Abstract
MicroRNAs (miRNAs) are short sequences of noncoding single-stranded RNAs that exhibit inhibitory effects on complementary target mRNAs. Recently, it has been discovered that certain viruses express their own miRNAs, while other viruses activate the transcription of cellular miRNAs for their own benefit. This review summarizes the viral and/or cellular miRNAs that are transcribed during infection, with a focus on the biomarker and therapeutic potential of miRNAs (or their antagomirs). Several human viruses of clinical importance are discussed, namely, herpesviruses, polyomaviruses, hepatitis B virus, hepatitis C virus, human papillomavirus, and human immunodeficiency virus.
Introduction
Several types of small noncoding RNAs have been discovered that affect a multitude of biological pathways within the cell. One such class includes microRNAs (miRNAs), short sequences of noncoding single-stranded RNAs that exhibit inhibitory effects on complementary target mRNAs. Research on miRNAs can provide insights into the development, treatment, and monitoring of diseases, including viral diseases. This review aims to provide an overview of recent research characterizing the role of miRNAs during viral infection. We focus on several viruses of clinical importance in order to assess the potential of host- and virus-derived miRNAs as therapeutic targets or biomarkers of disease.
A
Biogenesis of miRNAs
Several excellent reviews are available that explain the details of miRNA biogenesis.2,3 Most miRNAs are transcribed through the actions of RNA polymerase II from templates found within introns of protein-coding genes or directly from independent genes.3,4 In the cytoplasm, the miRNA guide strand remains associated with Argonaute (Ago) within the RNA-induced silencing complex (RISC), while the complementary strand, referred to as the miRNA* (star strand) or passenger strand, is degraded. Unlike most cellular miRNAs, certain viral miRNAs can be derived from both strands of the double-stranded miRNA molecule, leading to the convention of naming the strands with −5p or −3p suffixes.
The guide miRNA is the primary mechanism for targeting the RISC complex to mRNAs. Although miRNAs are ~22 nucleotides in length, the mRNA target is generally recognized through complementary base pairing of the seed sequence comprising nucleotides 2–7 at the 5′ end of the miRNA strand. 5 Consequently, one miRNA:RISC complex can silence hundreds of mRNAs with complementarity to the same seed sequence, regardless of their translational products. Once bound by the RISC, miRNAs usually target the 3′-untranslated region (UTR) of mRNAs, 6 resulting in the repression of translation and/or degradation of the target mRNA.3,7 It is now well established that miRNAs influence an extensive number of biological pathways in this manner through regulation of protein-coding genes.
Techniques for miRNA Identification and Verification
Host- and virus-derived miRNAs can be identified computationally using bioinformatics or through functional screening assays. A variety of bioinformatics programs are now available that predict miRNAs (Table 1). Although bioinformatics has been extremely valuable in bringing together possible sets of miRNAs and their targets, the advent of relatively inexpensive next-generation sequencing (deep sequencing) platforms and high-density oligonucleotide arrays has simplified the functional screening of viral miRNAs. Most viral miRNAs were initially identified through a modified rapid amplification of cDNA ends protocol. Briefy, poly-acrylamide gel-purified small RNAs were modified with 5′ or 3′ oligonucleotides that functioned as primers for PCR after reverse transcription and the amplified segments were cloned and sequenced. Next-generation sequencing techniques have since then replaced the time-consuming process of cloning and can process millions of sequence reads in parallel. An elegant use of this technique to identify biologically relevant miRNAs is termed
Approaches used by miRNA target prediction software tools.

Following their identification through bioinformatics or high-throughput techniques, putative miRNAs and their targets must be experimentally validated. Luciferase reporter assays are commonly used to verify direct binding of a miRNA to a particular mRNA sequence. The 3′-UTR region predicted to interact with a miRNA target is placed downstream of the luciferase gene in a reporter construct. The vector is then transfected into cells alongside an miRNA expression vector or a control vector, and luciferase expression is reduced if the miRNA targets the MRE. Site directed mutagenesis or target protector nucleotides can also be used to verify MRE sequence complementarity to the miRNA. Antisense oligonucleotides (molecular sponges) show specificity of a particular miRNA seed sequence for an MRE by complementary binding to the miRNA sequence. 11 Viral miRNA deletion mutants are also used to verify the biological function of individual or clusters of miRNAs.
Biomarker and Therapeutic Potential of miRNAs during Viral Infection
The majority of known virally encoded miRNAs are found within DNA viruses that replicate in the nucleus. The first viral miRNAs discovered were found to be expressed by Epstein-Barr virus (EBV). 12 Since then, the great majority of miRNAs have been discovered in herpesviruses. A few viral miRNAs have also been reported in other human DNA virus families, including polyomaviruses (described below) and possibly adenoviruses (in very low abundance from virus-associated RNAs). In addition, miRNAs have been reported in DNA viral families that do not infect humans, namely, asco-viruses (Heliothis virescens ascovirus), baculoviruses (Bombyx mori nuclear polyhedrosis virus), and nimaviruses (white spot syndrome virus). Notably, miRNAs from human papillomaviruses (HPVs) have yet to be definitively discovered.
Retroviruses possess a replication stage that involves nuclear DNA as a result of reverse transcription, and miRNAs have indeed been discovered in a few retroviruses, including avian leucosis virus subgroup J, African green monkey simian foamy virus, and bovine leukemia virus.13,14 As will be discussed later, miRNAs have been reported from the human immunodeficiency virus (HIV)-1 transactivation response (TAR) RNA of CD4 T cells,15–17 although other studies have failed to find biologically relevant viral miRNAs from HIV-1-or human T-lymphotropic virus-infected cells.18–20
The existence of miRNAs in cytoplasmic RNA viruses has yet to be detected, and several reasons have been proposed as to why they are unlikely in cytoplasmic viruses. The nuclear location of Drosha and DGCR8, necessary for genesis of pre-miRNAs, is one of the leading explanations for this absence. Interestingly, Rouha et al engineered a nuclear-replicating RNA virus to contain a miRNA-precursor stem-loop sequence element within its RNA genome. They found that a functional miRNA was produced without affecting viral replication, 21 implying the possible existence of Drosha-independent miRNA generation pathways within the cytoplasm.
Herpesviruses
Herpesviruses are large, double-stranded DNA viruses that infect a range of invertebrate and vertebrate animals. Nine human herpesviruses (HHVs) exist, possessing genomes of ~125–230 kb in length. The great majority of miRNAs reported thus far have been found in herpes viruses, and several of the herpesviruses each encode over 20 predicted miRNAs (many of which have not yet been shown to be biologically functional). As DNA viruses that replicate in the nucleus, herpesviruses have access to the nuclear proteins Drosha and DGCR8 that are necessary for the processing of primary miRNAs (pri-miRNAs). Several herpesviruses appear to have auto-regulatory miRNAs, including those that maintain latency, while others use miRNAs to modulate cellular responses during infection (Table 2).
Notable herpesvirus-encoded miRNAs.

Alphaherpesvirinae subfamily members: herpes simplex virus-1 (HSV-1), herpes simplex virus-2, and varicella zoster virus
Members of the
The process of virion replication takes ~18–20 hours to complete and occurs in three stages corresponding to the expression of immediate early (IE; α), early (E; β), and late (L; γ) genes. During latency, there is little detectable transcription of IE, E, or L genes in infected neurons, even though it is known that they produce high levels of untranslated latency-associated transcripts (LATs). An unstable spliced 6.3 kb transcript and two stable LAT introns of 2 kb and 1.5 kb are spliced from an unstable 8.3 kb primary transcript.22–25 Although protein products of the LAT gene locus have not been discovered, the site has been associated with the repression of IE gene transcription and the maintenance of latency.26–28 More recently, miRNAs of both virus and host origin have been connected with maintaining HSV-1 or HSV-2 latency.
According to miRBase.org, 29 18 virus-derived pre-miRNAs have been identified from HSV-1 that encode 27 mature miRNAs. Many of these have been identified using bioinformatics or sequencing analysis and have yet to be characterized in functional biological assays. The best characterized miRNAs are HSV1-miR-H1 through miR-H6. miR-H1 and miR-H6 are located in the LAT promoter, while miR-H2, miR-H3, miR-H4, and miR-H5 are derived from the primary LAT transcript (Fig. 1). 30

Genomic location of selected HSV-1 pre-miRNAs.
The targets of most HSV-1 miRNAs are still unknown, but those that have been characterized point to a role in preventing viral reactivation from latency through regulation of
The earliest studies of HSV-1 miRNAs described their roles in maintaining latency. It is now known that many of these are also present and differentially expressed during productive infection.30,34,35 For example, miR-H1 and miR-H6 are more highly expressed than miR-H2, miR-H3, and miRH4 during productive infection than latency. 30 The opposite is true during latency, implying that miRNA expression plays a role in controlling the ordered expression of viral genes. Flores et al took the analysis of HSV-1 miRNA a step further by examining which of the miRNAs are loaded into the RISC as an indication of miRNA biological function. Surprisingly, only nine HSV-1 miRNAs were found to be associated with the RISC. In addition, the miRNAs were found to associate with the RISC at differing rates, 33 suggesting that several may not be functional. A recent study by Belter et al shows that miRNAs in high concentration are able to form secondary structures resembling RNA aptamers, 36 molecules whose specific 3D shapes allow them to bind targets with high affinity in the absence of the RISC.
A recent report indicates that host-derived miRNAs may also play a role in promoting HSV-1 latency. Pan et al found that the neuron-expressed miR-138 downregulated the expression of ICP0. A mutant HSV-1 virus lacking an ICP0 mRNA site complementary to miR-138 showed a twofold to fourfold decrease in ICP0 protein in neuronal cells. 37 Notably, the expression of ICP0 was unchanged in Vero cells infected with the mutant virus, further emphasizing that cell type is an important factor in determining the biological relevance of both viral and host miRNAs.
HSV-2 is the causative agent of genital herpes. Although viral miRNAs are not necessarily conserved between related viruses, HSV-2 shares ~85% homology with HSV-1 and has several miRNAs located at corresponding locations within its genome. HSV-2 miRNAs often share a high degree of homology with their HSV-1 counterparts when considering the seven-base pair seed regions of the miRNAs. 30 HSV-2 miRNAs have also been shown to exert similar regulation upon orthologous HSV-2 genes. For example, both HSV-2 miR-H2 (ie, miR-III) and HSV-1 miR-H2 are able to repress ICP0. 32 Similarly, HSV-2 miR-H3 and miR-H4 (ie, miR-I and miR-II, respectively), which are found in high copy number in neurons, are complementary to and lead to the downregulation of ICP34.5.32,38 There are exceptions to this conservation that demonstrate that the relative expression of each miRNA is not necessarily the same between the two viruses and that they may differentially regulate the expression patterns of viral genes. Specifically, miR-H2 is the most highly expressed miRNA during HSV-1 infection, while miR-H3 is most abundant during HSV-2 infection. 39 Additionally, not all miRNAs are identical between the two viruses: HSV-2 lacks an ortholog of miR-H1 and the miR-H6 found in HSV-2 uses the 5′ strand of the miRNA duplex, whereas HSV-1 miR-H6 uses the 3′ strand.30,31 Interestingly, the seed sequence of the HSV-2 miR-H6 5′ strand is identical to the seed sequence of the HSV-1 miR-H1, although the targets of either miRNA are currently unknown. 30
HSV-2 encodes a total of 18 virus-derived pre-miRNAs that derive 24 mature miRNAs. HSV-1 miR-H1, miR-H8, miR-H14, miR-H18, miR-H26, and miR-H27 have no known HSV-2 counterparts, while HSV-2 uniquely encodes miR-H9, miR-H10, and miR-H19 through miR-H25.
Varicella zoster virus (VZV; officially HHV-3) is the final member of the
Gammaherpesvirinae subfamily members: EBV and Kaposi's sarcoma-associated herpesvirus
EBV (HHV-4) and KSHV (HHV-8) are the two herpesviruses within the
EBV causes 90% of the cases of mononucleosis in teenagers or adults. The virus is also associated with Burkitt's lymphoma (BL), Hodgkin's lymphoma, primary effusion lymphoma (PEL), nasopharyngeal carcinoma (NPC), and gastric carcinomas (GaCas).
EBV was the first virus demonstrated to express miRNAs.
12
There are two clusters in the EBV genome that encode 25 pre-miRNAs that ultimately produce 44 mature miRNAs. The BamHI fragment H rightward open reading frame 1 (BHRF1) cluster is found within the BHRF1 locus and encodes three miRNAs: miR-BHRF1-1, miR-BHRF1-2, and miR-BHRF1-3. miR-BHRF1-1 is located in the promoter of BHRF1, whereas miR-BHRF1-2 and miR-HBRF1-3 are encoded in the 3′-UTR region of the gene. The second cluster of EBV miRNAs is located within introns of the BamH1-A region rightward transcript (BART) locus. This cluster encodes miR-BART1 through miR-BART22, with the exception of miR-BART2, which is found downstream of the BART locus between
EBV establishes and maintains persistent infection through a series of transcription programs characterized by regulated viral gene expression, 48 and the expression of certain miRNAs correlates with the latency program of infected cells. The first transcription program, known as Latency 3 or the growth transcription program, occurs when EBV infects a naïve B cell. This causes the cell to differentiate into a lymphoblast and proliferate. As with normally activated B lymphoblasts, the cell migrates to the lymph node germinal center follicle and continues to proliferate. It is at this point that the cell switches to the Latency 2 transcription program, also known as the default transcription program, which induces cell differentiation that causes the cell to leave as a resting memory B cell. In the periphery, the Latency 0 program initiates and protein translation ceases, except during the Latency 1 program, when the cell divides due to the expression of EBNA1. The process of productive viral replication and shedding into saliva is provoked by memory cells that return to the tonsil and undergo differentiation into antibody-producing plasma cells. 48
The BHRF1 miRNAs are highly expressed during the Latency 3 transcription program of infected BL and in lymphoblastoid cell lines (LCLs), which are generated through EBV infection of resting B cells. 49 Studies that created mutant viruses by introducing mutations into the BHRF1 miRNAs showed that these miRNAs are involved in contributing to B cell transformation by promoting cell-cycle progression and inhibiting apoptosis.50,51 In stark contrast, BHRF1 miRNAs were not detectable in NPC, PEL, or BL cell lines in Latency 1 or 2 transcription programs, whereas BART miRNAs were expressed to high levels. 49 In addition, NPC and GaCa epithelial tumors exhibited 13- and 8-fold higher expression, respectively, of BART miRNAs compared to LCLs. 52 Taken together, this suggests that BART miRNAs appear to be preferentially expressed in epithelial cells (such as NPCs), while they are moderately expressed in B cells and dispensable for in vitro EBV-induced transformation. 53
Several viral and host mRNA targets of EBV miRNAs have been identified. miR-BART2 is complementary to
Also, EBV miRNAs can target host mRNAs to prevent apoptosis. Gene ontology analysis of the host mRNA targets of the 12 most abundant EBV miRNAs indicated that 132 apoptosis-associated host genes may be targeted by EBV miRNAs.
55
Notably, the p53 upregulated mediator of apoptosis (PUMA), a pro-apoptotic gene induced by p53, has been shown to be targeted by miR-BART5 and miR-BART19.55,59 Also related to promoting apoptosis, the BCL2 family member
EBV miRNAs have also been implicated in evasion of NK cells through miR-BART2-mediated downregulation of MHC Class I Polypeptide-Related Sequence B (MICB), a stress-induced NK cell ligand, in epithelial cell lines.55,63 In addition, the T cell-attracting chemokine CXCL11, produced by B cells during EBV infection, is downregulated by EBV miR-BHRF1-3.12,64 EBV miR-BART18 decreases the histone acetylase cyclic AMP-responsive element-binding protein (CBP),
65
which along with p300 associates with IRF3 and IRF7 to induce the transcription of Type 1 interferon (IFN) genes. Blocking miR-BART18 in Akata A.15 cells, an EBV+ BL cell line, led to increased Type 1 IFN signaling, as measured in terms of
The other human gammaherpesvirus is Kaposi's sarcoma-associated herpesvirus (KSHV) (HHV-8). The greatest risk of infection with KSHV is for immunocompromised individuals, who more frequently develop Kaposi's sarcoma. Like EBV, KSHV infection is also associated with PEL and multicentric Castleman disease. 66
According to miRBase.org, 29 13 pre-miRNAs and 25 miRNAs are found within the KSHV genome. The miRNAs are all located as a cluster within the KSHV latency-associated region (KLAR), which encodes four genes expressed during latency and lytic infection: latency-associated nuclear antigen (LANA), v-Cyclin, v-FLIP, and Kaposin (K12). All the miRNAs map to the K12 locus: miR-K12-1 through miR-K12-9 and miR-K12-11 are encoded within the K12 intron, while miR-K12-10 and miR-K12-12 map to a K12 open reading frame and the 3′-UTR, respectively (Fig. 2).18,67,68 Although the relative expression levels can vary in different cell types, the 10 miRNAs encoded on the K12 intron are generally expressed as a group in latently infected cells.9,69,70 In contrast, miR-K12-10 and miR-K12-12 are expressed alongside the K12 gene during lytic infection. 71

Genomic location of KSHV pre-miRNAs. KSHV pre-miRNAs cluster within the KLAR, which also contains genes for LANA, v-Cyclin, v-FLIP, and Kaposin. All the miRNAs map to the K12 locus: miR-K12-1 through miR-K12-9 and miR-K12-11 are encoded within the K12 intron, while miR-K12-10 and miR-K12-12 map to a K12 open reading frame and the 3′-UTR, respectively.
The cluster of KSHV miRNAs is expressed alongside the KLAR genes during latency, and similar to HSV-1 and HSV-2 miRNAs, it has been shown to play a role in preventing reactivation to the lytic cycle. The expression of the KSHV replication and transcription activator (RTA), the master regulator of the latent-lytic switch, is negatively regulated by miR-K12-9 and miR-K12-7.72,73 Similarly, miR-K12-3 and miR-K12-11 target cellular activators of RTA.
74
Inhibiting or eliminating these KSHV miRNAs (or a cluster including these miRNAs) induced elevated lytic gene expression and spontaneous lytic reactivation in fibroblasts, endothelial cells, and PEL cells.72–75 This further emphasizes that several KSHV miRNAs are involved in the maintenance of latency in both lymphoid and non-lymphoid cells. It is also interesting to note that host-encoded miR-498 and miR-320d have also been shown to target RTA in a PEL cell line,
76
and 5 of the 99 cellular miRNAs induced by ectopic expression of HIV
Recent work by McClure et al and Bai et al mapped the 3′-UTRs of KSHV genes and discovered that these regions are important in the negative regulation of KSHV genes.78,79 A total of 28 potential KSHV gene targets of known KSHV miRNAs were identified that correspond to all stages of viral replication, 79 indicating that many miRNA targets have yet to be investigated.
In addition to viral transcripts, KSHV miRNAs target many host mRNAs. Notably, KSHV miRNAs are thought to target >1000 putative host genes,9,70 although not necessarily directly.
80
Several classes of cellular genes consistently targeted include apoptosis, angiogenesis, cellular metabolism, lymphocyte activation, and immune modulation genes.9,80,81 For example, several KSHV miRNAs target cellular thrombospondin 1, which is thought to negatively regulate angiogenesis and proliferation.
81
KSHV miR-K12-K1 binds the 3′-UTR of
As a herpesvirus, KSHV encodes several viral orthologs of host genes, including
In addition to encoding homologs of cellular miRNAs and its own miRNAs that target host genes, KSHV also expresses proteins that induce several infection-promoting cellular miRNAs. For example, knocking down cellular miR-21 and miR-31 prevented KSHV K15M-mediated motility of PEL cells, 89 and cellular miR-132 regulates interferon-stimulated genes in KSHV-infected lymphatic endothelial cells. 90 It is not surprising that several herpesviruses use miRNAs to interfere with Type 1 IFN signaling; miR-K12-12 also appears to target CBP, which is involved in the activation of Type 1 IFN genes. 65
Betaherpesvirinae subfamily members: human cytomegalovirus, HHV-6A, HHV-6B, and HHV-7
Human cytomegalovirus (HCMV; officially HHV-5), HHV-6A, HHV-6B, and HHV7 establish latency in leukocytes and are characterized by slower replication than the other herpes viruses. Healthy individuals generally do not display symptoms when infected with HCMV, although 10–20% of infectious mononucleosis cases are attributed to the virus. The virus can reactivate and cause life-threatening disease in immunocompromised individuals, and HCMV is the most prevalent congenital infection in industrialized countries.
According to miRBase.org, 29 15 pre-miRNAs encode 26 mature HCMV miRNAs. Unlike KSHV or EBV, the HCMV miRNAs are scattered throughout the viral genome. They can be found on both strands, present in 3′-UTRs or within intergenic regions. 91
HCMV miRNAs target both viral and host genes. Most HCMV miRNAs correlate with the expression of lytic genes.92,93 miR-UL112-1 targets several HCMV genes, including IE1, IE72, UL112/113, UL114, and UL120/121.94–96 The expression of UL138, a viral protein that contributes to latency, 97 is decreased 46% in miR-UL36-transfected HEK293 cells, 98 suggesting that this miRNA may be involved in maintaining productive infection. However, no HCMV miRNA has been found to be essential for replication.
Although the majority of its miRNAs have been characterized during lytic infection, HCMV may also encode a subset of miRNAs that self-regulate to maintain a latent state. As mentioned above, miR-112-1 has been shown to downregulate IE72, an abundant HCMV transactivator necessary for induction of E and L gene expression. 95 The HCMV IE1 and IE2 transactivators are central to reactivation from latency, 99 and HCMV miR-112-1 binds to the 3′-UTR of IE1 and reduces reporter expression in transient-transfection assays of HEK293T cells. 96 In support of this, miR-US33 transfection of human embryo lung fibroblast cells reduced infectious viral titers and IE1/IE2 proteins at a multiplicity of infection of 0.01 (although not 0.1, 1, or 5). 93 Thus, although few HCMV miRNAs are found to be expressed during latency, several putative miRNAs may have a significant effect upon the suppression of required IE genes.
In addition to targeting viral genes, HCMV also encodes miRNAs that target cellular genes. For example, miR-US25-1 and miR-US25-2 target several cellular targets, many of which are associated with cell-cycle control, and infection with an miR-US25-1 knockout virus resulted in increased cyclin E2 expression in human primary fibroblast cells. 100 Like other herpesvirus miRNAs, HCMV also encodes miRNAs that target cellular genes in an effort to evade host immune responses. Notably, HCMV miR-UL112 was the first miRNA shown to inhibit expression of MICB. It also acts in concert with the HCMV-encoded UL16 protein to inhibit expression of MICB when UL16 protein levels are reduced. 101 CCL5, a chemokine that attracts T cells, is downregulated in human foreskin fibroblasts by HCMV miR-UL148D, 102 while MHC Class I peptide loading is inhibited by the miR-US4-1-mediated downregulation of endoplasmic reticulum (ER) aminopeptidase 1 (ERAP1). 103 Taken together, HCMV encodes several miRNAs that function to regulate host responses.
Whereas HHV-6A is an orphan virus, HHV-6B and HHV-7 cause roseola infantum and are associated with febrile seizures of children below two years of age. Deep sequencing has identified four miRNAs, termed hhv6b-miR-Ro6-1 through miR-Ro6-4, encoded within the left and right direct repeat regions of HHV-6B. 104 Hhv6b-miR-Ro6-3, miRRo6-2, and miR-Ro6-1 are antisense to the predicted B1, B2, and B3 open reading frames (ORFs), respectively, that are thought to encode IE genes. 105 Future studies will determine if these HHV-6 miRNAs are involved in regulating early lytic infection or maintaining latency. Thus far, no viral miRNAs have been identified for HHV-7.
Biomarker and therapeutic potential with respect to herpesvirus miRNAs
Herpesviruses encode proteins that subvert host immune responses, and it is clear that the viruses also encode several miRNAs that interfere with innate and adaptive antiviral mechanisms. Type 1 IFN signaling activates a variety of cellular antiviral pathways and is a potent inducer of NK cell activation and cytotoxicity. 106 Not surprisingly, HCMV, EBV, and KSHV encode miRNAs that target Type 1 IFN signaling, 65 and these viruses also encode miRNAs that downregulate MICB.55,63,101 Blocking miR-BART18 in an EBV+ BL cell line resulted in increased Type 1 IFN signaling, 65 highlighting that efforts to inhibit herpesvirus miRNAs could be fruitful in reestablishing IFN-induced antiviral pathways. Likewise, inhibition of viral miRNAs may be useful to reverse IFN refractoriness of EBV+ or KSHV+ tumor cells 65 or increase the efficacy of interferon therapy for herpesvirus conditions, such as HSV epithelial keratitis. 107 Conversely, these herpesvirus miRNAs could assist in the regulation of aberrant Type 1 IFN responses in systemic lupus erythematosus and other diseases. 108
Adaptive immune responses are also circumvented through herpesvirus miRNAs, either directly or indirectly. Inhibition of Type 1 IFN signaling through miRNAs mentioned above can negatively regulate antigen-presenting cell (APC) and CD8 T cell responses.109,110 In addition, T cell-attracting chemokines CXCL11 and CCL5 are targeted by EBV and HCMV miRNAs, respectively,12,64,102 and HCMV miR-US4-1 downregulates ERAP1 to inhibit MHC Class I peptide loading in the ER. 103 Correspondingly, inhibition of herpesvirus miRNAs during infection would be expected to modulate APC and T cell responses.
Herpesvirus miRNAs have huge potential as biomarkers of diagnosis and disease progression. As an example, HCMV is the most common infection of patients after organ transplantation and can drastically affect morbidity, mortality, and organ rejection. Diagnostic results assist in monitoring of infection following transplantation and guide decisions to give preemptive therapy. 111 The current standards to test for acute infection include serology for IgM/IgG titers, an antigenemia assay to test for HCMV pp65 antigen, and quantitative nucleic acid testing (QNAT) using PCR. However, false-positive serology results can occur in patients with EBV or HHV-6 infections, and IgM antibodies take weeks to appear. This underscores the limitations of traditional serology to quickly assess new infection. Moreover, the pp65 antigenemia assay cannot be used in patients with neutropenia, and QNAT is extremely sensitive and faster than culture but does not differentiate between shedding of virus in the absence of active disease. 111 Certain HMCV miRNAs are expressed at different stages of viral replication and could therefore be used to further monitor primary infection or reactivation from latency. Samples could be easily obtained from biopsies, whole blood or plasma, known areas of shedding such as saliva, or other fluids such as cerebrospinal fluid. In support of using herpesvirus miRNAs as biomarkers, Kawano et al found that certain EBV miRNAs were elevated in plasma of patients with chronic active EBV infection compared to control patients or patients with infectious mononucleosis. 112 A recent study by Zhang et al found that EBV miR-BART7 and miR-BART13 levels in plasma specimens produce a 90% predictive value for NPC. In addition, both miRNAs were downregulated following radiotherapy. 113
Exosomes, which are lipid microvesicles released from cells, are thought to provide extra stability to their encapsulated miRNAs. Exosome miRNA can be separated from free miRNAs, virion-associated miRNAs, or genomic nucleic acid. 114 miRNAs have been detected within exosomes in plasma from human patients with KSHV and from EBV+ LCL and NPC cell lines.69,114–116 Exosome-associated miRNAs in plasma could provide more stable samples for analysis, leading to better diagnostics to monitor active infection or herpesvirus-associated malignancies.
Although some herpesviruses exhibit conservation in their miRNA seed regions, diagnostic miRNA microarrays could provide a means to differentiate between herpesvirus infections that cause similar conditions, such as roseola infantum caused by HHV-6B or HHV-7. The plausibility of this idea is supported by a study by Marshall et al that showed that the majority of KSHV-encoded miRNAs were highly conserved within clinical isolates. 69 Because reactivation from latency is a critical step in the replication of herpesviruses, comparison of lytic versus latent miRNAs may also be useful in diagnosis, monitoring, and treatment. Although excellent work has been done to elucidate the biological functions of the miRNAs present during herpesvirus infection, additional serum profiling studies from infected individuals need to be performed in order to have a more thorough understanding of which viral and cellular miRNAs are present during the different stages of herpesvirus infection.
Polyomaviruses
Another family of double-stranded DNA viruses that replicate in the nucleus is the
Infections of humans with polyomaviruses can result in serious diseases, although it is usually quite rare. For example, infection with Merkel cell polyomavirus (MCPyV) can lead to Merkel cell carcinoma 117 and BK polyomavirus (BKPyV) can cause polyomavirus allograft nephritis, polyomavirus hemorrhagic cystitis, and bladder cancer in some cases.118–125 Additionally, JC polyomavirus (JCPyV) can cause progressive multifocal leukoencephalopathy (PML) and has been associated with cases of colorectal cancer,126,127 while trichodysplasia spinulosa-associated polyomavirus can cause trichodysplasia. 128
As nuclear DNA viruses, polyomaviruses are good candidates to encode miRNAs. Unlike the herpesviruses that each encodes a handful of miRNAs, most or all polyomaviruses encode a single pre-miRNA within the large T antigen (LTAg) gene (Table 3). Although the exact location of the pre-miRNA varies, it is always oriented in the opposite direction of the LTAg and is transcribed as part of late transcription events. Two mature miRNAs are produced from the pre-miRNA that downregulate the expression of the LTAg, which is necessary for the early/middle phase of the infection cycle. Therefore, the main role of the polyomavirus miRNAs appears to be in helping the transition from the early to late phase of the infection cycle.
Summary of human polyomavirus-encoded miRNAs.
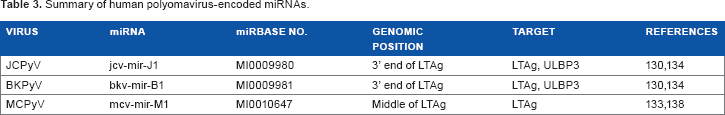
Cellular targets of polyomavirus miRNAs have also been identified and appear to target immune responses. Both JCPyV and BKPyV miRNAs bind to the 3′-UTR and reduce translation of UL16-binding protein 3 (
The 5′ and 3′ miRNAs encoded by polyomaviruses appear to have potential as biomarkers of infection and are detectable in plasma, urine, brain tissue, colon tissue, and feces.134–136 JCPyV miRNA was isolated from both plasma and urine in healthy asymptomatic individuals using qRTPCR, which was more sensitive in detecting the presence of the virus than traditional serological methods. 136 JCPyV miRNA has been found in postmortem brain tissues of PML patients, 134 and miR-J1-5p was also detectable in colon tissue and feces of healthy subjects. 135 Interestingly, miRNA expression was lower in diseased versus normal adjacent tissue and inversely correlated with expression of LTAg, possibly supporting its role in downregulating expression of the protein. 135 Currently, serology for JCV proteins is most commonly used to indicate infection, but together with the apparent stability of the miRNA, 134 the ease in obtaining polyomavirus miRNAs from patient samples indicates that they may be powerful biomarkers for detecting asymptomatic forms of polyomavirus infections. On the other hand, using host-encoded miRNAs that are induced by PyV infection as potential biomarkers may not be as fruitful: in a miRNA profiling study, Lagatie et al found that there was no significant difference in host-encoded miRNAs between individuals with or without JCPyV. 137
Using polyomavirus miRNAs or their respective antagomirs (complementary oligonucleotides used to silence miRNAs) as therapeutics will most likely be difficult. A major problem lies in the fact that most individuals are infected asymptomatically by PyVs and never develop serious health consequences. Later, when the virus is assumed to be in a latent form of infection, the miRNA is not being expressed and thus there is no target to antagonize. In cases where tumorigenesis has occurred and a proviral form exists, the virus is likely to be expressing only E genes, 138 which does not include the miRNA. Since tumorigenic activity is believed to largely come from the action of LTAg, 139 it is interesting to speculate that the PyV miRNA could serve as a therapeutic in malignant cells to theoretically reduce the amount of LTAg ex pressed. However, this is also likely to induce the lytic cycle, which may have its own unintended consequences.
Hepatitis B virus
The hepatitis B virus (HBV) is a small, enveloped DNA virus belonging to the
As a DNA virus that enters the nucleus, HBV is a good candidate to encode miR NAs. Nevertheless, no HBV miRNAs have been verified experimentally, and only one has been identified through computational methods. 143 On the other hand, there are a large number of host-encoded miRNAs described that interact with HBV and are differentially expressed during the course of infection (Table 4). Additionally, miRNAs have been identified that are specifically involved in the progression of HBV-related diseases. 144 Thus, cellular miRNAs, rather than viral miRNAs, may have the potential to become powerful biomarkers of HBV pathology.
Host miRNAs involved in HBV-related health conditions.

The most studied miRNA involved in the course of HBV infection is perhaps miR-122. It represents 50–70% of the miRNAs found in normal liver cells145–147 and plays a role in maintaining homeostasis, as well as regulating metabolic pathways involving lipid and cholesterol metabolism.148,149 miR-122 has been shown to downregulate cyclin G1 and heme oxygenase 1 (HMOX1), resulting in inhibition of HBV replication.150–152 Additionally, Chen et al found that miR-122 has viral mRNA target sequences located in the HBV coding region of the reverse transcriptase and the 3′-UTR of the HBcAg core protein.153 The virus counters with production of X protein, which binds peroxisome proliferator-activated receptor gamma to inhibit the transcription of miR-122. 154 In addition, all four HBV mRNAs have complementary sites for miR-122 that act as sponges to sequester endogenous copies of the miRNA. 155
Not surprisingly, miR-122 is downregulated in a number of HBV-associated hepatic cancers. 156 Fan et al showed that miR122 negatively regulates tumor-promoting N-myc downstream-regulated gene 3, which is upregulated in HCC+ patient liver samples compared to normal adjacent tissue. 157 The biological importance of miR-122 was further emphasized when the malignant phenotype of an HBV-related HCC cell line could be reversed by transient transfection of miR-122 into the cells. 157 Inhibition of miR-122 also leads to the increased expression of pituitary tumor transforming gene binding factor, resulting in liver cancer cell proliferation, invasion, and tumor growth. 158
Other cellular miRNAs are targeted in a similar manner by HBV. Two well-known tumor suppressor miRNAs, miR-15a and miR-16-1, are found in multiple human cancers and are decreased in HBV-infected cells by HBV X protein.159–161 As occurs with miR-122, HBV mRNAs have a site complementary to miR-15a and miR-16-1 and act as sponges to sequester the miRNAs. As a result, the oncogene Bcl-2, a regulatory target of miR-15a/16-1, was increased significantly in HBV-transfected cells. 162 Viral targeting of these miRNAs, specifically miR-15a, likely evolved in response to the ability of the host miRNA to target HBp and HBx transcripts to suppress HBV infection. 159
Several other cellular miRNAs are induced by infection, although not all result in a benefit to the host. HBsAg expression and HBV proliferation are directly suppressed by miR-199a-3p and miR-210, 163 and miR-125a-5p downregulates the expression of HBsAg. 164 On the other hand, Zhang et al showed that miR-1 indirectly enhances HBV replication by targeting histone deacetylase 4 and E2F transcription factor 5, the activation of which arrests the cell cycle and reverses the cancer phenotype. 165 In support of this, it has been found that miR-1 is epigenetically regulated and found in lower levels in HCC cells. 166 Paradoxically, the HBV HBx protein promotes the expression of cellular miR-148a, which enhances tumorigenesis and promotes cell proliferation, cell migration, and anchorage-independent growth of HepG2 and Hep3B cells.167,168 Further research is needed to discern the relationship between tumorigenic and antitumorigenic miRNAs during HBV infection.
Traditional detection methods and notable miRNA biomarkers of HBV infection
Alanine aminotransferase (ALT) and aspartate aminotransferase (AST) are the most commonly used biomarkers to assess liver damage, 169 although levels can be altered through conditions unrelated to the liver.170,171 A more definitive diagnosis can be obtained by liver biopsy, but biopsy is costly, inconvenient, and may not be accessible to all individuals. 172 Furthermore, complications such as pain, mental and physical anguish, bleeding, and even death are possible.172–174 Another problem with liver biopsy lies in the sampling nature of the biopsy itself. Typically, liver samples taken at biopsy represent 1/50,000th of the total liver, which could result in random sampling error. 172 While ultrasound is a tempting noninvasive alternative and good for assessing late stage liver damage, it is not very useful in assessing earlier stages of disease. 175
For specifically identifying HBV as a causative agent of liver damage, current serological methods consist of antibody tests for HBsAg and HBeAg antigens. PCR assays may also be used for direct determination of HBV genomic DNA in serum. For identification of HBV in tissues, detection of HBsAg and HBcAg by immunohistochemical staining or HBV DNA by Southern hybridization, in situ hybridization, or PCR is performed. 176
Considering the above challenges, miRNAs have emerged as an alternative biomarker method with the possibility of monitoring the progression of disease in individuals. Multiple studies have already shown that noninvasive testing for miRNAs has the potential to be used as biomarkers of HBV infection and HBV-positive HCC.177,178 A number of specific, differentially expressed host miRNAs have been identified in the last decade; a partial list is shown in Table 5.
Host miRNAs reported to be differentially expressed in serum/plasma during HBV infection.

One of the most widely used miRNA biomarkers for HBV is miR-122, not surprising due to it being one of the most thoroughly studied miRNAs in HBV infection and liver disease. Waidmann et al found that the serum levels of miR-122 were effective as a biomarker in HBV-infected patients as they discriminated infected from healthy subjects, discriminated inactive carrier patients with high or low levels of HBsAg, and correlated with the levels of ALT, HBV genomic DNA, and HBsAg. 179 It has also been shown that both miR-122 and miR-18a are released in the blood and could be used for HBV-related HCC screening.160,179,180 An miRNA profiling study on HBV and hepatitis C virus (HCV) found that miR-122 was significantly upregulated in serum of patients with both viruses, whereas elevated miR-22, miR-99, and miR-125b levels were more characteristic of chronic HBV infection and may be useful in discriminating between infection with the two viruses. 181
Other panels of miRNAs have also been found to be differentially expressed between healthy and chronic HBV patients. Zhang et al identified 34 miRNAs dysregulated in chronic hepatitis B patients, with miR-122, miR-572, miR-575, and miR-638 upregulated and miR-744 downregulated significantly. 182 A study examining miRNAs in the plasma of chronically infected children identified 16 miRNAs to be upregulated in HBeAg+ compared to children with HBeAg–: miR-99a, miR-100, miR-122, miR-122*, miR-125b, miR-192, miR-192*, miR-193b, miR-194, miR-215, miR-365, miR-455-5p, miR-455-3p, miR-483-3p, miR-885-5p, and miR-1247. 183 Another miRNA serum profiling study also identified miR-99a-5p, miR-122-5p, and miR-192-5p as significantly overexpressed between chronic hepatitis B adult patients and inactive carriers. Using an MiR-B-Index that normalized these three miRNAs to internal control miRNAs (miR-126, miR-320a, and miR-335), the same study showed that the serum miRNA profile of patients responding to pegylated (PEG)-IFN alpha resembled that of inactive carriers, while nonresponders and relapsers matched baseline measurements of patients with chronic hepatitis B. 184 This comprehensive study illustrated that miRNAs could be useful not only in assessing disease status (inactive carriers vs. chronic hepatitis B patients) but also in identifying responders to certain treatments, such as PEG-IFN. The study also emphasized that although a single biomarker is desirable for simplicity and financial reasons, panels of biomarkers can also be effective in determining disease state and response to treatment. Further support for biomarker panels is provided by several studies.177,185,186 Notably, Li et al used a panel of 13 miRNAs to distinguish control patients from those with HBV infection (miR-10a, miR-223, miR-375, and miR-423), control patients from those with HBV-HCC (miR-23a, miR-23b, miR-92a, miR-342-3p, miR-375, and miR-423), patients with HBV from those with HCV infection (miR-92a up in HCV and miR-375 up in HBV), and patients with chronic HBV from those with HBV-positive HCC (miR-19a and miR-125b). 177
Liver injury can produce similar biomarker responses, regardless of the causative agent. An interesting miRNA profiling study by Ura et al examined the expression of 188 miRNAs from HBV-HCC, HCV-HCC, and normal patients. They identified 19 miRNAs that were differentially expressed between HBV and HCV. 187 In all, 31 miRNAs were associated with liver disease, regardless of the virus, but 6 were specific for HBV and 13 for HCV. Notably, miR-105, miR-134, and miR-211 were over fourfold upregulated in HBV and miR34c was over fourfold increased in HCV. 187 However, other studies have produced conflicting results,188,189 and thus, the miRNAs identified are likely to need further validation.
Role of miRNAs as therapeutics against HBV
The potential that miRNAs hold to serve as therapeutic agents against HBV has long been recognized,190–193 and a number of studies have attempted to employ miRNA to combat HBV
Multimeric miRNA expression cassettes have been used to effectively target multiple sites within the virus genome using mouse models of HBV infection,
214
and a common delivery method for the miRNA therapeutic uses adeno-associated virus (AAV) or lentivirus systems.215,216 AAV-delivered miR-26a resulted in cell-cycle arrest
Hepatitis C virus
The HCV is a small, enveloped, positive-strand RNA virus belonging to the
As a cytoplasmic RNA virus, HCV is unable to encode miRNAs using traditional nuclear processing machinery, and no HCV-encoded miRNAs have yet been described. Additionally, encoding a miRNA within its genome would be a risky strategy for an RNA virus, as the miRNA processing cell machinery might target the genome for cleavage. HCV does encode suppressor of RNAi silencing proteins via the HCV core protein and envelope protein E2,219,220 and a plethora of host-encoded miRNAs have been described that interact with HCV and are differentially expressed during the course of an infection. In fact, as the disease progresses from initial infection all the way to HCC, corresponding changes in miRNA levels have been detected in profiling studies.187–189,221–223 If a reproducible pattern of specific host miRNAs could be documented as disease progression occurs, it could identify miRNAs as powerful biomarkers of HCV pathology. Table 6 provides a partial list of miRNAs shown to be differentially expressed or found to interact with HCV during infection.
miRNAs involved in HCV infection.
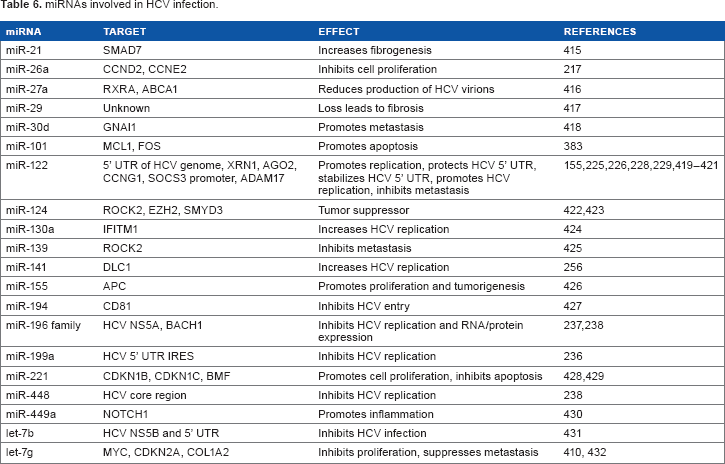
Like HBV, most of the attention regarding miRNAs and HCV has centered around miR-122, which, as mentioned above, represents 50–70% of the miRNAs found in normal liver cells.145–147 It is important to note that while miR-122 is an inhibitor of infection by HBV,150,151,153,157 it
Traditional detection methods and putative biomarkers of HCV infection
The initial testing for HCV is usually done by enzyme immunoassay to detect host antibodies directed against HCV 230 Liver damage associated with HCV is also screened for using ALT and AST enzyme tests.230,231 Positive results in initial screenings are usually confirmed via molecular testing to detect the presence of HCV RNA, sometimes followed by patient biopsy to assess the extent of liver damage, if necessary. 230 As described above for HBV, liver biopsies are not without serious drawbacks.173,174
Algorithms are also used that include a variety of additional criteria and biomarkers such as age, sex, levels of other liver enzymes, or platelet numbers. Examples include Firbo-Test and HepaScore.232,233 Unfortunately, many results fall in the
Compared to standard diagnostic methods, the potential advantages of using miRNAs as HCV biomarkers include the noninvasive nature of retrieval, generation of fairly rapid results, and possible lower costs compared to other methods. Profiling studies are often used as the first step in identifying candidate miRNAs that could serve as biomarkers, and there are already several studies that compare HCV-related disease states and healthy individuals.187–189,221–223,235 These studies have begun to reveal the potential of miRNAs to be a useful tool for diagnosing HCV-related disease as well as monitoring the progression or regression of the disease.
Several miRNAs are differentially regulated as a result of HCV or HCV-induced conditions (Tables 7 and 8). Notably, studies have demonstrated the utility of using miR-122 to identify HCV, HCC, and hepatocyte injury,181,234 and this miRNA is also extensively used to identify HBV status.160,177,179 Other host miRNAs directly target HCV genomic sequences: miR-199a reduces HCV replication by targeting the 5′-UTR of the genome internal ribosome entry site (IRES) region. 236 Additionally, the expression of miRNA-196b appears to be upregulated by the IFN pathway and can thus inhibit HCV by directly targeting NS5A or by indirectly targeting BACH1 and HMOX1.237–239 However, there have been conflicting results reporting whether it is upregulated or downregulated during infection,239–241 possibly making its implementation as a biomarker difficult. As a way of effectively distinguishing HCV versus HBV infection, miR-92a and miR-375 are more highly upregulated in HCV+ or HBV+ patients, respectively. miR-10a and miR-125b are not upregulated in HCV+ but highly upregulated in HBV+ individuals, 177 although miR-10a is upregulated in HCV-HCC cases. 223 Several biomarkers overlap in these and other non-viral liver conditions, emphasizing the importance of biomarker panels to properly differentiate these diseases.
Host miRNAs upregulated during HCV infection.

Host miRNAs differentially expressed in liver tissue biopsies during HCV infection, HCC, or HCV-associated HCC.

miRNAs as therapeutics for HCV
Current therapeutics for HCV include the use of PEG-IFN and ribavirin,242–244 and more recently, sofosbuvir, ledipasvir, and boceprevir have been used effectively.245–248 While many therapies are successful, there are still a significant number of patients in whom the virus is not fully cleared.249,250 Additionally, therapies are costly, not easily accessible to all infected individuals, and can have severe side effects.251,252
Recently, miR-122 has emerged as a promising candidate in the implementation of miRNAs as therapeutics. The observation that this abundant liver miRNA is upregulated and promotes replication during HCV infection made it an obvious choice to antagonize therapeutically. A small effector molecule against miR-122 inhibited HCV replication in liver cells, 253 and Lanford et al used an antisense locked nucleic acid (LNA; an oligonucleotide with greater strength of binding due to chemical modification) derivative of miR-122 as a therapeutic in HCV-infected chimpanzees. A sharp reduction in HCV RNA and improved liver histology was observed without significant side effects or detectable resistance from the virus. 254 Phase 1 clinical trials of miravirsen, an LNA that targets miR-122, were completed in 2009 by sponsor Santaris Pharma. In Phase 2a studies of null responders to PEG-IFN with HCV genotype 1 infection, miravirsen reduced mean HCV RNA levels up to three orders of magnitude with no drug-associated severe or serious adverse events. 255 Notably, HCV RNA levels increased following cessation of the drug. Long-term administration studies are ongoing (ClinicalTrials.gov identifiers NCT01727934 and NCT02031133). These results will possibly usher in a new era of miRNA-based biologics.
While the results using miR-122 are promising, there are still several other miRNAs worth considering to this end. Pedersen et al described mimics of five miRNAs (miR-196, miR-296, miR-351, miR-431, and miR-448) that reduced HCV replication and infection when transfected into HCV+ cells. 238 Antagomir-mediated knockdown of miR-141 effectively inhibited HCV replication in infected hepatocytes. 256 Moreover, the mimics of three miRNAs, miR-196b, miR-199a-3p, and miR-29, have been expected to function in anti-HCV therapy. 257
An interesting related approach was considered by Yang et al who created artificial miRNAs based on five preselected targets on the HCV genome. When introduced into cells, they dramatically reduced the replication of cell culture-propagated HCV without causing hepatocellular toxicity. 258 This may suggest that knowledge of the viral target sequence may be just as valuable, if not more so, than targeting naturally occurring miRNAs. Since artificial miRNAs and their derivatives are relatively easy to create and introduce into a host, this could help alleviate the need for extensive profiling studies to identify aberrantly expressed miRNAs as targets.
Human papillomavirus
HPV is a small, non-enveloped, double-stranded DNA virus that infects epithelial cells. Over 120 different types have been identified. While most infect cutaneous epithelium, ~40 types infect mucosal epithelium, some of which are associated with oncogenesis. Types 16 and 18 together account for ~70% of cervical cancer cases. HPV is also associated with less common cancers, including cancer of the anus, penis, vulva, and vagina. In the United States, HPV is the most common sexually transmitted disease. The virus is also associated with the vast majority of head and neck cancers in nonsmokers. 259
The majority of HPV infections are eventually cleared from the host, but a small proportion result in persistent infections that can progress to cervical intraepithelial neoplasias (CINs). High-grade CINs (CIN2 or CIN3) are considered a precursor to cervical cancer, which may occur within years or decades. 259 As such, much effort has been invested to characterize the role of the virus in oncogenesis, particularly pertaining to the roles of viral oncoproteins E5, E6, and E7.
Recently, host miRNAs have also been implicated in the process. Several miRNAs have been identified as specifically targeted by E5, E6, or E7 (Table 9), and their subsequent cellular effects in many cases have been characterized. E6 targets p53,260,261 a transcription factor that is thought to be used for transcription of many cellular miRNAs.262,263 As a result, many miRNAs may be drastically underexpressed and might lead to adverse cellular effects. miR-23b, miR-34a, miR-203, and miR-218 are all found to be downregulated in E6-expressing cells,264–269 which is particularly noteworthy because of the association of these miRNAs with oncogenic processes, in part through direct or indirect effects of p53 inhibition.
HPV proteins known to alter host miRNAs.
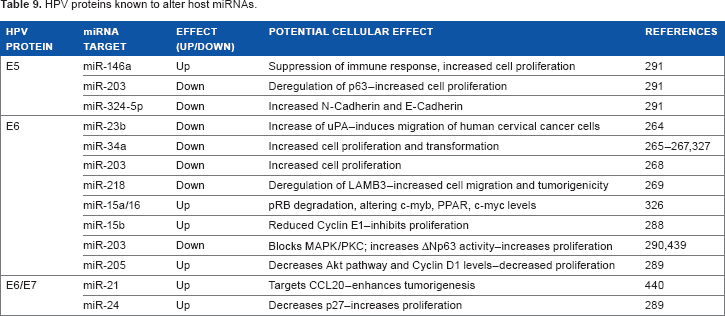
Although all the above miRNAs have been implicated in cervical cancer, they have also been reported as tumor suppressors in a variety of other cancers: miR-23b in prostate, bladder, and breast cancers270–272; miR-34a in neuroblastomas, HCC, and ovarian, colon, pancreatic, and bladder cancers266,273–276; miR-203 in lung, prostate, and esophageal cancer, as well as leukemia and glioma277–279; and miR-218 in prostate, colon, gastric, and bladder cancer.280–282
E7 degrades the tumor suppressor protein pRB, which releases E2F from the pRB–E2F complex, 283 freeing E2F to activate transcription. The promoter regions of several miRNAs contain E2F binding sites,280,284,285 which could possibly lead to adverse effects on the cell if oncogenic miRNAs are transactivated. In contrast to E6-induced miRNAs, most of the miRNAs shown to be dysregulated through the action of E7 are upregulated, such as miR-15b/16-1, miR-24, and miR-205.286–289 The downregulation of miR-203 has been credited to the indirect blocking of the MAPK/PKC pathway by E7. 290
Less is known about the actions of E5 on cellular miRNAs. A recent study found that several miRNAs were altered in HPV16 E5-expressing HaCaT cells. 291 Notably, the authors of that study found that miR-146a was induced, while miR-203 and miR-324-5p were repressed. miR-146a and miR-203 had previously been shown to be dysregulated in the same manner in cervical cancer tissues or HPV+ cell lines.290,292–300
miRNAs as potential biomarkers of HPV infection and progression
It is suspected that the expression of many miRNAs may be altered during the course of HPV infection, possibly contributing to oncogenesis. Several studies have examined miRNAs dysregulated in cervical cancer or head and neck squamous cell carcinoma (HNSCC), two cancers that are likely to be the result of HPV infection. This section of our review focuses on those fewer studies that specifically addressed HPV+ cell lines or HPV+ cancer cells/tissues for the purposes of identifying HPV-specific biomarkers (Table 10), pointing out when individual miRNAs have also been identified in cancerous cells or tissues with a possible contribution by HPV.
Selected miRNAs identified in HPV-associated conditions.

miR-21 has been implicated as an oncogenic miRNA for many cancers. 301 Indeed, almost all profiling studies to date of cervical and head and neck cancers have found miR-21 to be overexpressed in the cancerous state,292,293,296,300,302–314 and miR-21 has been shown to be upregulated in HPV-related cancers as well.308,315 Lajer et al compared HPV+ and HPV–patients with tonsillar squamous cell carcinoma (TSCC) and found miR-21 to be upregulated in HPV+ patients. Additionally, miR-21 was upregulated in HPV+ or HPV– HNSCC as well as HPV+ cervical squamous cell carcinoma (CSCC), 315 implying that it may serve as an HPV-specific biomarker as well as a general cancer biomarker. The fact that miR-21 overexpression is detectable in plasma makes it even more appealing as a biomarker. 308
Like miR-21, miR-31 is detectable in saliva 316 and has been implicated as an oncogenic miRNA. 317 Several oral cancer tissue profiling studies have found miR-31 to be upregulated,293,304,309,310 and it has also been documented to be upregulated in HPV+ cancers. A study comparing HPV+ oral squamous cell carcinoma (OSCC) patients to HPV– control patients found that miR-31 was the most upregulated miRNA of OSCC biopsies. 318 Interestingly, a follow-up study by the same group comparing miRNA expression in HPV+ or HPV– TSCC tissue revealed that miR-31 was one of the most downregulated miRNAs in HPV+ TSCC, 315 further indicating the potential utility of the miRNA in the differential diagnosis of HPV+ carcinomas.
Other interesting miRNAs were revealed in this study of HPV+ TSCC, including the upregulation of miR-363 and downregulation of miR-375. In accordance with these results, miR-363 has also been shown to be upregulated in both HPV+ and HPV– pharyngeal squamous cell carcinoma (PSCC) and in HPV16+ HNSCC cell lines.318,319 miR-375 has been found to be significantly downregulated in HNSCC and OSCC.300,302,311,320
miR-155 is another miRNA that has been found to be upregulated in several oral cancer profiling studies304,310–312 however, it has been documented as being downregulated when comparing HPV– and HPV+ cell lines or oral cancer tissues with control tissues.294,319 It is possible that this miRNA may behave differently in tissue culture as compared to actual tissue. This was the case for miR-181a, which was found to be downregulated between HPV– versus HPV+ tissue culture, 319 despite that several profiling studies using cancerous tissues have identified miR-181a as upregulated.294,302,304,321 Nonetheless, the consistent appearance of miR-155 in HPV-related conditions is compelling and worth further investigation.
Similarly, miR-203 has been shown to be upregulated or downregulated in different studies. miRNA was downregulated in keratinocyte cell lines expressing HPV oncoproteins,290,291 which is further supported by several profiling studies that used tissues from HNSCC or CSCC.294,296–300,306 Martinez et al, however, found miR-203 to be upregulated when comparing HPV+ and HPV– keratinocytes. 269 Nonetheless, the repeated appearance of miR-203 in HPV-related conditions warrants its inclusion in further studies as a potential biomarker. It is also noteworthy that it has been detected in plasma in patients with laryngeal squamous cell carcinoma (LSCC). 299
Other notably upregulated miRNAs are miR-200c and miR-205. When 28-fold increased, miR-200c was found to be the most overexpressed miRNA in HPV16+ cell lines compared to HPV– cell lines. 269 miR-205 is known to be induced via E7 and has been shown to be upregulated in HPV+ cervical cells 286 and in several oral cancer profiling studies, as well.304,322–324 However, one study has indicated that miR-205 is downregulated in HNSCC and is significantly associated with poor survival. 303
In addition to the upregulated miRNAs mentioned above, several miRNAs have been shown to be downregulated during HPV infection. The expression of miR-143 and miR-145 is reduced in HPV+ cervical cancer cell lines compared to HPV– cell lines.269,286 Both miRNAs are similarly downregulated in tissues from patients with cervical carcinomas, hypopharyngeal squamous cell carcinoma (HSCC), LSCC, or HNSCC.297,298,300,304,306,307,311,325 miR-34a is known to be negatively regulated by the HPV E6 protein, and its expression was downregulated in HPV16+ cervical cells and HPV+ cervical tissue.325–327 It has also been shown to be downregulated in OSCC.294,305
One of the more promising HPV biomarkers may exist in miR-218, also regulated by the HPV E6 protein. It has been identified as significantly downregulated in multiple studies using HPV-infected cells.269,300,319 In fact, a study comparing HPV+ versus HPV– cervical cancer cell lines found miR-218 to be (the only) significantly underexpressed miRNA. 269 It has been shown to be downregulated in HPV+ squamous cell carcinoma of the head and neck cell lines and in the HSCC tissue. Collectively, these studies identify miR-218 as a potential HPV-specific biomarker.
Human immunodeficiency virus
HIV is an enveloped lentivirus belonging to the
In 2004, five putative HIV miRNAs were computationally identified,
332
and it was later reported that an HIV miRNA derived from the
Deep sequencing studies of HIV-infected T cells identified several HIV small RNAs, including an 18-nucleotide RNA antisense to the HIV tRNA primer binding site. 336 Similarly, Schopman et al used deep sequencing to identify small RNAs from HIV-infected T cells. One set of small RNAs originated from secondary hairpin structures formed by the viral genomic RNA and were attributed as a class of miRNAs. Although they were found to be in very low proportion in the total small RNAs, three of them were able to prevent luciferase activity when cloned into a reporter construct, including the previously reported TAR miRNA. Antagomirs also increased HIV replication of infected 293T cells. 337
A recent study by Whisnant et al pointed out that miRNAs <22 nucleotides are more consistent with RNA breakdown products than other miRNAs and that a decisive seed region on the 5′ end of the miRNA must be considered. 20 They also highlighted that low-frequency miRNAs, representing <0.1% of the total miRNA pool, are not likely to be biologically relevant. 338 They examined small RNAs in two HIV-infected cell lines, human primary CD4+ peripheral blood mononuclear cells (PBMCs), and human macrophages, and did not identify any HIV miRNAs. 20 By using PAR-CLIP, they found a small subset of expressed cellular miRNAs that exhibited complementarity to HIV genome sequences. However, they determined that these miRNAs are largely unable to interact with the HIV RNA genome due to extensive viral secondary structures. Since HIV-1 has been implicated in the induction of several cellular miRNAs from infected cells (described below), this finding generates another layer of controversy in the matter. Together, these studies emphasize that additional confirmatory evidence is needed to establish a body of evidence to support or refute whether HIV-1 indeed encodes its own miRNA.
Cellular miRNAs induced by HIV-1 infection
Studies have begun to shed light on the involvement of cellular miRNAs during HIV-1 infection. As mentioned above, additional studies will be needed to verify the biological relevance of these antisense oligonucleotides at the concentrations found in infected cells. For biomarker discovery, the relative presence or absence of miRNAs (or breakdown products) during infection is a more important consideration than their absolute concentration, as long as they are consistently detectable in a validated manner.
During HIV infection, several cellular miRNAs have been reported to be modulated that indirectly impact the replication of the virus. In 2007, Triboulet et al used microarray studies of HIV-infected Jurkat or PBMC to identify 11 cellular miRNAs that were upregulated upon infection and were dependent upon Drosha and Dicer machinery. Viral replication occurred faster in the absence of these miRNAs. Notably, miR-122, miR-297, miR-370, and miR-373* were only expressed in HIV+ cells, while the miR-17/92 cluster (that includes miR-17-5p/3p, miR-18, miR-19a, miR-19b-1, miR-20a, and miR-92-1) was downregulated. 339
A challenge of using PBMCs to examine HIV-associated cellular miRNAs is that miRNAs are expressed differently in the various peripheral blood cells in the context of different environments. In an attempt to address this concern, Chang et al infected the CD4+ SUP-T1 T cell line with HIV-1 or UV-inactivated HIV-1 and used deep sequencing to identify miRNAs. At 24 hours post infection, 65 and 39 cellular miRNAs were upregulated or downregulated, respectively, in the HIV-1-infected cells. 340 Similarly, HIV-1 infection of the CEMx174 hybrid B/T cell line resulted in the increased or decreased expression of 72 and 106 miRNAs, respectively, as assessed by the miRNA microarray. 341 The results varied with different cell types or HIV strains, making comparison of the studies difficult.
In addition to affecting viral replication, cellular miRNAs have also been reported to directly target HIV-1 regulatory and accessory genes. A cluster of cellular miRNAs have been shown to target the 3′ end of the HIV-1 RNA, common to nearly all HIV-1 mRNA 3′-UTRs, to inhibit HIV-1 infection of primary CD4 T cells. Termed the “anti-HIV miRNAs,” this cluster includes miR-28, miR-125b, miR-150, miR-223, and miR-382. 342 Combined inhibition of these five miRNAs increased virus production in CD4 T cells isolated from HIV-1+individuals on highly active antiretroviral therapy (HAART), indicating the miRNAs may be involved in maintaining latent infection in resting CD4 T cells. Furthermore, another group has shown that miR-150 and miR-223 are expressed at high levels in CD4+ T cells isolated from healthy controls compared with the HIV-1-infected individuals. 343 A study using microarray to examine miRNA levels in the PBMC of HIV-1+ individuals with high viral load found that miR-223 was downregulated in this patient population, but miR-150 expression was unchanged. 344
It has been speculated that the anti-HIV miRNAs may be reduced in the differentiation of monocytes to macrophages, thereby creating an HIV-promoting intracellular environment. However, microarray studies showed that only miR-223 exhibited downregulation upon monocyte differentiation into macrophages. Meanwhile, miR-28-3p expression only slightly declined, miR-125b and miR-382 did not differ from background levels, and miR-150 was strongly upregulated. 345
Several studies have examined the role of the mi-R29 family during HIV infection, and target sites for miR-29a and miR-29b have been predicted within the HIV-1
miRNAs as potential biomarkers of HIV infection
Several groups have examined the regulation of cellular miRNAs during HIV infection to validate the potential of miRNAs as biomarkers (Table 11). Moreover, recent studies have specifically examined the potential of miRNAs to distinguish between different groups of HIV-infected individuals. Munshi et al recently analyzed miR-16, miR-146b-5p, miR-150, miR-191, and miR-223 levels in the PBMCs and serum of HIV/AIDS patients in different stages of disease. miR-150 and miR-146b-5p were found to distinguish HIV+ individuals as well as those on HAART or developing drug resistance. The levels of miR-150 were found to be decreased in the PBMCs of HIV/AIDS patients before treatment, and HAART was able to restore levels of miR-150. Individuals who developed drug resistance reduced miR-150 expression to the levels observed in the symptomatic HIV/AIDS patients. 349 In contrast, plasma levels of miR-150 were increased during infection and reduced upon treatment, prompting the authors to propose that miR-150 in the plasma may be derived from damaged tissues or released exosomes. A positive trend was also generally observed between PBMC miR-150 and CD4 T cell counts.
Biomarker potential of host miRNAs associated with HIV-1 infection (>2-fold change in miRNA): selected studies.

Some individuals infected with HIV, known as elite suppressors (ES), maintain an undetectable viral load and high CD4 T cell counts in the absence of any treatment. A study that characterized the miRNAs present in the PBMCs of ES versus untreated HIV/AIDS patients determined that the two groups actually share expression patterns of several miRNAs, although a few miRNAs were differentially expressed. miR-31, miR-31*, and miR-150 were significantly downregulated, while miR-155 and miR-22 were moderately upregulated in active HIV/AIDS versus ES patients.
350
This study also found a correlation of particular miRNAs with CD4 T cells counts in infected individuals compared to uninfected controls: miR181b (negative) and miR-31, miR-31*, miR-29a, and miR-150 (positive) all significantly correlated with CD4 T cell count (
While viral titers and CD4 T cell counts are important for predicting disease progression, it has been suggested that CD4 T cell counts do not necessarily correlate well with virologic failure in patients being treated with ART, 352 emphasizing the importance of the characterization and validation of potential HIV biomarkers.
Potential miRNA therapeutics for HIV
At first thought, the cluster of anti-HIV miRNAs (miR-28, miR-125b, miR-150, miR-223, and miR-382) is of interest as potential antiretroviral drugs due to their reported ability to target HIV-1 mRNAs. However, several of these miRNAs are expressed in normal CD4 T cells, 353 which are still capable of being infected by HIV. Recently, Whisnant et al found that HIV-1 transcripts are refractory to miRNA binding, likely due to secondary structure of viral mRNAs. 20 This implied that these anti-HIV miRNAs may not be able to target the virus directly. This does not preclude their possible role, however, in modulating expression of genes that could create a cellular environment that is not susceptible or permissive to infection. Considering that a single miRNA is capable of affecting the transcription of many genes and therefore possibly inducing off-target effects, additional research is warranted concerning the utility of miRNAs as antiretroviral drugs.
Concluding Remarks
Over 200 biomarker-related clinical trials involving miRNAs are listed on the Clinicaltrials.gov website (https://clinicaltrials.gov/ct2/results?term=miRNA), demonstrating that miRNAs continue to have great potential as biomarkers of diagnosis, disease progression, and treatment efficacy. As such, the use of miRNA biomarkers during diseases of viral origin also warrants further investigation. Several DNA viruses of clinical importance, namely, herpesviruses and polyomaviruses, encode unique viral miRNAs that could prove promising to differentiate viruses that have similar clinical symptoms. The use of miRNAs specifically expressed at different stages of infection could also help in accurately determining disease status. For the RNA viruses or retroviruses described above, profiling panels of cellular miRNAs could provide a means of identifying viruses or classifying disease stage. miRNAs that can be detected in PBMCs, plasma, or exosomes and mirror expression at the local site of infection or pathology are particularly attractive, providing a means to avoid invasive biopsies and better monitor disease using less-invasive methods. However, future miRNA assays based upon next-generation sequencing will require additional technical expertise and standardization. In addition, the presence of specific miRNAs in individuals of different backgrounds has not been thoroughly studied, and the conservation of miRNAs between clinical strains of the same virus also needs to be further explored.
Elucidating the biological functions of specific miRNAs has allowed us to contemplate the role of specific miRNAs as therapeutics or as therapeutic targets. However, drugs based upon RNAi have not been without multiple technical difficulties, including vehicle design, antisense oligonucleotide bio-stability, targeted delivery to cells of interest (and to necessary intracellular compartments), and off-target effects. Nonetheless, new strategies have been and continue to be developed to combat these initial roadblocks and build better therapeutics.354,355 Preliminary results from Phase 2a studies of miravirsen, an LNA that targets miR-122 during HCV infection, are cautiously promising. 255 The results of additional clinical trials of miravirsen and other antagomirs will be of great interest in ascertaining the therapeutic potential of miRNAs and their inhibitors.
Author Contributions
Made critical revisions and approved the final version: JL, MB. All listed authors wrote sections of the manuscript and approved the final manuscript.
