Abstract
Background
Direct-acting antiviral (DAA) therapy has dramatically increased sustained virological response rates in HCV-infected patients. However, resistance-associated substitutions (RAS) interfering with NS3- and NS5A-targeted therapy, still emerge. This real-life study analysed the type and frequency of RAS in rare cases of patients failing DAA regimens in 12 clinical centres in Israel.
Methods
Blood samples and clinical data from 49 patients who failed various DAAs were collected. RAS identified in the NS3 and NS5A regions by population (Sanger) and next-generation sequencing (NGS) were compared by treatment regimen and HCV subtypes.
Results
The majority (71.4%, 35/49) of patients were infected with the genotype (GT)1b strain, while 12.2% (6/49) carried GT1a and 14.3% GT3a/b (7), GT4a (1) and GT1b/GT3a. RAS were identified in 85.7% (42/49) of failures, of which 90.5% (38/42) were clinically relevant RAS (known to be associated with a specific GT and DAA in patients failing therapy or those with more than twofold change in in vitro replicon assays). The most abundant RAS were 168A/E/Q/G/N/V (32.6%, 16/49) identified in NS3, and 93H/N (61.2%, 30/49), 31I/M/V (34.7%, 17/49) and 30R/H/K (12.2%, 6/49), identified in NS5A. Significantly more clinically relevant RAS were identified in NS5A (82.2%, 37/45) than in NS3 (35.7%, 10/28; P<0.01). While RAS were identified in all GT1a, GT3b and GT4a failures (100%, 10/10), only 71.8% (28/39) of GT1b or GT3a failures had RAS (P=0.09). In four cases, NGS identified additional clinically relevant RAS and in one patient, NGS deciphered coexistence of GT3a and GT1b infections.
Conclusions
Our findings, together with additional real-life data, will contribute to the optimization of retreatment in DAA failure, when cost-related and suboptimal regimens must be employed.
Introduction
The recent development of new therapies targeting specific viral proteins has revolutionized the fate of HCV-infected patients. These advances have resulted in successful outcomes and real hopes for global HCV eradication. However, <5% of patients treated with direct-acting antivirals (DAA), fail therapy. These include difficult to cure HCV cases, such as those infected with HCV genotype (GT)3, who suffer from liver cirrhosis, are coinfected with hepatitis B or HIV, or who have already failed targeted therapy [1,2]. Ideally, these patients could be treated with the new recently approved pan-genotypic DAAs, which target the replication complex NS5A protein (pibrentasvir and velpatasvir), the NS3/4A protease (glecaprevir and voxilaprevir) and the NS5B polymerase (sofosbuvir) and could cure even such hard-to-treat patients [3]. Nevertheless, until these new regimens are accessible worldwide, retreatment decisions need to be carefully planned to ensure successful outcomes. Correct determination of HCV genotype and subtype and identification of resistance-associated substitutions (RAS), remain critical factors.
NS3, and later on, NS5A and NS5B resistance testing, was proposed upon approval of the first-generation DAAs, but studies have shown that baseline RAS do not directly predict treatment failure [4,5]. On the other hand, preliminary data suggest that in patients that fail DAA therapy, retreatment can be optimized based on NS5A RAS testing results [6]. NS3 RAS, for example, R155K in HCV GT1a and D168E/V in GT1b, which were shown to emerge after failure on a simeprevir- or a paritaprevir-based treatment [7], may also be relevant when protease inhibitors are considered for retreatment. NS5B RAS are rarely detected (1% of failures) even after failure of a regimen containing a polymerase inhibitor [8,9] and therefore, NS5B resistance testing is not considered even before retreatment.
RAS are usually determined by population sequencing (Sanger sequencing), or by next-generation sequencing (NGS). While population sequencing detects variants present in at least 15% of the quasispecies, NGS can detect variants with a prevalence of >1.5% of the viral quasispecies [10]. NGS may be beneficial especially in patients who failed DAA therapy and are about to start a new treatment regimen when the majority of the viral population already constitutes wild-type variants [11] although up to date, no study has shown the clinical benefit of variants with prevalence smaller than the Sanger-like cutoff of 15% as recommended by EASL guidelines [12]. Different RAS emerge following exposure to different DAAs. Moreover, different amino acid substitutions emerge in the background of different HCV genotypes and subtypes [13]. Accordingly, RAS are only considered clinically relevant if they were previously found to be associated with a specific DAA in patients failing therapy or if they conferred a greater than twofold change in susceptibility compared with a reference strain in in vitro replicon assays [7]. Thus, amino acid substitutions identified at failure that, according to the current literature, cannot be ascribed to the treatment regimen at failure, are herein designated non-clinically-relevant RAS. Although such RAS cannot account for the treatment failure, they should be taken into account before retreatment, if drugs directed at such RAS are to be considered.
To date, real-life data on RAS in DAA treatment failures is still accumulating [7,14,15]. In this study, we characterized the type and frequency of NS3 and NS5A RAS identified in Israel using both population sequencing and NGS in plasma samples from patients who failed multiclass DAA treatments.
Methods
Study design and participants
Plasma samples (n=49) and clinical data from HCV patients who failed various DAA regimens, were collected between 2015 and 2018 from 12 medical centres in Israel. Plasma samples were collected <1 year post failing regimen (or longer, due to technical difficulties) and stored at −70°C prior to analysis. The study was approved by the Ethical Committee of the Sheba Medical Center (approval number 9329-12-SMC). Fisher's exact test was used for statistical analysis.
Amplification of NS3 and NS5A regions
RNA was extracted from 0.5 ml plasma, using the NucliSENS Easy MAG total RNA extraction system (Biomerieux, Marcy l'Etoile, France), according to the manufacturers’ protocol. Amplification of the NS3 region was performed using high fidelity HiFi (Thermo Fisher Scientific, Waltham, MA, USA) for Sanger sequencing, with 7 ml HCV RNA, as previously described [4], or using the DeepChek®-HCV NS3 DR assay when NGS was applied (ABL, Luxembourg City, Luxembourg). Amplification of the NS5A region was performed using the DeepChek®-HCV NS5A DR Assay (ABL).
Sanger sequencing
Direct sequencing of all PCR products was performed using an automatic sequencer (ABI PRISM 3100 genetic analyzer DNA Sequencer; Applied Biosystems, Foster City, CA, USA) and BigDye Terminator v1.1 Cycle Sequencing kit (Applied Biosystems). Nucleotide sequences from all genotypes were assembled using the The OpenGene® DNA Sequencing System (Siemens Healthineers, Erlangen, Germany). All sequences (GT1–4) were aligned with reference sequences (GeneBank accession numbers NC_004102 or M62321 for HCV genotype GT1a, D90208.1 for genotype GT1b, D17763 for GT3a, D49374 for GT3b and Y11604 for GT4).
NGS Analysis
NS3 and NS5A PCR products used for Sanger sequencing were also used for NGS analysis. Following removal of primers and free nucleotides and size-selection with Agencourt AMPure XP (Beckman Coulter, Inc., Brea, CA, USA), the concentration of the PCR products was determined with Qubit 2.0 and the dsDNA HS kit (Life Technologies, Carlsbad, CA, USA). NS3 and NS5A fragments purified from each original sample were pooled together in an equimolar ratio and diluted to a final concentration of 0.2 ng/ml. Library preparation was conducted with the Nextera XT kit (Illumina, San Diego, CA, USA), according to the manufacturer's instructions. Briefly, 1 ng pooled PCR products (5 ml of 0.2 ng/ml) was fragmented, index adapters were ligated and libraries were amplified. Following size-selection and primer removal (with AMPure XP beads), size distribution was assessed using Tape Station 2.0 (Agilent, Santa Clara, CA, USA) and normalized libraries were pooled in a single tube. Prior to sequencing, the pooled libraries were denatured and diluted to 4 nM. Sequencing was performed using a 500 cycle (2 X 250bp paired -end) Miseq reagent kit v2 and Miseq sequencer (Illumina). Miseq sequencer was programmed to measure Phred quality score (Q score) and filter out reads with Phred score lower than 30 [16]. Hence, the generated FASTQ files contain only ≥Q30 reads (base call accuracy of 99.9%). A total of 5,000–100,000 reads per sample was obtained as detailed in Additional file 1.
Analysis of sequencing results
DeepChek-HCV software (a commercial software for resistance analysis of NGS and Sanger results from ABL) was used for interpretation of all sequencing (NGS and Sanger) results [16,17].
RAS were identified by DeepChek and the clinically relevant RAS were determined based on their association with a specific genotype and DAA in patients failing therapy or with twofold change in in vitro replicon assays as described by Sorbo et al. [14]. RAS at amino acids positions 28, 30, 31, 58 and 93 for NS5A, and 36, 54, 55, 80, 122, 155, 168, 170 and 174 for NS3 were assessed. A patient was considered as having a clinically relevant RAS if Sanger or NGS analysis identified such a RAS. Threshold value for mutation frequency in NGS was 1.5% [10]. HCV genotype was determined by Geno2pheno [18].
Results
Characterization of patients failing various DAA regimens
During the study period (between September 2015 and February 2018), approximately 3,300 HCV patients were treated with various DAAs (not including the first-generation NS3 inhibitors, boceprevir and telaprevir) in 12 clinical centres in Israel. While in most cases, a sustained virological response was achieved, 49 patients (1.5%) failed therapy. NS3 or NS5A RAS were found in 42 (85.7%) of these 49 patients; 38/49 (77.6%) had clinically relevant RAS. Comparison between the clinical and demographic characteristics of patients failing therapy with clinically relevant RAS to those without clinically relevant RAS is presented in Table 1. Most (69.8%, 30/43 for whom treatment response was recorded) of the patients failing therapy experienced virological relapse following undetectable virus levels at the end of treatment. Different treatment regimens (with or without ribavirin) were used: 44.9% (22/49) were treated with sofosbuvir-based regimens (combined with either simeprevir [SIM]/ledipasvir [LDV]/velpatasvir [VEL] or daclatasvir [DCV]); 30.6% (15/49), failed treatment with ombitasvir/paritaprevir/ritonavir/+dasabuvir (OBV/PTV/r/DSV), while 14.3% (7/49) failed grazoprevir/elbasvir (GRZ/EBR) treatment.
Demographic and baseline characteristics of patients failing DAAs by clinically relevant RAS

Number of patients for whom clinical information was available.
Four cases of genotype misclassification were noted: GT1b reported as GT1a; GT1b reported as GT1a; GT3a having both GT3a and GT1b and GT1b misclassified as GT2. The corrected genotype is presented. AE, adverse events; ASV, asunaprevir; DAA, direct-acting antiviral; DCV, daclatasvir; EBR, elbasvir; GRZ, grazoprevir; LDV, ledipasvir; RAS, resistance-associated substitution; RBV, ribavirin; SIM, simeprevir; SOF, sofosbuvir; VEL, velpatasvir; 3D, ombitasvir/paritaprevir/ritonavir/+dasabuvir.
Overall, 45 of the patients were treated with a regimen that included an NS5A-targeted therapy and 28 with regimens that included NS3 targeted therapy. While 82.2% (37/45) of the NS5A exposed patients presented with clinically-relevant RAS, only 35.7% (10/28) of the patients failing a regimen that included NS3-targeted DAAs, had NS3 clinically relevant RAS (P<0.01).
All patients with GT1a, GT3b and GT4a (n=9) had clinically relevant RAS. On the other hand, only 72.5% (29) of the 40 patients with GT1b (n=35) or GT3a (n=5) infections, showed clinically relevant RAS (P=0.09; Table 2).
Sanger- and NGS-determined RAS, by treatment regimen and HCV subtype
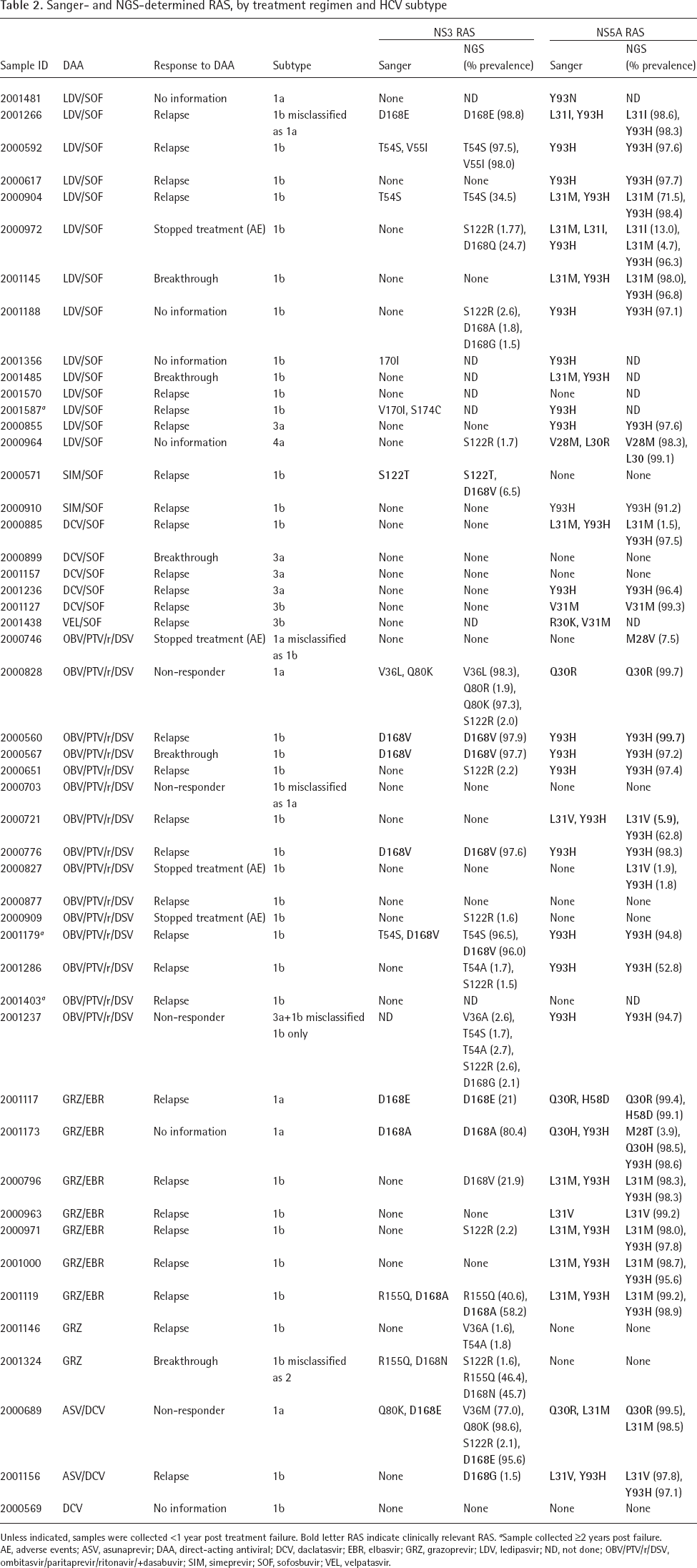
Unless indicated, samples were collected <1 year post treatment failure. Bold letter RAS indicate clinically relevant RAS.
Sample collected ≥2 years post failure. AE, adverse events; ASV, asunaprevir; DAA, direct-acting antiviral; DCV, daclatasvir; EBR, elbasvir; GRZ, grazoprevir; LDV, ledipasvir; ND, not done; OBV/PTV/r/DSV, ombitasvir/paritaprevir/ritonavir/+dasabuvir; SIM, simeprevir; SOF, sofosbuvir; VEL, velpatasvir.
RAS Prevalence by Treatment Regimen, HCV Subtype and Sequencing Technology
Table 2 summarizes all RAS identified by Sanger and NGS, in each of the analysed patient samples. The most abundant RAS were 168A/E/Q/G/N/V (32.6%, 16/49) in NS3. 93H/N (61.2%, 30/49), 31I/M/V (34.7%, 17/49) and 30R/H/K (12.2%, 6/49) were the most abundant in the NS5A. In 53.1% (26/49) of the cases, more than one RAS was identified at failure and in 36.7% (18/49) cases more than one RAS was identified in the NS5A protein. Specifically, L31 and Y93 double NS5A RAS were observed in 14/49 of the failures. Multiclass (NS3+NS5A) RAS were detected in 46.9% (23/49) of the patients of which 30% (7/23) could be directly attributed to the failing regimen.
All the seven patients who failed GRZ/EBR treatment (two with GT1a and five with GT1b) had at least one NS5A RAS, all of which were clinically relevant, and five of these patients had an additional NS3 RAS. While the two GT1a patients failing OBV/PTV/r/DSV had RAS (one had M28V NS5A RAS, detected by NGS at 7% prevalence, and the other Q30R, detected by both sequencing technologies), no RAS were identified in 4 of the 13 GT1b OBV/PTV/r/DSV failures. It should be taken into consideration, however, that these patients could also have RAS in the NS5B, which was not sequenced. Interestingly, 3/4 patients who stopped treatment due to adverse events had RAS, all of them clinically relevant.
NGS versus Sanger-Based Sequencing and Misclassification of HCV Genotypes
All 49 samples were assessed by Sanger sequencing and 42 of them were also analysed by NGS. Overall, more RAS were identified in both the NS3 and the NS5A by the NGS technology, however, all additional RAS detected by NGS showed a prevalence <15%. In four patients, the RAS detected by NGS were clinically relevant: in two patients, NGS identified NS5A RAS and in two patients, the RAS were in NS3 (Table 2).
In four cases (8.2%), sequencing analysis revealed that pre-therapy HCV genotype had been misclassified. One of these patients (2001266), was classified as a GT1a carrier, while both Sanger and NGS analyses revealed GT1b infection. Another case (2000703), was treated as a GT1a carrier, while both sequencing methods identified infection with GT1b. Patient 2001324 carried a GT1b misclassified as GT2. Patient 2001237 was a drug user who was infected initially with genotype GT3a, treated accordingly and considered to be cured. This patient failed the second round of treatment following reinfection with GT1b. While GT1b was determined for the NS3 region and GT3a for NS5A using Sanger sequencing, NGS revealed the coexistence of the two HCV strains, GT3a and GT1b, following analysis of reads spanning both NS5A and NS3 regions (Table 2).
Discussion
In this real-life national Israeli study, NS3 or NS5A RAS were identified in almost all patients failing various DAA therapies. High prevalence (77.5%, 38/49) of clinically relevant RAS was observed. Others reported lower prevalence of clinically relevant RAS in patients failing DAA therapy (54.8% [19] and 49.5% [20]) Suboptimal regimens used herein in several cases and the collection of most of the samples for resistance testing early after treatment failure, may account for this higher frequency.
The prevalence of clinically relevant NS3 RAS in patients treated with regimens containing protease inhibitors (35.7%, 10/28), was significantly lower than the prevalence of clinically relevant NS5A RAS (82.2%, 37/45) in patients treated with a combination that included NS5A inhibitors. This and earlier reports of similar observations [19,20] can be attributed to the fact that NS3 RAS tend to revert to the wild-type amino acid within 1 year after treatment failure, while NS5A RAS generally remain stable for years [21]. Indeed, in nearly all cases with clinically relevant NS3 RAS (7/8), samples were tested less than 1 year post failure. Nevertheless, it should be taken into account that resistance to protease inhibitors may remain unnoticed, even after mutant strains become undetectable by NGS [22].
In our study, 36.7% (18/49) of the treatment failures had more than a single RAS in the NS5A protein. Multiple RAS in NS5A after treatment failure have been previously observed [9,20], and the significance of this phenomenon was demonstrated in vitro: variants with any single mutation showed mild to moderate resistance to NS5A inhibitors, whereas variants with double mutations (for example, L31 and Y93 mutants) had a high fold of resistance [23]. Here, L31 and Y93 double RAS were observed in 14/49 of the failures.
Multiclass resistance was also frequently (46.9%) detected. Interestingly, in five cases, RAS in the NS3 or NS5A were identified, regardless of the specific anti-NS3 or NS5A treatments. Others found such ‘extra target’ RAS in around 9.0% of failing patients [20] and suggested that these are natural pretreatment variants. Unfortunately, baseline resistance testing was unavailable to support this assumption. However, since patients with multiclass RAS are still the most difficult to retreat, these findings are of clinical importance.
The case of the patient who was infected with both GT1b and GT3 strains emphasizes the added value of NGS over the conventional methods for deciphering HCV genotypes. The benefit of NGS was also demonstrated by the larger number of NGS-detected RAS compared to Sanger sequencing, some of which were clinically relevant. Nevertheless, due to the very low number of HCV treatment failures requiring resistance testing, the usage of deep sequencing technology should be carefully considered especially in light of its technical complexity and price. Furthermore, the prevalence of the resistance mutations detected by NGS only was low (between 1.5–8% of the viral quasispecies) and the clinical relevance of <15% resistance variants is debatable [24]. Indeed, systemic next generation sequencing is currently not recommended [24].
Overall, clinically relevant RAS were identified in all patients infected with GT1a, GT3b and GT4a strains. In GT3a infections, three patients had clinically relevant RAS, and two (failing DCV/SOF) had no resistance mutations. However, both of these GT3a patients had cirrhosis, a condition that may negatively affect treatment outcomes [25] and lead to failure. In 9 of the 35 treatment failures bearing the GT1b strain, no clinically relevant RAS were identified and four patients were RAS-free. Included was a case that was misclassified and treated as GT2, a case with chronic renal failure (due to hereditary neuropathy) comorbidity to the viral hepatitis and a chronic HCV patient erroneously considered to be acutely infected and treated as such. Moreover, RAS analysis in some of the patients without clinically relevant RAS was performed on a sample taken more than 1 year after failure, a period during which the wild-type virus may have evolved to become the dominant variant [21].
In four cases, suboptimal treatment was administrated due to genotype misclassification by commercial genotyping assays. These results highlight the need for HCV sequencing as the most efficient method for subtype/genotype assignment [4].
This study had several limitations. Although the data were collected from all major medical centres, it did not include results from health maintenance organizations in the country. Analysis was restricted to samples taken after treatment failure; data regarding baseline resistance are not routinely collected and were not available. An additional inherent limitation related to the small DAA treatment failure sample size. However, the study findings clearly demonstrated that in real-life, when treatment is cost related and suboptimal regimens may be employed, RAS characterize treatment failures. Therefore, the current recommendations of the European Association of the study of the liver (EASL) to perform resistance testing in patients failing DAA therapy should be employed [6]. Recently, Dietz et al. [7] described patterns of RAS in treatment failures and proposed several retreatment options with broadly available DAAs according to the pattern of identified RAS. These can be used to guide retreatment strategy. DAAs with a high genetic barrier and longer treatment duration should be considered in patients with RAS. Our findings, together with more real-life data, will contribute to the optimization of retreatment in DAA failure patients.
