Abstract
Background
Neuraminidase (NA) inhibitors (NAIs), including oseltamivir and zanamivir, play an important therapeutic role against influenza infections in immunocompromised patients. In such settings, however, NAI therapy may lead to the emergence of resistance involving mutations within the influenza surface genes. The aim of this study was to investigate the evolution of NA and haemagglutinin (HA) genes of influenza A(H1N1) pdm09 virus in an immunocompromised patient receiving oseltamivir then zanamivir therapies.
Methods
Nasopharyngeal swab (NPS) samples were collected between 27 January 2018 and 11 April 2018 from a haematopoietic stem cell transplant recipient. These include 10 samples collected either pre-therapy, during oseltamivir and zanamivir treatment as well as after therapy. The A(H1N1)pdm09 HA/NA genes were sequenced. The H275Y NA substitution was quantified by droplet digital RT-PCR assay. A(H1N1)pdm09 recombinant viruses containing HA mutations were tested by HA elution experiments to investigate in vitro binding properties.
Results
Oseltamivir rapidly induced the H275Y NA mutation which constituted 98.33% of the viral population after 15 days of oseltamivir treatment. The related HA gene contained S135A and P183S substitutions within the receptor-binding site. After a switch to zanamivir, 275H/Y and 119E/G/D mixed populations were detected. In the last samples, the double H275Y-E119G NA variant dominated with S135A and P183S HA substitutions.
Conclusions
This report confirms that oseltamivir can rapidly induce the emergence of the H275Y substitution in A(H1N1)pdm09 viruses and subsequent switch to zanamivir can lead to additional substitutions at codon E119 resulting in multi-drug resistance. Such data additionally suggest a potential compensatory role for HA substitutions near the receptor binding site.
Introduction
Influenza viruses are responsible for acute respiratory diseases that constitute a significant public health priority worldwide. During typical seasonal epidemics, influenza viruses affect 10% to 20% of the human population [1]. In most people, seasonal influenza strains replicate in the upper respiratory tract causing a self-limited disease. However, in high-risk individuals, such as immunocompromised patients, viral replication may progress towards the lower respiratory tract, potentially triggering secondary bacterial infections and pneumonia and leading to more severe infections and sometimes death [2]. Haematopoietic stem cell transplant (HSCT) recipients are among populations at higher risk of developing seasonal influenza infections with fatal outcomes [3]. Since their appearance in 2009, influenza A(H1N1)pdm09 viruses continue to circulate worldwide and have become important seasonal respiratory pathogens. A recent report showed that viral shedding in HSCT patients with A(H1N1)pdm09 infections was more prolonged compared with infections with other seasonal influenza A subtypes and B strains [4]. The use of antivirals in this situation has the potential to increase the risk of developing drug resistance.
Anti-influenza agents are expected to play a major role against severe influenza infections. Before the recent approval of baloxavir marboxil in 2018 [5], neuraminidase inhibitors (NAIs) constituted the only class of antivirals recommended for the control of influenza infections. Since their development in late 1990s, oseltamivir and, to a lesser extent, zanamivir have been the most frequently used NAIs worldwide [6]. With the advent of the 2009 influenza A(H1N1) pandemic, the use of oseltamivir has grown globally for both prophylactic and therapeutic purposes increasing the potential for the emergence of NAI resistance [7,8]. In immunocompromised patients, A(H1N1)pdm09 variants harbouring the H275Y NA substitution conferring oseltamivir resistance emerged at rates that could exceed 50% [9,10]. As A(H1N1)pdm09-H275Y variants retain susceptibility to zanamivir, the latter constitutes the alternative antiviral option against influenza A(H1N1)pdm09-H275Y variants. However, we and others have recently shown that the switch to zanamivir could induce zanamivir resistance mediated by substitutions at codon 119 (E119A/D/G) within the framework of the NA enzyme [11–14]. Noteworthy, A(H1N1) pdm09 variants with the double H275Y-E119D/G substitution are resistant to all available NAIs [11,12].
Herein, we report a clinical case of multidrug resistance in an HSCT recipient patient who received oseltamivir and zanamivir therapies. By analysing the evolution of surface (HA and NA) genes from sequential respiratory samples, we confirmed the molecular pattern leading to the cross-resistance phenotype, as suggested by previous reports, and highlighted the potential compensatory role of HA mutations within the receptor binding site.
Methods
Molecular detection of influenza A(H1N1)pdm09 virus and antiviral treatment
A total of 10 nasopharyngeal swabs (NPS) were collected from an immunocompromised patient between 27 January 2018 and 11 April 2018 (Figure 1). During that time, the patient received two courses of oseltamivir (from 28 January 2018 to 14 February 2018 and from 27 February 2018 to 1 March 2018) and one course of zanamivir (from 1 March 2018 to 20 March 2018). Plasma samples were also collected at some of these dates. RNA was extracted from clinical samples using the QIAsymphony SP automated instrument and served for a multiplex real-time PCR detection of respiratory pathogens using the 21 Fast Track Diagnostics system [15].

Molecular characterization of HA and NA genes of influenza A(H1N1)pdm09 strains from an immunocompromised patient who received oseltamivir and zanamivir therapies
Sequencing of viral surface genes
The HA and NA genes from NPS samples of A(H1N1) pdm09 viruses were amplified by gene-specific RT-PCR tests and sequenced using the ABI 3730 DNA Analyzer (Applied Biosystems, Carlsbad, CA, USA). The deduced amino acid sequences were compared after an alignment using the ClustalW program (EMBnet, Lausanne, Switzerland).
Viral quantification using droplet digital PCR (RT-ddPCR)
Viral RNA was extracted from thawed specimens using the QIAmp Viral RNA Mini Kit (QIAGEN, Mississauga, ON, Canada). The droplet digital (dd)PCR workflow and data analyses were performed with the OneStep RT-ddPCR supermix according to the manufacturer's instructions. The primers and probes targeting the H275 (WT) and Y275 (mut) variants of influenza A(H1N1)pdm09 virus were previously described [16]. The cycling and post-cycling steps were done as previously reported [17]. The cycled plate was then transferred and read in the FAM and HEX channels of the QX200 reader (Bio-Rad, Mississauga, ON, Canada).
Generation of 7:1 reassortant influenza A(H1N1) pdm09 viruses
The pre-therapy and post-zanamivir HA genes from respiratory swabs were amplified by RT-PCR and cloned into the bidirectional pLLB-A plasmid as previously described [18]. Reverse genetics was used to generate 7:1 reassortant influenza viruses in the background of A/Québec/144147/09 (an A(H1N1)pdm09-like virus; GenBank accession numbers FN434457-FN434464) as previously reported [19].
Haemagglutination and elution assays
In order to assess the impact of HA substitutions present in post-therapy isolates on the binding affinity to α2,3 and α2,6sialic acid receptors, haemagglutination assays were performed in U-bottom microtitre plates using 50 ml of serial twofold dilutions of 7:1 reassortant (generated as described above) and 50 ml of a 1% suspension of guinea pig or chicken red blood cells (RBCs) containing high proportions of α2,6-sialic acid (SA) receptors or α2,3-SA receptors, respectively. Plates were incubated at 4°C for 1 h for determination of haemagglutination titres. For elution experiments, an equivalent of 16 HA units of each isolate were used for haemagglutination assays as stated above and the plates were transferred to 37°C for recording elution times.
Analysis of cell surface expression of recombinant NAs
The potential role of the E287K NA substitution on the surface expression of the NA enzyme was assessed using recombinant NAs. Influenza A/Quebec/144147/09 (H1N1) WT and mutant (H275Y, E287K and H275Y+E287K) NA genes were inserted into a pHDM plasmid (kindly provided by J Bloom, California Institute of Technology, Passadena, CA, USA) in which the NA protein-coding sequence is directly fused to the V5 epitope tag, followed by an internal ribosome entry site (IRES) expressing the green fluorescent protein (GFP). Recombinant pHDM plasmids (1 μg) were used to transfect 293T cells in 12-well plates (4x10 5 cells/well). 20 h post-transfection, the cells were collected after a brief trypsin-EDTA treatment and resuspended in an isotonic buffer for the determination of surface NA expression as previously described [20].
Patient description
A 72-year-old man with acute myeloid leukaemia underwent an allogenic HSCT on 6 December 2017. He was hospitalized on 22 January 2018 for suspected intestinal graft versus host disease (GvHD). 5 days after admission, the patient presented influenza-like symptoms and a real-time RT-PCR test, performed on an NPS, was positive for influenza A(H1N1)pdm09 virus. Double dose oseltamivir treatment (150 mg twice daily, subsequently adapted to the patient's glomerular filtration rate [GFR]) was initiated on 28 January. A chest computerized tomography (CT) scan performed the following day revealed the presence of bilateral ground glass infiltrates. Despite viral positivity on an NPS collected on 12 February (Figure 1), oseltamivir was stopped on 14 February 2018, after 18 days of treatment, due to further decline in renal function. At the respiratory level, the patient's clinical condition was stable at that time. However, oxygen saturation subsequently decreased again, and a chest CT scan performed on 27 February showed an increase of the pulmonary infiltrates. An NPS was collected and oseltamivir was re-introduced on 27 February (the dosage was adapted to a decreased GFR). In the absence of clinical improvement, intravenous (iv) zanamivir was started on 1 March, replacing oseltamivir. While subsequent influenza A RT-PCR cycle threshold (Ct) values initially suggested a decrease in viral load in NPS samples, the Ct values dropped again after 12 March despite continuous iv zanamivir (Figure 1). However, the patient improved clinically, and a chest CT scan performed on 20 March, showed a regression of the infiltrates, and zanamivir was therefore stopped. This exam also revealed an increase in bronchiectasis, indirectly supporting a fibrosing process, possibly due to pulmonary GvHD. On 20 April, the patient experienced a novel episode of respiratory deterioration, and a chest CT scan performed on the same day showed a new increase of the ground glass infiltrates. Zanamivir was temporarily introduced again on 20 April but, due to a rapid clinical improvement upon broad spectrum antibiotic treatment initiation, antiviral treatment was interrupted after 3 days. Thereafter, the patient became dyspneic again and a chest CT scan performed on 3 May displayed an increase in the pulmonary infiltrates. The patient's condition deteriorated, particularly at the respiratory level and he died on 23 May. Autopsy revealed multiple pneumonia foci, of different ages, with active abscess forming pneumonia, organizing pneumonia foci and constituted fibrosis.
Results
Molecular detection of influenza A(H1N1)pdm09 virus in clinical respiratory samples
The real-time PCR for influenza A was positive for the 10 NPS samples collected from 27 January to 11 April with Ct values ranging between 16 and 23 (Figure 1). Plasma samples collected on 25 February 2018 and 26 February 2018 were also positive (Ct of 37 and 38, respectively) whereas 4 other ones (collected on 22 January 2018, 29 January 2018, 14 February 2018 and 5 March 2018) were negative (not shown). Further sequencing experiments identified the influenza A viruses as A(H1N1)pdm09 variants and the sequences of NA and HA genes were determined.
Analysis of the NA genes
The focus was initially on the detection of the H275Y substitution responsible for resistance to oseltamivir. In addition to Sanger sequencing that demonstrated the presence of mixed 275H/275Y chromatogram peaks, dd-RT-PCR tests were performed for further quantifi-cation of 275Y and 275H viral populations. As summarized in Figure 1, the H275Y substitution could be detected as a major population (98.33% versus the WT) in the 12 February 2018 sample (collected after the first course of oseltamivir therapy). After a relative drop to 30.63% in the sample of 27 February 2018 (13 days after cessation of the first oseltamivir treatment), the variant persisted in the remaining post oseltamivir and post-zanamivir samples at relatively important rates (between 64.3% for sample 19 March 2018 to 99.64% for sample 11 April 2018). In addition to the H275Y substitution, the NA protein of samples collected during the first course of zanamivir therapy contained mixed residues at positions 287 (E/K), 94 (V/I), 95 (S/G) and 119 (E/G/D). Finally, the NA of the last three samples collected in the absence of therapy harboured an H275Y/E119G variant (Figure 1).
Analysis of the HA genes
The receptor binding site (RBS) of influenza viruses consists of three structural elements including two loops (130-loop and 220-loop) and one helix (190-helix) [21]. The HA gene of clinical samples contained three substitutions (V19L, S135A and P183S) that appeared in NPS collected after the first oseltamivir course and were subsequently maintained. Changes at residues 250 (V/A) and 265 (V/I) could also be transiently detected. The alignment of HA amino acid sequences (Additional file 1) showed that the S135A and P183S substitutions were located near the RBS of the HA molecule. Residue 135 is part of the 130-loop whereas residue 183 is near the 190-helix.
In haemagglutination assays using A(H1N1)pdm09 7:1 reassortants, those containing the post-treatment HA (19L, 135A and 183S) eluted more rapidly guinea pig RBCs than the reassortants containing the pre-therapy HA (19V, 135S and 183P; 1 h versus >4 h; Table 1). The two reassortants had the same elution time (1 h) when using chicken RBCs.
Receptor binding specificities of A(H1N1) viruses using HA and HA elution assays
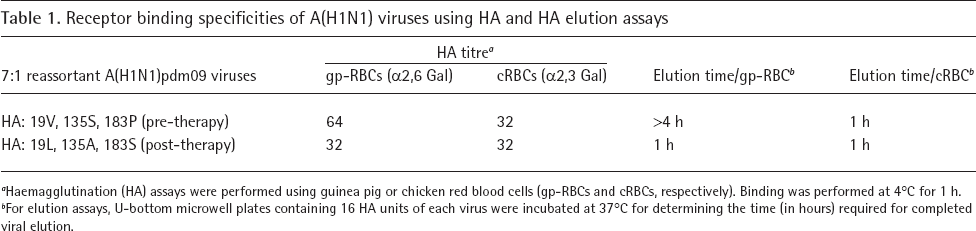

Haemagglutination (HA) assays were performed using guinea pig or chicken red blood cells (gp-RBCs and cRBCs, respectively). Binding was performed at 4°C for 1 h.
For elution assays, U-bottom microwell plates containing 16 HA units of each virus were incubated at 37°C for determining the time (in hours) required for completed viral elution.
We further tried to investigate whether these HA changes could improve the fitness of NAI-resistant variants. Unfortunately, despite many attempts, no recombinants could be rescued when using the NA gene containing the E119G/H275Y substitutions. Noteworthy, in previous clinical cases, A(H1N1)pdm09 isolates containing E119G [12] and E119D [11] NA substitutions demonstrated a deficient growth in vitro.
Surface NA activity
The impact of E287K and H275Y substitutions on NA surface activity when assessed in 293T cells transfected with pHDM plasmids confirmed the role of H275Y in reducing the surface NA activity (54% versus the WT) and revealed that the E287K change alone did not significantly alter the NA surface activity (89% versus the WT; Figure 2). In addition, the H275Y+E287K mutant showed 32.5% of surface activity versus the WT demonstrating that E287K did not improve the NA surface activity of the H275Y mutant protein.
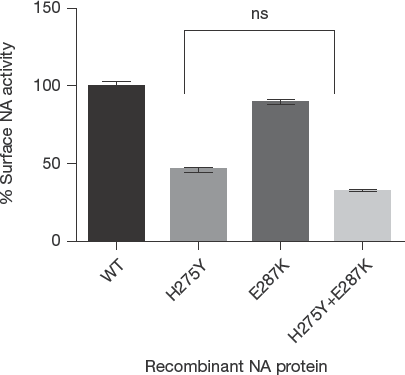
Cell surface activity of recombinant A(H1N1)pdm09 NAs
Discussion
Reduced inhibition to NAIs in influenza viruses is mainly mediated by changes (substitution/deletions) in or near the active site of the NA enzyme [22]. Although NAIs are analogues of the sialic acid (DANA: 2,3-dehy-dro-2-deoxy-N-acetylneuraminic acid), zanamivir and oseltamivir can be distinguished by important structural differences that may influence the nature and rate of mutations of resistance [23]. Zanamivir, which is structurally closer to DANA compared with oseltamivir [24], has been infrequently associated with the emergence of viral resistance [6]. Accordingly, in vitro passaging of influenza A(H1N1)pdm09 virus under zanamivir pressure failed to induce drug resistance [25]. In contrast, in A(H1N1)pdm09 infections, prophylactic or therapeutic exposures to oseltamivir readily selected for NAI-resistant H275Y variants in immunocompromised patients [7,12]. Besides NA changes, reduced inhibition to NAIs could also involve changes in or near the receptor binding site (RBS) of the HA protein due to the important functional balance between HA and NA surface proteins [26].
In this, report, we took advantage of having a series of respiratory samples from an HSCT patient including pre-therapy NPS and samples collected after oseltamivir (used as a first-line antiviral agent) and zanamivir (administered after detection and persistence of the H275Y substitution) for analysing the evolution of influenza A(H1N1)pdm09 NA and HA genes. Similarly to previous reports, we found that oseltamivir rapidly selected for the H275Y NA mutation (Figure 1). In fact, a very low proportion of that variant (0.06%) could be detected by ddRT-PCR in the first NPS collected before oseltamivir therapy. Oseltamivir pressure enhanced the H275Y level to 98.3% in the subsequent NPS. As previously described [11–13], the switch to zanamivir also induced the occurrence of a second NA substitution at the E119 framework residue (E119G). This substitution was maintained during 1 month in the double H275Y-E119G variant (Figure 1). Tamura et al. [12] reported that the dual H275Y-E119G mutation recovered from an immunocompromised child resulted in ≥100-fold reduced inhibition against zanamivir, oseltamivir, peramivir and laninamivir. This was also the case for another H275Y-E119D A(H1N1) pdm09 variant described by us [11]. Of interest, viral cultures could not be obtained for the two double mutants suggesting attenuating effects [11,12]. The role of the H275Y-E119D changes in altering viral fitness was further confirmed both in vitro and in animal models [27].
While the impact of the framework H275Y and E119G substitutions on the resistance phenotype to NAI is well documented [12], the effect of NA substitutions at residues 94, 95 and 287 has not been reported. Of interest, residue 287 is a surface exposed amino acid that was suggested to play a role on virion release impacting in vitro virus infectivity and surface NA activity [28]. Thus, there seems to be a rationale for investigating the possible compensatory role for E287K. Our results showed that the introduction of the E287K mutation in the NA gene containing the oseltamivir-resistance H275Y substitution did not improve surface NA activity when using recombinant proteins (Figure 2). Noteworthy, none of the three changes at positions 94, 95 and 287 was retained in the last multidrug-resistant samples.
As shown in Figure 1, besides NA changes, the clinical A(H1N1)pdm09 variants contained HA substitutions under NAI pressure. As S135A and P183S changes are located in or near the HA RBS, there was a possibility for altered HA binding property. For instance, a recent report showed that in vitro passaging of A/California/04/2009 (H1N1) virus under lectin pressure induced the S183P HA substitution, which significantly increased binding to α2,6 Sa-linked glycans [29]. In a recent clinical report, NAI therapy against A(H1N1)pdm09 infection in a child with severe combined immunodeficiency disease (SCID) induced the P183S HA change [14]. In our HA elution experiments, the reassortant containing the pre-therapy HA (135S, 183P) had a higher affinity to the guinea pig RBC (rich in α2,6 Sa-linkage) from which it could not elute after 4 h of incubation at 37°C, contrasting with the complete elution within 1 h for the reassortant containing the post-therapy (183S) HA (Table 1). Of note, while most influenza H1N1 strains of 2018 harboured HA with 183P residue, the HA of previous 2009 and 2015 vaccine strains (A/California/07/2009 and A/Michigan/45/2015, respectively) had S at this position. Therefore, the NAI pressure seemed to cause a reversion to the original A(H1N1)pdm09 genotype. Thus, due to the reduced NA activity caused by the H275Y substitution, the isolate had to select for HA substitutions with the potential to reduce affinity to α2,6 receptors for restoring the HA/NA balance, as demonstrated in our HA elution experiments. Unfortunately, the fact that we do not possess the clinical isolates with the different HA/NA genotypes to compare their fitness constitutes a limitation of this study. Moreover, we could not rescue recombinant viruses harbouring the post-therapy NA protein (H275Y/E119G) despite many attempts. Finally, the potential compensatory role of changes in other viral segments is also to be considered.
In conclusion, we provide herein an additional clinical report on the emergence of multidrug-resistant A(H1N1)pdm09 viruses in immunocompromised patients receiving sequential oseltamivir and zanamivir therapies. In conjunction with previous reports, there seems to be a consistent molecular pathway leading to the multi-drug resistance phenotype (Additional file 2). It consists on the rapid selection of the H275Y NA substitution, following oseltamivir therapy, followed by E119G/D changes while on zanamivir therapy. The resulting double NA mutant displays highly reduced susceptibility to all NAIs. Contrasting to previous reports [11,12], we provide here insights for possible involvement of HA changes in or near the RBS on the drug resistance phenotype. Due to limited antiviral options for the control of influenza infections in immunocompromised individuals, the occurrence of such molecular events may have major clinical implications. This report highlights the need for the development of novel anti-influenza compounds.
Footnotes
Acknowledgements
This work was supported by a Canadian Institutes of Health Research (CIHR) foundation grant to GB (grant No. 229733) for a research programme on the pathogenesis, treatment and prevention of respiratory and herpes viruses.
The authors declare no competing interests.
