Abstract
Background
Patients at risk of breast cancer are submitted to mammography, resulting in a classification of the lesions following the Breast Imaging Reporting and Data System (BI-RADS®). Due to BI-RADS 3 classification problems and the great uncertainty of the possible evolution of this kind of tumours, the integration of mammographic imaging with other techniques and markers of pathology, as metabolic information, may be advisable.
Design and methods
Our study aims to evaluate the possibility to quantify by gas chromatography-mass spectrometry (GCMS) specific metabolites in the plasma of patients with mammograms classified from BI-RADS 3 to BI-RADS 5, to find similarities or differences in their metabolome. Samples from BI-RADS 3 to 5 patients were compared with samples from a healthy control group. This pilot project aimed at establishing the sensitivity of the metabolomic classification of blood samples of patients undergoing breast radiological analysis and to support a better classification of mammographic cases.
Results
Metabolomic analysis revealed a panel of metabolites more abundant in healthy controls, as 3-aminoisobutyric acid, cholesterol, cysteine, stearic, linoleic and palmitic fatty acids. The comparison between samples from BI-RADS 3 and BI-RADS 5 patients, revealed the importance of 4-hydroxyproline, found in higher amount in BI-RADS 3 subjects.
Conclusion
Although the low sample number did not allow the attainment of high validated statistical models, some interesting data were obtained, revealing the potential of metabolomics for an improvement in the classification of different mammographic lesions.
Significance for public health
The breast cancer risk is evaluated after mammographic exam by the BI-RADS classification of lesion. The BI-RADS 3 classified cases comprise a wide class of lesions and their treatment must be subjected to multidisciplinary discussion and consideration. Metabolomic analysis of plasma from subjects undergoing mammography may give new information on metabolite content and allow a better classification for BI-RADS 3 cases.
Introduction
Breast cancer is considered a major Public Health concern for its high morbidity and mortality rates: EUROSTAT reports that breast cancer accounted for 1.8% of all deaths in the EU-28 in 2015 and 3.6% of deaths in women. 1 The breast cancer risk is evaluated worldwide by the Breast Imaging Reporting and Data System (BI-RADS®) since its introduction in 1992 by the American College of Radiology. The BI-RADS System was designed to serve as a guide to provide standardized terminology in breast XR imaging.2,3 This recommended reporting structure includes final assessment categories with management recommendations and a framework for data collection and auditing. The BIRADS ® for mammography was designed to standardize breast image reports and to reduce confusion in breast image interpretations. It also facilitates the monitoring of results and quality assessment. But, despite the initial wide diffusion of the System, and considering the effort to produce free-access tool available on the web for the BI-RADS reporting numerous reports about the not univocal tumour classification by this system, as it is used, are reported with challenging to reproduce classification, especially for the categories associated with BI-RADS 3 classes.
Indeed, BI-RADS provides a highly questionable positive predictive reference (PPV) ranges for 3 (<2%), for 4 (2-95%), and in some cases provides subclassification (4a, b, c) considered as unnecessary by several authors. Due to this uncertainty, radiologists continue to have different PPVs for identical lesions evaluated by the Radiological Mammography.
The increasing participation of the population to breast screening programs has led to an increase in the diagnosis of the mammographic lesions, particularly for the lesions classified with a degree of malignancy (BI-RADS 3) in the BI-RADS classification system. According to the most recent literature data, the risk of malignancy of these kinds of lesions varies in the range 9.9-35.1%. The benign lesions group includes atypical lobular hyperplasia (LIN 1), classic lobular carcinoma in situ (LIN 2) and pleomorphic lobular carcinoma in situ (LIN 3). 4 Further, there are the ductal carcinoma in situ (DCIS), with different grading, DCIS of low nuclear grade, DCIS of intermediate nuclear grade and DCIS of high nuclear grade. 4
To complete the classification of the benign lesion, sclerosing lesions (sclerosing adenosis, radial scar), benign phyllodes tumour lesions (the most phyllodes tumors are benign, but in rare cases, they can be malignant), breast micro glandular adenosis, mucocele- like lesions and adenomyoepithelioma, must be included.5,6 Some of these kinds of lesions are usually classified in the BIRADS 3 group.
Due to BI-RADS 3 classification problems, the great uncertainty of the possible evolution of these kinds of tumours and the need of integration of mammographic imaging with other techniques and markers of pathology, as metabolic information, must be recommended.
This study aims to quantify plasma metabolites in patients with mammograms classified from BI- RADS 3 to BI-RADS 5 and assessing if there may be a similarity between the metabolome of patients with BI-RADS 5 lesions (certainly malignant after VAB Vacuum Assisted Biopsy) and patients with BI-RADS 3 lesions (with uncertain potential for malignancy after VAB). Such a broad classification uncertainty should be corrected by a multidisciplinary approach to the pathology classification. In the last years many metabolomic studies on breast cancer tried to identify useful biomarkers for early diagnosis and pathology degree classification: different bio-samples were analysed (plasma, serum, urine, saliva, bioptic tissue) employing different analytical platforms (LC- or GC-MS, NMR) but, despite the large number of data and useful suggestions, no definitive and univocal molecular biomarker has been identified. 7 In this paper, the Metabolomic support to the breast cancer diagnosis and classification is presented. This pilot project aimed at establishing the sensitivity of the metabolomic classification of blood samples of patients undergoing breast radiological analysis, to improve the tumour classification and follow up of these subjects.
Design and Methods
Study population
The study was conducted at the AOU Cagliari University Hospital-Italy, from June 2016 to October 2017. The written informed consent was obtained from subjects before inclusion in the study. All procedures were in accordance with the Helsinki Declaration of 1975, as revised in 2008. Clinical data of the study population are reported in Table 1.
Clinical characteristics of the study population. Group 1: cases, women submitted to mammography; group 2: healthy controls.

Blood samples were collected from 38 patients submitted to radiological mammography (Amulet Innovality, Fujifilm) in the clinical laboratories of the Radio diagnostic Complex Structure of AOU Cagliari University Hospital (Italy): 32 subjects were diagnosed with BI-RADS 3 to 5 classification. Control samples were collected from 10 healthy subjects. All blood samples were centrifuged at 2000 rpm for 10 minutes; the surnatant plasma was transferred in Eppendorf safe-lock tubes and immediately frozen and stored at -80°C until analysis.
Sample preparation
Plasma samples were analysed as reported. 8 In brief, samples were thawed at 4°C, and 400 μl were treated with methanol, mixed with a vortex mixer and then centrifuged. The upper phase was transferred in glass vials and evaporated to dryness in an Eppendorf vacuum centrifuge. Fifty μl of methoxylamine hydrochloride (0.24 M in pyridine) were added to each sample and left to react for 17 h at room temperature. Then 50 μl of MSTFA (N-Methyl-N-trimethylsilyltrifluoroacetamide) were added and left to react for one h at room temperature. Samples were diluted with 100 μl of hexane containing the tetracosane (0.015 mg/ml) internal standard and analysed on an Agilent 5977B Mass Spectrometer interfaced to the GC 7890B equipped with a DB-5ms column (J & W). Each acquired chromatogram was analysed using the free software AMDIS (Automated Mass Spectral Deconvolution and Identification System; http://chemdata.nist.gov/mass-spc/amdis) supported by an in-house made library, including 300 metabolites. This strategy allowed for the detection of 81 compounds: following the identification levels defined by the Metabolomics Standards Initiative (MSI), 9 60 were “confidently identified compounds” (level 1), 8 “putatively annotated compounds” (level 2), 5 “putatively annotated compound class” (level 3), and eight unknown compounds. AMDIS analysis produced an electronic sheet data matrix (Microsoft® Excel®, Microsoft Co., Washington DC, USA) that was submitted to statistical analysis as previously described. 10
Statistical analysis
The AMDIS data matrix was processed with the integrated web-based platform MetaboAnalyst (http://www.metaboanalyst.ca/). 11 Missing values were replaced with half of the minimum positive value. After normalization by sum, data were log-transformed and then categorized using the Pareto scaling procedure. Statistical procedures include univariate analysis, partial least square discriminant analysis (PLS-DA) and orthogonal partial least square discriminant analysis (OPLS- DA). Variable importance in projection (VIP) score for each model was calculated. PLS-DA models were tested with the leave-one-out cross-validation (LOOCV) method for the evaluation of statistical parameters (correlation coefficient R 2 , cross-validation coefficient Q 2 ), which allowed us to determine the optimal number of components for the model description.
Results
The first statistical analysis examined group P, pathologic subjects, compared to group C, healthy controls. Univariate analysis (t-test) revealed four metabolites as statistically different between groups: 3-aminoisobutyric acid, fructose, 3-hydroxybutyric acid and 2-hydroxybutyric acid. The first was found more abundant in controls, while the others were more abundant in the pathological group. The PLS-DA model reveals an overlapping between samples but reaches an acceptable level of statistical significance, as reported in Figure 1A. The model reveals all metabolites as more abundant in pathologic samples except for 3-aminoisobutyric acid, cholesterol, stearic, linoleic and palmitic fatty acids, cysteine, and phosphate.
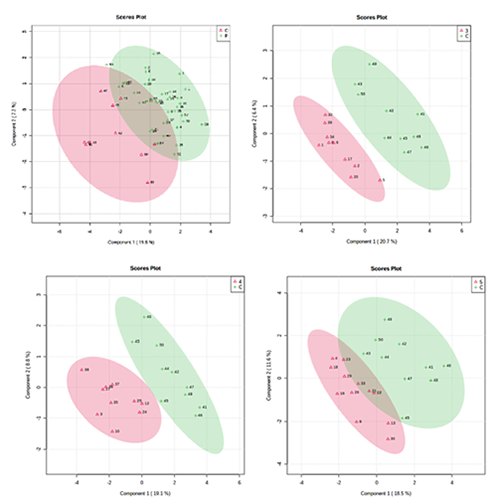
PLS-DA score plot between the first two components of the model: A) Pathological subjects P (green) vs healthy subjects C (red) (accuracy=0.79167; R2=0.41472; Q2=0.16346. B) BI-RADS 3 subjects (red) vs healthy subjects C (green) (accuracy=0.7; R2=0.61611; Q2=0.29495). C) BI-RADS 4 subjects (red) vs healthy subjects C (green) (accuracy=0.7; R 2 =0.66387; Q2=0.33798). D) BI-RADS 5 subjects (red) vs healthy subjects C (green) (accuracy=0.63636; R2=0.51355; Q2=0.0068).
The low level of Q2 (Q2=0.16) reveals that a simple model with two classes for plasma samples from subjects marked by BIRADS score from 3 to 5, together with the great uncertainty for the attribution of BI-RADS score in some cases, justify the low predictive power of the model also indicating the need to stratify the samples by the BI-RADS score, providing the comparison of each BIRADS class with the control group.
The comparison between BI-RADS 3 subjects (10) and controls (10) resulted in the PLS-DA model described in Figure 1B. The comparison between BI-RADS 4 subjects (10) and controls (10) generated the PLS-DA model described in Figure 1C. The comparison between BI-RADS 5 subjects (12) and controls (10) generated the PLS-DA model described in Figure 1D.
Table 2 reports the most important metabolites (VIP score >1.0) for the above reported PLS-DA models, with the corresponding trend.
PLS-DA most important metabolites (VIP = variable importance in the projection; VIP score > 1) and the relative abundance differences: ↑ more abundant in pathological class (all pathological samples (P), BI-RADS 3(3), BI-RADS 4(4), BI-RADS 5(5) compared to controls C); ↓ less abundant in pathological compared to controls. Chemical class: AA (Amino acid), HA (Hydroxy acid), A (Acid), FA (Fatty acid), PO (Polyol), Am (Amine), S (Sugar), St (Steroid), I (Inorganic).

These data revealed some common features between the BIRADS groups, when compared with the control group. 2-hydroxyand 3-hydroxybutyric acids, ethanolamine and fructose were found in higher amount in all pathological groups, while 3-aminoisobutyric acid and cholesterol were less abundant.
Finally, we compared the BI-RADS 5 with the BI-RADS 3 in order to establish the sensitivity of our approach towards differences between different malignity of tumours. The PLS-DA model did not reach statistical significance, being characterized by accuracy= 0.68182, R2=0.55635, Q2=- 0.040531. The removal of an outlier (sample 16) did not improve the model predictivity, while the corresponding OPLS-DA model was characterized by good discriminatory power (Figure 2).

OPLS-DA model obtained from the comparison between BI-RADS 3 and BI-RADS 5. R2X=0.0607, R2Y=0.437, Q2=0.071. Score plot (left) and most important metabolites (features) (right).
One metabolite, the 4-hydroxyproline, resulted as the most important in the discrimination between classes BI-RADS 3 and BI-RADS 5, being more abundant in the BI-RADS 3 group.
Discussion
This study reports a novel investigation approach for the breast tumours analysis and classification. Usually the BI-RADS system has a significant amount of failures in tumours classification especially for the BI-RADS 3 class, a class with borderline characteristics and with difficult categorization of a broad typology of tumours. Table 3 reported the most common benign lesions classified in BI-RADS 3.
Common benign lesions classified in BI-RADS 3.

In case of malignancy suspect after mammography, a biopsy is required. The cells or tissue from biopsy are withdrawn by VAB/Core-Biopsy CB or by Excisional Biopsy of the Breast. These samples will be histologically classified by means of the Anatomic Pathology techniques to verify the presence of tumor.12–13 In all these cases, an upgrade toward malignant lesions diagnosis of 5-7% of cases is expected. For this reason, the surgical extensions procedures after the VAB recently increased (and in some case also by excisional surgery procedures, that is highly discussed in the scientific community), even in case of benign lesions, with a general increase of cost for the surgical procedures and patient's stress and overload. Recently, several guidelines have been published by the most influential scientific associations about the management of lesions with the BI-RADS 3 uncertain malignancy potential, but, to date, there is no univocal positive predictive diagnostic system able to indicate, a priori, which lesions, histologically catalogued as BI-RADS 3, should evolve towards higher risk classes also after extended surgery procedures. Since there is no characteristic radiological pattern, it is not easy to define whether the biopsy collection has wholly removed the lesion or there may be neoplastic or pre-neoplastic alterations accompanying the surrounding parenchyma, so it must be based on the positive predictive value (PPV) of these injuries. Although the latest edition of the NCCN guidelines always recommends surgical excision, many studies do not justify this position. 14 In fact, a European Consensus has recently been published limiting the use of surgery to adequately selected BI-RADS 3 lesions during the multidisciplinary discussion. 15 The need to proceed with surgical excision should be based on clinical-radiological and histological data, taking into particular consideration the patient's family history, 16 after adequate informed consent. This is particularly true, especially after VAB sampling, in cases where microcalcifications are entirely removed by post-biopsy mammography. 12
In order to improve the sensibility and the specificity, we tested the potential contribution of Metabolomics with a pilot study to investigate if GC-MS analysis of plasma samples of patients could give a better understanding of the difference between the categories of benign tumors usually classified in the BI-RADS 3 class.
The comparison between pathological samples (BI-RADS 3 to 5) with controls (all together or each single BI-RADS group) revealed 2- and 3-hydroxybutyric acid, together with ethanolamine and fructose more abundant in pathological samples, while 3- aminoisobutyric acid and cholesterol were found in higher amount in healthy subjects.
2- and 3-hydroxybutyric acid were detected and proposed as diagnostic biomarkers in ovarian cancer patients 17 and in breast cancer. 18 These molecules and their oxidation products are classified as ketone bodies: the higher amount found in cancer patients when compared with healthy controls, may be ascribed to an upregulated fatty acid oxidation due to the higher energy demand of tumour cells.
Aminoisobutyric acid was found in higher amounts in healthy controls: this molecule mainly derives from the breakdown of the DNA pyrimidine base thymine (Figure 3).

Biochemical reaction leading to 3-aminoisobutyric acid.
Ethanolamine constitutes the polar head of phosphoglycerides, being the second most abundant constituent of membranes of this class of lipids. Upregulation in phospholipid metabolism was reported in different studies employing a multiplatform approach.18–19
4-Hydroxyproline resulted the most important metabolite in the discrimination between classes BI- RADS 3 and BI-RADS 5, being more abundant in the BI-RADS 3 group. This molecule is an amino acid found almost exclusively in collagen, being responsible of the correct folding of its helix polypeptide chains. The proline 4-hydroxylation is a post-translational process catalysed by Prolyl 4-hydroxylase (P4H). An increase of 4-hydroxyproline amount has been connected to collagen degradation. 20 Our study proofs the ability of Metabolomics to distinguish between BI-RADS 3 e BIRADS 5 classes, also with limited sample numerosity. Although the statistical models did not reach high predictivity values, nevertheless interesting information were obtained about the metabolic profile of different BI-RADS classes.
Conclusion
In this pilot study, we reported the potential contribution of Metabolomics to the radiological classification of breast cancer images. Metabolomics represents a powerful tool for the extraction of features about the health status of patients with a suspect of breast tumour. As reported in recent publications about metabolomics and cancer,7,8,19,21 a fingerprint with the capability to improve specificity and sensibility of the radiological classification of tumours can be extracted from the peripherical plasma or specific tissue. BI-RADS classification can be enhanced by the quantification of the metabolites list from Metabolomics analysis. We found a particular model for the differences between BI-3 and BI-5 using peripherical plasma. To establish the correct fingerprint and the proper features, the number of samples must be increased, having the reasonable assurance that the metabolomic model can work.
Finally, this study proves that Metabolomic analysis opens an essential gate to the omic data use in the Radiological Diagnosis, the Radiometabolomics.
