Abstract
Introduction
In Dahl rats’ kidney cortex, the alternatively spliced form of the epithelial sodium channel α subunit (α ENaC-b) is the most abundant mRNA transcript (32+/-3 fold > α ENaC-wt) as was investigated by quantitative RT-PCR analysis. α ENaC-b mRNA levels were significantly higher in Dahl R versus S rats, and were further augmented by high salt diet.
Objectives
In the present study, we described the molecular cloning and searched for a possible role of α ENaC-b by testing its potential expression in COS7 cells as well as its impact on α ENaC-wt expression levels when co-expressed in COS7 cells in a dose-dependent manner.
Methods
Using RT-PCR strategy, the full-length wildtype α ENaC transcript and the alternatively spliced form α ENaC-b were amplified, sequenced, cloned, subcloned into PCMV-sport6 expression vector, expressed and co-expressed into COS7 cells in a dose-dependent manner. A combination of denaturing and native western blotting techniques was employed to examine the expression of α ENaC-b in vitro, and to determine if an interaction between α ENaC-b and α ENaC-wt occurs in vitro, and finally to demonstrate if degradation of α ENaC-wt protein does occur.
Results
α ENaC-b is translated in COS7 cells. Co-expression of α ENaC-b together with α ENaC-wt reduced α ENaC-wt levels in a dose-dependent manner. α ENaC-wt and α ENaC-b appear to form a complex that enhances the degradation of α ENaC-wt.
Conclusions
Western blots suggest a novel mechanism in α ENaC regulation whereby α ENaC-b exerts a dominant negative effect on α ENaC-wt expression. This is potentially by sequestering α ENaC-wt, enhancing its proteolytic degradation, and possibly explaining the mechanism of salt-resistance in Dahl R rats.
Keywords
Introduction
The amiloride-sensitive epithelial sodium channels (ENaC) consists of at least three subunits denoted by α, β, and γ, each of which possesses two transmembrane domains.1–3 The α ENaC subunit is a key molecule for the formation of a functional Na+ channel complex in vivo, whereas the β and γ subunits greatly potentiate the channel activity. 3 α ENaC gene knockout in mice was lethal within 40 h of birth because of fluid-filled lungs, confirming the critical role of α ENaC in forming functional Na+ channels in vivo. 4 Naturally occurring α ENaC alternatively spliced forms have been identified in mice, chicken, humans and rats.5–8 To date, there are two alternatively spliced forms of the α subunit of ENaC (α ENaC-a and -b) that have been published in rats, in addition to the major α ENaC transcript. 7 α ENaC-a transcript is a low abundance transcript compared to the full-length α ENaC and has been studied in terms of expression, functionality and binding to ENaC blocker. 7 On the other hand, α ENaC-b is yet to be characterized and appears to play a role in modulating ENaC for the following reasons: a) it shares the same splice site with human and chicken spliced forms; b) our preliminary results showed that unlike α ENaC-a, α ENaC-b mRNA levels are ~32 ± 3-fold higher than full-length α ENaC in kidneys of Dahl rats indicating an increased stability/synthesis of the former; 9 c) α ENaC-b mRNA concentrations are significantly higher in Dahl R versus Dahl S rats, 9 suggesting a putative dominant negative effect on α ENaC activity in a model of suppressed ENaC activity (Dahl R rats); 10 d) α ENaC-b is a salt-sensitive transcript, because high versus normal salt diet caused a remarkable increase in α ENaC-b mRNA levels (P < 0.05) in Dahl R rats. 9
Owing to the fact that non functional α ENaC alternatively spliced forms, such as α ENaC-b, have been proposed to serve as negative regulatory components for ENaC activity, 8 we were prompted to invest in basic research with a view to identifying novel targets for diagnosis, prevention and treatment of disorders related to ENaC dysfunction. The overall objectives of these studies were to investigate whether α ENaC-b is translated in vitro, whether it is co-expressed with α ENaC-wt, and whether α ENaC-b expression suppresses α ENaC-wt expression when co-expressed in COS7 cells in a dose-dependent manner.
Methods
RNA extraction, RT-PCR and PCR
Kidney cortex of Dahl S rats was cut out microscopically and homogenized with polytron. Poly (A)+ mRNA from kidney cortex of Dahl S rats was isolated using Trizol reagent (Invitrogen, CA, U. S.A.) according to the manufacturer's protocol. Isolated RNA was subjected to DNAse treatment for removing potential genomic DNA contamination using DNase treatment kit (Ambion, ON, Canada). First strand cDNA was synthesized from 2 μg of mRNA with superscript II RNase H-Reverse transcriptase (Invitrogen, CA, U.S.A.).
For α ENaC-wt and α ENaC-b, specific primers (Table 1) were designed to flank the open reading frames of α ENaC-wt and -b form. High fidelity Expand Long Range, dNTPack (kind gift from Roche Applied Science®, QC, Canada) is employed to amplify the full length α ENaC and α ENaC-b. Touchdown PCR method was employed to improve the efficiency of gene amplification as follows (start temperature (Tm) at 68 °C and the annealing Tm reduced 2 °C per cycle for the next cycles up to Tm of 58 °C, then the remaining cycles were performed at a 58 °C annealing temperature). After amplification, the specific fragments of 2267 bp (α ENaC-wt) and 1586 bp (α ENaC-b) were visualized in 1% agarose gels (Fig. 1).
Primers used for reverse transcription polymerase chain reaction and expected sizes of cDNA fragments.
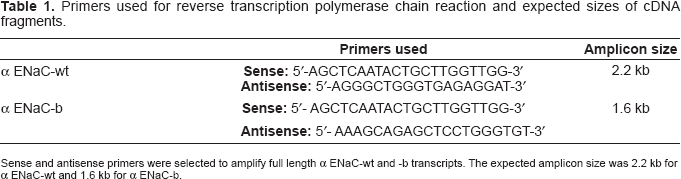
Sense and antisense primers were selected to amplify full length α ENaC-wt and −b transcripts. The expected amplicon size was 2.2 kb for α ENaC-wt and 1.6 kb for α ENaC-b.
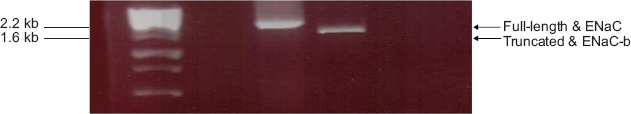
Amplification of ENaC-wt and alternatively spliced form from total RNA. Total RNA extracted from kidney cortex of Dahl S rats was utilized as a template to generate full lengths cDNAs of α ENaC-wt and alternatively spliced form α ENaC-b. High fidelity Expand Long Range, dNTPack was used to amplify α ENaC-wt and -b form. Touchdown PCR method was employed to improve the efficiency of gene amplification. After amplification, the specific fragments of 2267 bp (α ENaC-wt) and 1586 bp (α ENaC-b) were visualized in 1% agarose gels.
Plasmids and generation of constructs
PCR products for α ENaC-wt and α ENaC-b were ligated with pCR®2.1-TOPO® vector using the TOPO TA® cloning kit (Invitrogen, CA, U.S.A.), with blue/white screening of DH5α competent cells on ampicillin selective plates. A total of 14 individual colonies for each species were isolated. Positive clones were picked and cultured in LB broth. The DNA extracted from these cultures using Qiagen DNA miniprep kit, was sequenced using the M13 universal forward and reverse primers, in addition to embedded primers within α ENaC-wt and-b forms to verify their full lengths sequences. Sequencing was performed using the DYEnamic ET Terminator kit according to the instructions provided by the manufacturer (PE Applied Biosystems, Foster City, CA). Sequencing products were purified (DyeEx 2.0 spin kit columns; Qiagen Canada, Mississauga, ON, Canada) and analyzed on 3100 DNA analyser (Applied Biosystems, Foster City, CA). The sequence of the cloned α ENaC-b is identical to the one previously reported, 7 and the sequence of the α ENaC-wt is identical to the one previously reported by our group for Dahl rats. 11
Ligated pCR®2.1-TOPO®-α ENaC-wt clone was digested using EcoRI, while that of α ENaC-b was digested using EcoRV and HindIII. Digested α ENaC-wt and -b fragments were purified from 1% agarose gel (QIAquick Gel Extraction Kit, Qiagen™, ON, Canada) and then subcloned overnight at 16 °C into PCMV-sport6 vector (Invitrogen, CA, U.S.A.) downstream the T7 promotor using the EcoRI site for α ENaC-wt and the EcoRV and HindIII sites for α ENaC-b. The positive clones were verified by restriction digestion, then PCR of the inserted construct and later by direct nucleotide sequencing as described above. DNA yield from positive clones was later increased for transfection studies using Qiagen midi-prep kit (Qiagen™, ON, Canada) for amplification of the yield.
Cell culture and transfection
COS-African green monkey kidney cells were maintained in culture at 37 °C/5% CO2 using Dulbecco's Modified Eagle's Medium (DMEM, Hyclone® Thermo Fisher Scientific Laboratories, Logan, UT) supplemented with glutamine and 10% heat-inactivated fetal bovine serum (FBS, PAA Laboratories, Pasching, Austria). Transfections were performed using lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, U.S.A.) and up to 35 μg DNA per 10 cm plate in co-transfection experiments. COS7 cells were seeded at approximately one third confluency on 10 cm diameter plates 18 h before the transfection. Immediately before transfection, the culture medium was replaced with 5 ml of DMEM supplemented with 10% (v/v) fetal bovine serum. PCMV-sport6 /αENaC-wt and -b were transfected into COS7 cells separately at a concentration of 25 μg DNA /plate (Fig. 2), as well as co-transfected together in a dose-dependent manner as demonstrated in Figures 3 and 4. The empty PCMV-sport6 vector at a concentration of 25 μg DNA /plate was transfected as a control. DNA was added to 250 μl TE buffer and lipofectamine was then added (Invitrogen®, ON, Canada). The mixture was incubated at room temperature for 20 min before addition to the COS7 cell culture. The cells were incubated overnight at 37 °C in an air/CO2 (19:1) atmosphere before the medium was aspirated and the COS7 cell culture osmotically shocked for 2 min with 10% (v/v) PBS.

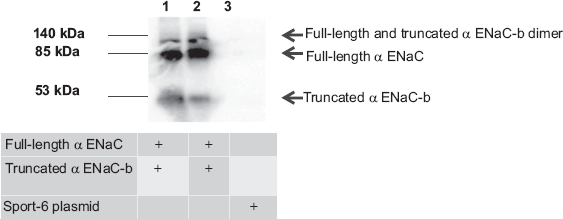
Expression of α ENaC-wt and α ENaC-b in COS7 cells. SDS/PAGE and Western blot analysis were performed using extracts from mammalian cells expressing either the alternatively spliced form α ENaC-wt or -b form or the empty vector in COS7 cells. Lane 1 represents proteins extracted from COS7 cells transfected with α ENaC-wt, lanes 2 and 3 represent COS7 cells transfected with the empty PCMV-sport6 vector, and lane 4 represents COS7cells transfected with α ENaC-b.
Analysis of α ENaC-wt and α ENaC-b expression by native Western analysis. α ENaC-wt and α ENaC-b interacts in vivo by the formation of a dimer of a molecular mass of 140 kDa. Lanes 1 and 2 represent proteins from COS7 cells co-expressing α ENaC-wt and α ENaC-b in equal proportions (1:1), lane 3 represents proteins from COS7 cells transfected with the empty PCMV-sport 6 vector. Bands are normalized against the house keeping gene β-actin and the experiment was done in triplicate.

Dose-dependent suppression of α ENaC-wt by increasing doses of α ENaC-b. Lanes 1–7 represent proteins of COS7 cells co-transfected by α ENaC-wt and α ENaC-b in the ratios provided. Denaturing Western analysis demonstrate a dose-dependent reduction in full-length α ENaC expression. Lane 8 represents proteins from cells transfected with the empty PCMV-sport 6 vector. A weak signal of α ENaC-b can be detected at 53 kDa, possibly indicating that dimerization of α ENaC-wt with α ENaC-b proteins degrades α ENaC-wt primarily and eventually degrades α ENaC-b. Bands are normalized against the house keeping gene β-actin and the experiment was done in triplicate.
Electrophoresis analysis
We used two different Western blot analyses namely SDS-PAGE or “denaturing gels” and native PAGE or “non denaturing gels”. While SDS-PAGE denatured α ENaC-wt and -b proteins, and allowed an easy access of the α ENaC antibody to the reactive epitope and subsequent separation of the α ENaC proteins based on molecular mass, native gel electrophoresis, on the other hand, allowed us to study the α ENaC-wt and -b proteins oligomerization potential. Migration for both SDS-PAGE and native gels was performed under 150 V and for 120 min.
SDS-PAGE and Western Blot analyses
Cells from transiently transfected COS7 cells expressing α ENaC-wt, or α ENaC-b, or the empty vector along with COS7 cells equally co-transfected with α ENaC-wt and -b form were washed twice with ice-cold PBS, scraped and then maintained for >2 h at 4 °C in RIPA lysis buffer (10 mM NaPO4, 150 mM NaCl, 1% deoxycholate, 1% Triton X-100, and 0.1% SDS, pH 7.2) supplemented with the protease inhibitor cocktail (Sigma, St. Louis, MO). Protein concentrations were measured by Bradford. Proteins were then separated by SDS-PAGE resolving gels containing 10% acrylamide, and 4% of the same solution for stacking gels. Samples were mixed with Laemmli buffer (Bio-Rad, 0.5 M Tris-Hcl, pH 6.8, 10% glycerol, 0.02% bromophenol blue) containing 0.1% SDS [sodium dodecyl-sulphate (w:v)] and supplemented with β-mercaptoethanol (v:v). Before loading, SDS samples were heated at 100 °C for 5 minutes. Proteins were separated by SDS-PAGE and western blots were performed according to procedures recommended by the Bio-Rad Protein III system (Bio-Rad Inc.).
Native gel electrophoresis and Western Blot analyses
Separation of proteins by PAGE under non denaturing conditions was performed similar to SDS-PAGE, except that SDS and β-mercaptoethanol were totally excluded and the samples were not boiled. This will preserve any protein interactions. Samples and protein markers were diluted in 0.5 M Tris-HCl, pH 6.8, 40% glycerol (v:v). and 0.02% bromophenol blue (w:v) (without SDS and β-mercaptoethanol), and without heating. In Native PAGE, 20 μg of proteins were separated on precast Novex® 8% Tris-Glycine Midi Gel (Invitrogen, CA, U.S.A.). Before loading the samples, the gel was cooled to 4 °C. After addition of native sample buffer (5X), samples were directly applied to the gel without any preconditioning. Native gel electrophoresis (18 cm × 16 cm) was conducted in the same apparatus as above. The running buffer (pH 8.3) contained Tris base (25 mM) and glycine (192 mM).
Polyclonal antibodies raised against the N terminal aa 46–68 [NH2-LGKGDKREEQGL-GPEPSAPRQPT-COOH] of the rat α ENaC were used to recognize the α ENaC-wt and -b proteins. Antibodies were purified from rabbit sera, concentrated to 1.3 ug/ul and diluted to 1:1000. Primary α ENaC antibodies were a kind gift from Dr. M. Amin. In Western blot, protein samples were transferred to nitrocellulose membranes (Bio-Rad Inc.) using Trans-Blot Cell system (Bio-Rad Inc., CA, U.S.A.). After transfer, nitrocellulose membranes were blocked for 1 h with 5% skimmed milk in 0.1% Tris-buffered saline containing 0.1% Triton X-100 (TBST) and incubated with primary antibody overnight. Blots were probed with horseradish peroxidase-conjugated anti-rabbit IgG secondary antibody (Promega Inc., ON, Canada) for 2 h at room temperature. Multiple washes for the membranes were performed using TBST after the primary and secondary incubations. Protein signals were detected using ECL substrate (Amersham Biosciences Inc. Chicago, IL) and quantified using Alpha Ease FC (FluorChem, 9900) software. All Western blots were stripped in 100 mm 2-mercaptoethanol, 62.5 mM Tris-HCl (pH 6.7), and 2% SDS for 30 min at 55 °C with constant agitation. After removal of antibodies, nonspecific interactions were reblocked by incubation in Tris-buffered saline-Tween 20 and 5% milk for 2 h before the blots were reprobed with the house keeping gene (β-actin).
Results
Molecular cloning and sequencing of α ENaC-wt and alternatively spliced form α ENaC-b
To determine whether α ENaC-b is translated in vitro, and if so, whether α ENaC-b binds to α ENaC-wt, we cloned α ENaC-wt and -b transcripts from Dahl S rats’ kidney cortex, using RT-PCR. Using PCR primers that were designed to amplify full-length coding sequences of α ENaC-wt and α ENaC-b transcripts, α ENaC-wt and α ENaC-b were obtained and cloned (Fig. 1). Sequences for both α ENaC-wt and -b form were confirmed by complete nucleotide sequencing analyses. α ENaC-wt sequence was consistent with our previously reported Dahl rat coding sequence. 11 The present α ENaC-b sequence was consistent with the previously reported α ENaC-b sequence. 7
Expression of full length α ENaC and truncated α ENaC-b alternatively spliced form in transiently transfected COS7 cells
Full length α ENaC and truncated α ENaC-b transcripts were studied in COS7 cells to confirm their ability to make a protein. In vitro translation of α ENaC-wt and α ENaC-b produced proteins of 85 and 53 kDa, respectively (Fig. 2).
Dimerization of α ENaC-wt and α ENaC-b
Using native Western analysis for lysates obtained from COS7 cells co-transfected with α ENaC-wt and -b forms, a dimer was formed at 140 kDa, equivalent to the summation of the molecular weights of α ENaC-wt and α ENaC-b. α ENaC-wt protein has a molecular mass of 85 kDa, while α ENaC-b protein has a molecular mass of 53 kDa (Fig. 3).
Dose-dependent suppression of full-length α ENaC expression by increasing doses of α ENaC-b
Western blot analysis using an α ENaC antiserum revealed that co-expression of α ENaC-wt with spliced form α ENaC-b decreased α ENaC-wt protein levels by increasing doses of α ENaC-b protein (Fig. 4).
Discussion
The present study reports the following: a) α ENaC-b transcript is translated in COS7 cells to a peptide of apparent mass of 53 kDa, b) α ENaC-b transcript is co-expressed with α ENaC-wt transcript, c) α ENaC-b forms a complex with α ENaC-wt resulting in dimer formation of a molecular mass of approximately 140 kDa, d) Increasing doses of α ENaC-b suppressed α ENaC-wt expression in a dose-dependent manner when co-expressed in COS7 cells.
Formation of alternatively spliced forms of α ENaC have been previously reported in four species: rats, humans, mice and chicken6–9,12 suggesting that alternative RNA splicing is possibly a mechanism for regulating α ENaC activity, besides transcriptional regulation. The biological impact of alternatively spliced forms, particularly those lacking functional domains such as α ENaC-b lacking the second transmembrane domain, may go as far as a drastic switchoff effect.13–15
Our previous findings showed that α ENaC-b mRNA levels were significantly higher in Dahl salt-resistant (R) versus salt-sensitive (S) rats. 9 Overrepresentation of α ENaC-b in Dahl R rats could be indicative of a dominant negative effect on α ENaC expression/function in this model. Additionally, owing to the fact that α ENaC-b is a saltsensitive transcript in Dahl R rats, 9 we were motivated to search for a biological role of α ENaC-b in vitro that might be in progress to protect the channel from over-activity in high salt environment in Dahl R rats, possibly by a dominant negative effect on α ENaC-wt. Therefore, α ENaC-b was amplified, sequenced and cloned. Expression of α ENaC-b alone and in the presence and absence of α ENaC-wt was examined.
Results show that the truncated α ENaC-b protein is translated in vitro and its co-expression with α ENaC-wt appeared to substantially reduce α ENaC-wt expression in a dose-dependent manner. α ENaC-b acts as a dominant inhibitor of α ENaC-wt by forming a complex with α ENaC-wt. These results support an important role for alternative splicing in α ENaC regulation, and suggest that α ENaC can form heteromers which may be important in regulating α ENaC expression and/or activity.
The present study is the first to demonstrate the expression of alternatively spliced α ENaC-b form in mammalian COS7 cells and expands our understanding of the putative mechanism of action of α ENaC-b as a negative expression regulator of α ENaC-wt; by directly interacting with α ENaC-wt as a dominant negative protein that ultimately hinders α ENaC-wt expression and potentially its activity. In co-expression studies (Fig. 4), a weak signal of α ENaC-b can be detected at 53 kDa, possibly indicating that dimerization of α ENaC-wt with -b protein degrades α ENaC-wt primarily and eventually degrades α ENaC-b. Our findings therefore uncover a new mechanism that may control the regulation of α ENaC expression in vivo by a potential spliced form α ENaC-b. A detailed understanding of the spectrum of alternatively spliced α ENaC-b is of importance not only for the understanding of ENaC regulation in Dahl rats, but also for the future drug development efforts. It will also provide insight into the beneficial impact of increasing α ENaC-b to possibly prevent salt sensitivity.
To this end, the present study suggests a dominant negative effect imposed by α ENaC-b on full length α ENaC protein expression in Dahl R rat kidneys. Our preliminary results suggest the intriguing possibility of a “genotoxic” effect of α ENaC-b on full-length α ENaC, potentially by accelerating proteolytic degradation of the latter in a dose-dependent fashion. There are at least two areas of potential clinical importance for these studies. First, identification of spliced forms that impair the hyperactivity of ENaC to sodium in the kidney of Dahl S rats would identify specific targets for the development of novel antihypertensive drug therapy for salt-dependent hypertension. Second, as gene therapy delivery systems continue to evolve, we will be examining in vivo delivery of adenoviruses carrying α ENaC-b and then assessing the resultant blood pressure effects. This may eventually gain relevance as a possible “proof-of-concept” gene therapy.
Footnotes
Acknowledgements
The author acknowledges Roche Diagnostics® Canada for generously providing free samples of Taq polymerase enzyme. Additionally, the author is grateful to Qiagen® Canada for providing free samples of the DyeEx 2.0 spin kit columns used for sequencing. MF Shehata is the recipient of the Memorial University of Newfoundland Horizon Award for 2007. MF Shehata has received numerous awards including the Canadian Institutes of Health Research/Pfizer/Canadian Hypertension Society Doctoral Research Award, Ontario Graduate Scholarship and the Ontario Graduate Scholarship for Science and Technology.
Disclosure
The author reports no conflicts of interest.
