Abstract
Background
HBV and HIV infections are highly endemic in sub-Saharan Africa and Nigeria while HBV–HIV coinfection is not uncommon. Antiretroviral (ART)-treatment for HIV can affect HBV whereby antiviral resistance mutations in the HBV genome can be selected. Here, we determined the prevalence of resistance mutations among ART-experienced and ART-naive HIV–HBV-coinfected patients in southwestern Nigeria.
Methods
A total of 81 serum samples from HBV–HIV-coinfected patients who were either ART-naive or received lamivudine (3TC)-containing ART-therapy and HBV-monoinfected patients were analysed. Hepatitis B surface antigen (HBsAg) was detected using ELISA. HBV-positive samples were confirmed by PCR amplification of the surface and polymerase regions. Mutations conferring drug resistance to HBV were analysed by direct sequencing. Phylogenetic analysis was performed to identify the HBV genotype.
Results
Of the 81 HBsAg-positive samples, 27 had detectable HBV DNA by real-time PCR with mean viral loads of 6.77 log IU/ml. Phylogenetic analyses showed a predominance of HBV genotype E. A high prevalence (22.2%; 6/27) of HBV resistance mutations among ART-experienced HBV–HIV-coinfected patients was detected. However, a relatively high selection rate of resistance mutations in drug-naive HIV–HBV-coinfected (3.7%; 1/27) and in HBV-monoinfected patients, potential drug resistance mutations (7.4%; 2/27) were also observed. HBV polymerase amino acid substitutions found included rtV173L, rtL180M, rtM204V, rtK212R, rtS213T, rtV214A, rtL229V and rtP237A/S.
Conclusions
Drug resistant mutations were detected frequently in ART-experienced HIV–HBV patients. Well-coordinated antiviral therapy for HIV patients coinfected with HBV should include proper HBV diagnosis and resistance testing to minimize the emergence and spread of antiviral drug resistance.
Introduction
HBV is a common cause of viral hepatitis and liver cancer in humans worldwide [1]. With the availability of an effective vaccine, the number of new HBV infections has markedly reduced globally over the years; however, about 350 million people remain chronically infected with this virus [2], while there are 4.5 million new HBV infections globally per annum, of which approximately a quarter progress to liver disease [3,4]. HBV is a para-retrovirus, which replicates through an RNA intermediate using the viral polymerase (reverse transcriptase [RT]) similar to that of HIV. The HBV reverse transcriptase is error prone and therefore leads to high heterogeneity of the virus. HBV is currently classified into ten genotypes (A–J) with varying geographical distribution. Genotype E is the predominant strain in west Africa (including Nigeria) and central Africa [5].
Most antiviral drugs currently licensed for HBV treatment target the HBV viral reverse transcriptase, an enzyme with multiple functions in viral replication and infection [6,7]. Inadequate suppression of HBV replication by antivirals under drug pressure and the prolonged usage of these drugs may result in the development of mutations within the polymerase region of the HBV genome that confers drug resistance or the ability to escape immune detection [8]. Therefore, a well-coordinated antiviral therapy should include proper diagnosis of HBV and resistance testing to minimize the emergence as well as the transmission of drug resistance mutations [9].
Hepatitis B is hyper-endemic in Nigeria with seroprevalence between 10–40% [3]. The HBV vaccine has been included in the national immunization programme in Nigeria since 1995 but was not widely available until 2004. Despite this, there is still a high burden of HBV infection in Nigeria [10]. With the 3.2% prevalence of HIV in Nigerian adults [11], HIV–HBV coinfections are not uncommon. The World Health Organization (WHO) has recommended that all HIV-infected patients should be tested for the presence of HBV infection [12], but this is rarely done in Nigeria and other regions of sub-Saharan Africa due to high cost or partly due to neglect of the disease and concentration on HIV treatment [13,14]. Since the HBV status of most HIV-infected patients remains unknown, and first-line anti-retroviral therapy regimen usually contains zidovudine and lamivudine (3TC) in Nigeria [15], this increases the risk of the emergence of HBV drug resistant strains. Moreover, studies have shown that HBV mutations can occur even in the absence of 3TC therapy in drug-naive HBV carriers, thereby contributing to the rapid development of drug resistance [16].
The effect of administration of ART-therapy in HIV–HBV-coinfected patients on HBV resistance in Nigeria is not well documented. In this study, we analysed the prevalence and pattern of HBV drug resistance mutations among HIV-positive hepatitis B surface antigen (HBsAg)-positive patients in Southwestern Nigeria.
Methods
Study samples and design
We carried out a retrospective study on samples collected by convenient sampling from patients at the University Teaching Hospital, Ibadan (UCH) and Obafemi Awolowo Teaching Hospital Ile-Ife (OAUTH), both located in southwest Nigeria, between 2016 and 2017. 81 HBsAg-positive patient samples were analysed in this study including 18 HIV-positive patients (13 ART-experienced and 5 ART-naive), and 63 HBsAg-positive infected patients. Serum samples were collected after informed consent and stored at −20°C. All the ART-experienced HIV-infected patients recruited in this study had 3TC as part of their regimen. All samples were tested negative for HCV, HDV and HEV.
Serology
Samples were initially tested for HBsAg using rapid immune-chromatographic strips (Micropoint Bioscience, Santa Clara, CA, USA). HCV, HDV and HEV antibodies were excluded by ELISA (Bio-Rad, Hercules, CA, USA; DiaSorin, Saluggia, Italy; Wantai Hepatitis E virus Diagnostics, Beijing, China) before selection. HBsAg was re-tested using Enzyme Immunoassay (Wantai Hepatitis B virus Diagnostics, Beijing, China) following the manufacturer's instructions.
PCR Amplification
Viral nucleic acid was extracted from 140 μl of each serum sample using the QIAamp Viral DNA Mini Spin kits with the Qiacube BioRobot Workstation (QIAgen, Hilden, Germany). Real-time PCR for HBV DNA was carried out using virus-specific primers and Taqman probes with the LightCycler 480 Instrument (Roche Diagnostics Corporation, Rotkreuz, Switzerland) as described previously [17,18]. Nested PCR amplification of the HBV RT gene (1145bp) or a partial coding sequence of the S-gene (332bp) followed by electrophoresis of the PCR products in 1.5% agarose gel stained with GelRed in 1% Tris-acetate-EDTA was performed as described previously [18].
Phylogenetic and mutation analyses
HBV genotyping was performed by sequencing of the PCR products followed by phylogenetic analysis as described [17,18].
Sequences from the RT-gene were analysed for drug resistant mutations using BioEdit version 7.2.5 and geno2pheno software. All primers used have been previously described [17,18].
In brief, Sanger sequencing was performed using the second-round PCR products as described previously [17,18]. Virus-specific primers [17] were used for sequencing of the PCR products using BigDye version 3.1 in an automated DNA sequencer ABI 3130xl (Applied Biosystems, Foster City, CA, USA). Sequencing PCR conditions consisted of a cycle of 96°C for 1 min, 25 cycles of (96°C for 30 s, 45°C for 15 s) and 60°C for 4 min. Sequence electropherograms were edited and analysed using Geneious R10.0.5 software. Nucleotide sequences were aligned with published reference sequences from GenBank using Geneious R10.0.5. The generated FASTA sequences were used for phylogeny. Mutation analysis was done using BioEdit version 7.2.5 and geno2pheno software. Sequences of isolates from this study have been submitted to NCBI under the accession numbers MH918641 to MH918655.
Statistical analysis
HBV viral copies are expressed in international log units per ml of serum (log IU/ml). Patients’ data are presented in averages or percentages. Sample data were compared between the groups using the Mann–Whitney U test and it was used to compare numerical variables with 95% CI, P-values less than 0.05 were considered statistically significant. Statistical data were analysed using Graphpad Prism. Phylogenetic trees were constructed using the maximum likelihood algorithm implemented in MEGA version 7.0 software. The evolutionary history was inferred by using the Maximum Likelihood method based on the Tamura-Nei model.
Ethical consideration
The study was approved by the Ethical review board of LAUTECH College of Health Sciences, Osogbo, Osun State, Nigeria and informed consent was received from every participant before sample collection. For this study, participants’ identities remain anonymous.
Results
Patient characteristics
Of the 81 participants in this study, the ratio of males to females was 36/45 with overall mean age of 34.1 years ±11.3 (range: 5–77 years). All samples were tested positive for HBsAg using enzyme immunoassay, and 27 (33.3%) of these had detectable HBV DNA with mean serum viral load of 6.77 log IU/ml. Table 1 shows characteristics of the patients including liver function data. Of those which had detectable HBV DNA, 16 were males and 11 were females with mean age 31.0 years ±11.1 (range: 5–52 years); 9 (33.3%) were HIV-positive patients and 18 (66.7%) asymptomatic HBV-monoinfected individuals. Seven of the nine HIV–HBV-coinfected patients were ART-experienced while two were naive.
Characteristics of study participants

ALT, alanine aminotransferase; AST, aspartate transaminase; HBeAg, hepatitis B e antigen; NA, not applicable; ND, not determined.
HBV Genotyping and Mutation Analyses
HBV genotyping was done by sequencing followed by phylogenetic analyses. Only genotype E was observed in the study cohort (Figure 1). The overall prevalence of resistance mutations was 25.9% (7/27) with respect to HBV-DNA-positive patients; with the highest rate detected among ART-experienced HIV–HBV-coinfected patients. Six (22.2%) ART-experienced HIV–HBV-coinfected patients showed HBV resistance. Notably, one (3.7%) ART-naive HIV–HBV-coinfected patient also showed HBV resistance. Out of the HBV-monoinfected patients, two (7.4%) showed potential HBV resistance mutations (Table 2). Seven patients showed resistance to lamivudine (3TC), all of which were HIV–HBV-coinfected; six ART-experienced and one ART-naive. The ratio of male to female among the patients with drug resistant mutations and/or potential resistance mutations is 2:1 with mean age 24.7 ±12.8 years (range: 5–40 years) and the mean HBV viral load was 5.67 log IU/ml of serum while a mean HBV viral load of 6.9 log IU/ml of serum was observed among patients without HBV resistance mutations (t=1.0159; P=0.31). There is no significant difference between the mean viral loads between the two groups.

Reconstructed phylogenetic tree of the polymerase region of HBV isolates
Eight amino acid substitution sites were identified with four (rtV173, rtL180, rtM204 and V214) in major resistance-associated positions and the remaining four (S213, K212, P237 and L229) at positions previously reported as potential or relevant mutation sites [19,20]. Two unique amino acid substitutions were found at position P237 (P237S/A), a previously identified putative adefovir associated resistance mutation site [20]. Multiple amino acid substitutions found include L180M+M204V, V173L+L180M+M204V, L180M+M204V+P237S, L180M+M204V+L229V and V214A+K212R+P237A. One single resistance-associated mutation detected was S213T (Table 3 and Figure 2).

Analysis of HBV resistance mutation
Characteristics of patients with HBV resistance mutations
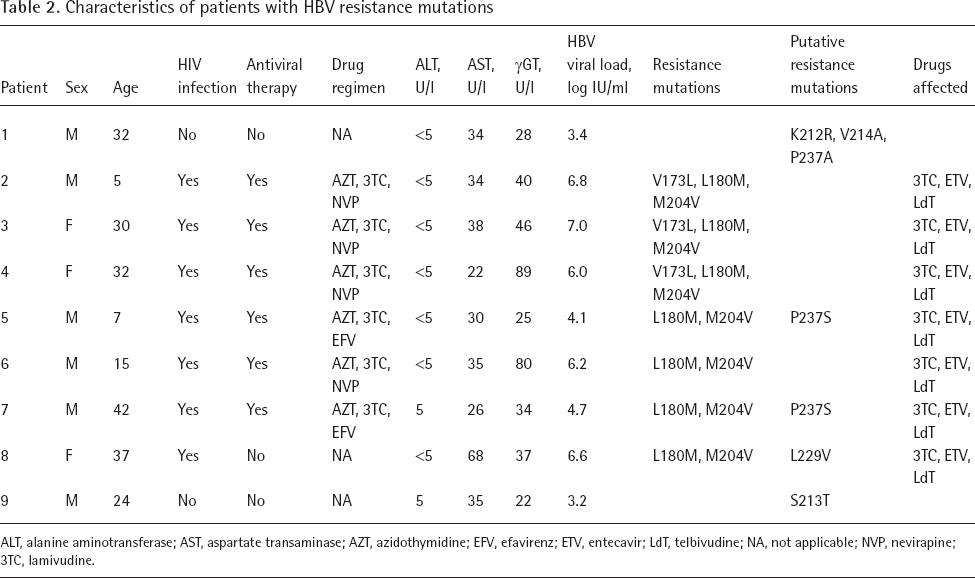
ALT, alanine aminotransferase; AST, aspartate transaminase; AZT, azidothymidine; EFV, efavirenz; ETV, entecavir; LdT, telbivudine; NA, not applicable; NVP, nevirapine; 3TC, lamivudine.
Mutations in the RT domain of the P gene in HIV–HBV-coinfected and HBV-monoinfected patients

Discussion
In this study of HBsAg-positive patients with or without HIV coinfection in southwestern Nigeria, the prevalence of HBV drug resistance mutations is high with 25.9% (7/27). The highest rates were found among HIV–HBV-coinfected ART-experienced patients (22.2%), all of them having lamivudine (3TC) as part of their ART-regimen. However, we also found HBV resistance among ART-naive HIV–HBV-coinfected patients (3.7%) as well as some potential drug resistance mutations in drug-naive HBV-monoinfected patients (7.4%). The most common amino acid substitutions detected were the well described 3TC resistance mutations rtL180M and rtM204V, found in seven (25.9%) HBV–HIV-coinfected patients. Primary drug resistance and secondary mutations were found in this study mainly among ART-experienced HIV–HBV-coinfected patients who also showed multiple HBV drug resistance. In total, seven patients had resistance to lamivudine which were all HIV–HBV-coinfected patients; six ART-experienced and one ART-naive. Similar results have previously been reported and WHO has recommended HBV and HCV testing before initiation of ART in HIV patients [12,21]. All participants had been infected with HBV genotype E. Our results showed that viral mutations related to lamivudine, adefovir, tenofovir, telbivudine and entecavir resistance circulate in patients and even in some who have never been treated.
HBV is highly heterogenic owing to lack of proofreading activity of the RT polymerase which leads to the production of numerous quasispecies within an infected host as well as wide inter-host variations. The need for prolonged use of polymerase inhibitors also mounts selective pressure on virus population, contributing to the development of drug resistance which may eventually lead to therapy failure in patients [22]. We identified eight amino acid substitution sites all of which have been previously reported [19,20]. Primary amino acid substitutions M204V were detected in seven HIV–HBV-coinfected patients, all of which are ART-experienced with lamivudine as part of their ART regimen; except one patient who is yet to begin therapy. This may be possibly due to initial infection with a mutated HBV strain. In agreement with previous studies [9,19], resistance to lamivudine was most frequently detectable in this study.
Resistance to tenofovir has been rarely reported and mostly in chronic HBV patients or HIV-coinfected patients undergoing drug therapy [22]. However, in this study we found in a drug-naive HBV-infected non-HIV patient, mutations that have been previously shown to be associated with resistance to adefovir and tenofovir [23]. We further found V214A and S213T substitutions in drug-naive HBV-monoinfected patients as well as the amino acid (AA) substitution K212R. This finding is in agreement with a recent report of a study conducted in Ethiopia where V214A and S213T amino acid substitutions were also found in drug-naive HBV-monoinfected patients [13]. It has been reported that the AA substitutions at V214 and S213 may be implicated in adefovir and tenofovir failure [19,23]. Amino acid change at K212 has been rarely reported and association with resistance is not fully described [18,21]. These drug-naive patients may have been infected from a patient who had developed resistance after antiviral treatment or were naturally occurring resistance mutations [13]; however, with the circulation of these resistant strains, the treatment options available will even be more limited.
HBV RT residues at position 236, 237 and 238 are known adefovir resistance sites known to induce resistance alone or in conjunction with others and are also known to vary among the genotypes of HBV [23,24]. In this study we identified two new substitutions at position P237 (A/S), unlike the P237H/T reported previously [13,19,21,24,25]. Two of the patients with this HBV P237 mutant were ART-experienced HIV– HBV-coinfected patients, while one was a drug-naive HBV-monoinfected individual. Amino acid substitution L229V was also found in one HIV patient in this study. This substitution has been described as a compensatory mutation for the impaired replication as result of AA substitution M204I [26].
In this study, most 3TC-related mutations were found in ART-experienced HIV–HBV-coinfected patients, however, the high prevalence of HBV drug resistance mutations detected in this study among HIV patients recommend the need for drug resistance testing before the initiation of therapy for HIV-infected individuals and chronic HBV patients. A well-coordinated antiviral therapy should therefore include proper diagnosis and resistance testing in order to further minimize the emergence and spread of drug resistance.
Comparison of the mean viral loads between patients with resistance-associated mutations and those without mutations showed no significant difference (P=0.31), this agrees with a previous study where it was also reported that there was no significant association between the viral loads of patients with or without HBV resistant mutations [8]. Marked increase in viral load has been a useful tool for monitoring antiviral therapy [27], however, this study is limited in that the sample size is low and there was no room for long-term monitoring of patients. In addition, the serum levels of transaminases between HIV–HBV-coinfected and HBV-monoinfected patients did not differ significantly. This is probably due to the fact that the liver enzyme levels may fluctuate in chronically infected patients and a one-time point analysis may not be sufficient to demonstrate clinical relevance; however, participants in this study were not followed up.
Limitations to this study are the relatively small number of patient samples and that the study was regionally limited to south-west Nigeria. Therefore, results may not be generalizable for the whole of Nigeria. ART-treatment in Nigeria includes mainly lamivudine (3TC) due to cost effectiveness; consequently the results of the frequency of resistance mutations shown here cannot be transferred to HIV patients under different treatment regimens. Due to the observational nature of this analysis the relationship between predictor factors and occurrence of HBV resistance mutations could not be assessed. It was also not possible to determine the ART adherence; however, suppression of HIV RNA levels and of HBV replication and the frequent selection of 3TC as well as other resistant mutations to drugs not used in this study group suggest other factors contributing to 3TC resistance. Thus, larger future studies are necessary to understand molecular mechanisms leading to therapy failure and selection of resistance mutations in HIV–HBV-coinfected individuals.
In summary, this study shows the frequent selection of HBV resistance mutations in ART-experienced HIV–HBV patients that strengthen the necessity for HBV testing before initiation of ART as recommended by WHO. The findings in this study also raise the concern about the circulation of drug resistant HBV strains among drug-naive HIV and non-HIV population. Therefore, a well-coordinated therapy and care for HIV patients should include proper HBV diagnosis and resistance testing to minimize the emergence and spread of antiviral drug resistance.
Footnotes
Acknowledgements
We thank the medical staff of the Gastroenterology Department UCH and HIV Clinic OAUTH for their assistance during collection of the samples. We are thankful to the laboratory staff of Robert Koch Institute, Berlin, Germany, namely Hannes Jarmuth, Steffen Zander, Roswitha Lorenz, Ute Obst and Kathrin Stanossek for technical support.
The study was supported by funding of DAAD (German Academic Exchange Service) as part of a bi-national scholarship to OAA and by a grant from the German Federal Ministry of Health with regard to a decision of the German Bundestag by the Federal Government (grant: ZMVI-2519GHP711, NiCaDe-Project). DH is supported by a scholarship from the Claussen-Simon-Stiftung (Claussen-Simon Foundation) ‘Dissertation Plus’ program, Germany. BW is supported by the China Scholarship Council (CSC), Beijing, China. The content is the responsibility of the authors only and does not represent the views of DAAD, BMG, Claussen-Simon-Stiftung and CSC.
OAA, OOO and CTB initiated, participated in conception and designed the study. OAA, ASO and ECO collected samples. OAA performed the experiments. OAA, BW, DH, FAO, OA and KK analysed and interpreted the data. OAA, OOO and CTB prepared the manuscript. All authors revised and approved the manuscript.
The authors declare no competing interests.
