compared between the baseline and follow-up images. None of the patients showed radiographic knee OA at baseline. Due to different scanning techniques and other scoring methods used at baseline and after 10-year follow-up, the MR outcomes from baseline and follow-up could not be accurately compared to assess the development of OA. Therefore, a certain degree of OA development in those patients without radiographic OA development but with cartilage defects visible on MR images could not be ruled out. In order to compare patients with radiographic OA development to a control group without any signs of OA development, only patients without cartilage defects visible on MR images were used as controls. Ultimately, a total of 124 (15%) patients were included in this study (Fig. 1).

Flowchart response to follow-up.
Radiographic knee examination and assessment
Standardized weight-bearing posterior-anterior knee radiographs next to supine lateral radiographs of the knee were made at baseline and 10-year follow-up. At baseline, one of the six musculoskeletal radiologists with at least four years of experience scored the radiographs for overall severity of OA using the Kellgren and Lawrence (K&L) system. 15 The follow-up radiographs were scored by an experienced musculoskeletal radiologist and a research fellow using the same K&L scoring method. 14 Individual development of OA was obtained by comparing the baseline K&L score to the follow-up K&L scores. Development of OA was considered to be present at a K&L score of 1 point or more.
Knee MR imaging
MR imaging examinations of the initially affected knee were performed after 10 years on a 3T system (Achieva 3T, Philips Medical Systems). Due to different MR scanning techniques used at baseline and 10-year follow-up, only the follow-up scans were used to assess cartilage defects. Focal cartilage defects were defined as an abrupt transition between the defect and the surrounding cartilage, and a diffuse cartilage defect was defined as a gradual transition between normal and thinned cartilage. 16 Cartilage defects outcome scores were used in a binary (absent vs present) fashion.
DNA
DNA was obtained from patients using saliva collecting containers (Oragene OG-250, DNA Genotek). Recent studies have shown that the use of saliva samples is a good alternative to blood samples to obtain genomic DNA of high quality. 17
The containers were sent to all the 326 eligible patients, including instructions for use. The patients were asked to wash their mouth once with water and to wait for at least 30 seconds. Then, they were asked to spit into the white container, cap the container with the blue lid, and finally shake the sample gently. The saliva samples were stored according to the manufacturer's instructions at room temperature until DNA extraction. On an average, the saliva samples were stored for 1.5 months.
DNA was extracted from the saliva samples using the Oragene kit (DNA Genotek) as described by the manufacturer. The saliva samples were incubated at 50 °C for two hours, after which 0.5 mL of the samples was transferred to a 1.5 mL Eppendorf tube. Then, 20 μL of Oragene purifier was added, and the sample was mixed by inversion and incubated on ice for 10 minutes. The samples were then centrifuged in a small, bench-top microfuge for three minutes at 18,000 × g at room temperature. The supernatant was transferred to a new tube and 0.5 mL of 95% ethanol was added, and the contents were mixed by inversion at least five times and incubated for 10 minutes at room temperature. The samples were again centrifuged in a small, bench-top microfuge for one minute at 18,000 × g at room temperature The resulting supernatants were discarded, and the DNA was dissolved in 100 μL of TE buffer (10 mmol/L Tris-HCl, 0.1 mmol/L ethylenediaminetetraacetic acid [pH 8.0]) and quantified. The DNA samples were stored at 4 °C until polymerase chain reaction (PCR) analysis.
For the candidate gene association study, we selected four SNPs in four candidate genes ASPN, GDF5, DIO2, and the 7q22 locus. Several distinct studies on primary OA have shown the replication ability of these susceptibility genes. 8 In this study, we hypothesize that these susceptibility genes already contribute to the development of radiographic knee OA in relatively young patients suffering from knee complaints. The P-values of Hardy–Weinberg test were 0.385 for DIO2, 0.916 for GDF5, 0.033 for ASPN, and 0.242 for the 7q22 locus. All genotypes were included in the equation. The four candidate genes selected for this study are described as follows.
DIO2
Iodothyronine deiodinase enzyme type 2 (DIO2) is a regulator of thyroid hormone metabolism in the growth plate, where the thyroid hormone triggers the terminal maturation of growth plate chondrocytes. 8 DIO2 has recently been shown to be a susceptibility gene for primary OA.3,18 In a genome-wide linkage scan and association analysis, an association was found between OA and the minor C-allele of SNP rs225014. 3 We hypothesize that the DIO2 gene also contributes to the development of radiographic knee OA in the current study population.
ASPN
Asporin (ASPN) inhibits transforming growth factor-β, which has a crucial role in the development and homeostasis of cartilage. 19 ASPN is expressed at low levels in normal cartilage, but expressed abundantly in OA articular cartilage. Due to polymorphisms in the ASPN gene, variant ASPN proteins arise with a variable number of aspartic acid (D) repeats in the amino-terminal end of the protein. 20 The T-allele of SNP rs13301537 is associated with variants of the ASPN gene encoding 13 (D13) and 14 (D14) aspartic acid repeats. 21 D14 was shown to be associated with knee OA in a Japanese and Han Chinese population, whereas D13 showed an association with a decreased risk of knee OA in the Caucasian population.21,22
GDF5
Growth differentiation factor 5 (GDF5) is a bone morphogenetic protein involved in the early development of joints in the embryonic tissues and is expressed in the synovial joint tissues throughout life. 23 The SNP rs143383 is the most widely replicated genetic association with knee OA, with the risk-associated T-allele showing reduced expression relative to the C-allele in OA. 6 The GDF5 rs143383 polymorphism is associated with knee OA in both Asians and Caucasians.6,24
7q22 locus
Conserved oligomeric Golgi complex subunit 5 (COG5) was recently discovered in a genome-wide association study as a novel gene involved in OA. The C-allele of SNP rs3815148 was associated with an increased risk of knee OA. 7
SNP genotyping
Genotyping was carried out in 217 samples at OA susceptibility SNPs from the four genes ASPN, GDF5, DIO2, and the 7q22 locus. The selected SNPs were fit in a Sequenom multiplex assay designed by the Assay Designer software version 3.1 (Sequenom). The SNPs were genotyped by mass spectrometry (the homogeneous MassARRAY system; Sequenom) using standard conditions. PCR reactions were carried out in a final volume of 5 μL containing 2.5 ng of genomic DNA. Genotypes were assigned using Genotyper version 3.0 software (Sequenom). The genotyping success rate was 98.02% ± 0.75% (SD). Internal genotyping controls were included with a concordance rate of 100%. All the genotype frequencies for the tested SNPs were in Hardy–Weinberg equilibrium (Table 1).
Multivariate regression analysis outcomes of OA development.
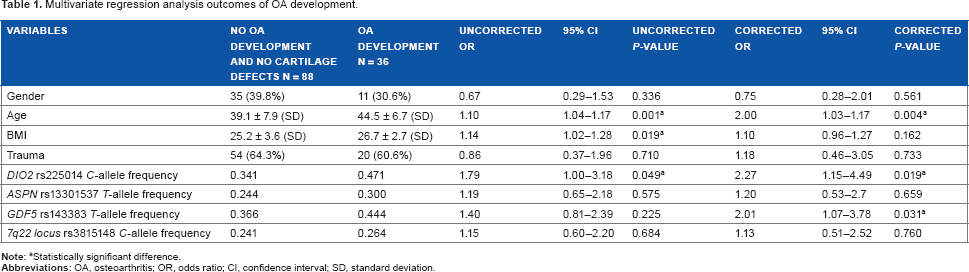
Statistically significant difference.
Statistical methods
A case–control approach was also used to assess the association between the susceptible risk allele frequencies of the seven selected SNPs in patients with knee OA development and in patients without the development of radiographic knee OA. Radiographic OA development was defined as a K&L score of ≥1. Patients who did not show the development of radiographic OA but who did have cartilage defects detected on the follow-up MR images were excluded from further analysis (Fig. 1). A logistic regression model was fitted to measure the strength of the association with the different genotypes (dose response), which is expressed as odds ratios (ORs) with 95% confidence intervals (95% CIs). In these analyses, we adjusted for age, gender, body mass index (BMI), and reported knee trauma. P-values < 0.05 were considered to indicate statistical significance. In order to assess the discriminating power of the different genotypes studied, we generated receiving operator characteristics (ROCs) using knee development as outcomes and age, sex, BMI, and trauma as conventional risk factors.
Results
Population characteristics
Of the 124 patients included in this study, five patients (4%) were of non-Caucasian descent. At follow-up, the mean age of the 124 patients included for the association study was 40.6 ± 7.9 years (SD), 46 (37.1%) of them were female, and the average BMI was 25.6 ± 3.5 (SD).
OA development
Of the 36 patients who developed radiographic knee OA after 10 years of follow-up, 27 patients had a K&L score of 1 and 9 had a K&L score of 2. Of the 172 patients without radiographic OA development, 84 (48.8%) patients showed cartilage defects detected on MR images and 88 (51.2%) did not. The latter patients were used as controls. Patients with development of radiographic knee OA were significantly older and had significantly higher BMIs than the controls without OA development and without cartilage lesions (Table 1).
As shown in Table 1, the minor C-allele frequency of the SNP rs225014 in the control group without OA development and without cartilage lesions was 0.341, whereas in the study group with OA development, the minor C-allele frequency was 0.471. There was a significant association between the presence of the minor C-allele of SNP rs225014 of the DIO2 gene and the development of radiographic knee OA (OR 1.8, 95% CI 1.00–3.18; P-value 0.049). This effect increased when corrected for age, BMI, and trauma (OR 2.3, 95% CI 1.14–4.49; P-value 0.019). The GDF5 T-allele frequency of the SNP rs143383 was 0.366 in the control group without OA development and cartilage lesions, while it was 0.444 in the OA development group. When corrected for age, BMI, and trauma, there was a significant association between the presence of the T-allele of SNP rs143383 of the GDF5 gene and OA development (OR 2.0, 95% CI 1.07–3.78, P-value 0.031). The OA susceptibility SNP in the ASPN and at the 7q22 locus did not show any association (Table 1).
Finally, we investigated whether the genes associated with increasing aspects of joint destruction as measured by radiographs and/or MR imaging characteristics after 10 years as defined by subjects without radiographic OA nor cartilage lesions visible on MR images (N = 88), no radiographic signs of OA but with cartilage defects visible on MR images (N = 84), and radiographic OA with cartilage defects visible on MR images (N = 36). As shown in Table 2, DIO2 showed a significant association (P-value 0.020) with increasing signs of cartilage destruction visible on both radiographs and MR images.
Multivariate regression analysis outcomes of OA development and cartilage lesions visible on MR images.

Statistically significant difference.
Increased risk prediction of DIO2 and GDF5 genotypes
To investigate the possibility to use the DIO2 and GDF5 genotypes as a predictive tool, we next performed an ROC analysis. After considering nonacute knee symptoms as an outcome, two models were fitted with knee OA development. Model one consisted of only age, gender, BMI, and trauma, while model two included age, gender, BMI, trauma, and DIO2 and GDF5 genotypes. The predictive value of the anthropometric traits alone as reflected by the area under the curve (AUC) was 0.647 (95% CI 0.530–0.764); however, when genotypes of the DIO2 and GFD5 genes were added, the AUC improved to 0.697 (95% CI 0.582–0.812; Fig. 2).

Receiver operating characteristics (ROC) analysis diagrams. The outcome is OA development a decade after knee complaints. (A) The model included age, gender, trauma, and BMI. The AUC is 0.647 (95% CI 0.530–0.764). (B) The model included age, gender, BMI, trauma, and DIO2 and GDF genotypes. The AUC is 0.697 (95% CI 0.582–0.812).
Discussion
In the present study, we showed that the DIO2 OA susceptibility SNP rs225014 and the GDF5 susceptibility rs143383 SNP are significantly associated with the development of knee OA (OR 2.3; 95% CI 1.14–4.49 and OR 2.0; 95% CI 1.07–3.78, respectively) in a relatively young patient group with a mean age of 44 years and with a history of knee complaints a decade ago. This effect appeared to be independent of other factors related to knee OA development, such as knee trauma, age, and BMI. Prediction of knee OA development improved from an average AUC of 0.647 for age, sex, BMI, and trauma alone to an AUC of 0.697 when including DIO2 and GDF5 genotypes in the risk prediction model, almost reaching a clinically relevant AUC value (≥0.7). Furthermore, an increased minor allele frequency of DIO2 is related to the signs of OA development visible on both radiographs and MR images (P-value 0.020); a similar trend is visible for GDF5, although not significant (P-value 0.206).
These data are in line with recently published data showing that articular cartilage expression of DIO2 is epigenetically regulated and that DIO2 rs225014 risk allele carriers, in particular, are less able to maintain cartilage homeostasis because of the fact that subtle changes in methylation, generally occurring due to environmental changes such as microtraumas, resulted in detrimental upregulation of DIO2. 9 Type II deiodinase (D2), expressed by the DIO2 gene, regulates the bioavailability of intracellular T3 in specific tissues such as the growth plate and facilitates terminal maturation of hypertrophic chondrocytes. Functional genomic studies showed high expression of DIO2 mRNA and D2 protein levels in osteoarthritic cartilage compared to healthy cartilage. 18 Furthermore, assessment of DIO2 allelic imbalance showed that the OA risk allele “C” was more abundantly present in articular joint tissues than the wild-type allele “T.” 18 Upregulation of DIO2 expression in a human in vitro model resulted in a marked reduction of the capacity of chondrocytes to deposit the extracellular matrix (ECM) components, including type II and type X collagen, while inducing OA-specific markers of cartilage matrix degeneration and mineralization. 9 In mice undergoing a forced running regime, it has been shown that DIO2 deficiency has a protective effect on the homeostasis of articular cartilage in the knee joints. 25
It is possible that trauma at a relatively early age affects the propensity of the highly specialized, maturational arrested articular chondrocytes to lose their maturational arrested state loss of epigenic control, among others, of the DIO2 gene.
Our results show that the T-allele of the GDF5 SNP rs143383 is significantly associated with the development of knee OA with an OR of 2.0 (95% CI 1.07–3.78). This is in line with earlier findings that GDF5 rs143383 polymorphism is associated with knee OA in both Asians and Caucasians.6,24,26 Recent research showed that GDF5 stimulation of human chondrocytes inhibits the expression of cartilage ECM-degrading enzymes MMP13 and ADAMTS4 and stimulates the expression of cartilage anabolic genes ACAN and SOX9. 27 GDF5 stimulation also inhibits the canonical Wnt signaling pathway through the expression of the DKK1 and FRZB inhibitors. The Wnt signaling pathway plays an important role during cartilage development, and activation of the pathway in the adult cartilage tissue leads to hypertrophy, initiation of calcification, and tissue degradation via increased expression of ECM-degrading components. 27 Therefore, altered expression of the GDF5 gene may result in ECM degradation and decrease of cartilage quality, which may induce the development of OA.
It is commonly accepted that a clinically useful diagnostic marker should have an AUC of ≥0.7. 28 Adding DIO2 and GDF5 genotypes in the model resulted in a significant increase of the AUC from 0.647 to 0.697. The latter underscores the fact that although patient characteristics and environmental factors are important, genetic factors play a substantial role in the development of knee OA.
None of the investigated SNPs of the ASPN and 7q22 locus genes were significantly associated with OA development in the current study population. These findings are in line with the outcomes of recent studies, which report conflicting evidence about these genes in the development of OA29,30
In total, five patients of non-Caucasian descent were included in this study; however, upon discarding these patients, DIO2 and GDF5 outcomes remained the same, indicating that these patients did not drive our associations and possibly that these genes confer risk to OA development in both Caucasian and non-Caucasian populations as previously shown.3,6,10,11,18,31,32
Several limitations of the study should be mentioned. Due to different scanning techniques and other scoring methods used at baseline and after 10-year follow-up, the MR imaging outcomes from baseline and follow-up could not be accurately compared to assess the development of OA. Therefore, a certain degree of development in those patients without radiographic OA but with cartilage defects visible on MR images could not be ruled out. In order to compare patients with radiographic OA development to a control group without any signs of OA development, only patients without cartilage defects visible on MR images were used as controls. This restriction resulted in the exclusion of 43% of the initial study group, leading to a significantly smaller study population. Being aware of the tendency of association studies to produce false-positive results, an additional replication is necessary. In this study, gender differences were not related to OA development. This may be explained by the fact that the female participants in this relatively young study population were mostly premenopausal.
Conclusion
The presence of the minor C-allele of the DIO2 SNP rs225014 and the T-allele of the GDF5 SNP rs143383 is associated with the development of OA in a relatively young study population 10 years after knee complaints. Subsequent ROC analyses showed that determining DIO2 and GDF5 genotypes significantly improves risk prediction toward clinically relevant values.
Author Contributions
Conceived and designed the experiments: KH, PV, JB, RN, IM. Analyzed the data: KH, PV, RN, IM. Wrote the first draft of the manuscript: KH, PV, IM. Contributed to the writing of the manuscript: KH, PV, JB, RN, IM. Agreed with manuscript results and conclusions: KH, PV, JB, RN, IM. Jointly developed the structure and arguments for the paper: KH, PV, JB, RN, IM. Made critical revisions and approved the final version: KH, PV, JB, RN, IM. All the authors reviewed and approved the final manuscript.
