Abstract
Background
Infections due to drug-resistant herpes simplex viruses (HSV) represent an important clinical concern, especially in immunocompromised patients. The present study was aimed at detecting acyclovir (ACV) susceptibility in HSV clinical samples.
Methods
A total of 13 HSV-positive clinical samples (5 HSV-1 and 8 HSV-2) recovered from patients (1 immunocompromised and 12 of unknown immune status) were included in the study. The genotypic analysis involved an initial UL23 (thymidine kinase) gene sequencing, followed by a confirmatory phenotypic assay using plaque reduction technique.
Results
Two novel amino acid changes, A37V and H283N, were detected in HSV-1 positive clinical samples, which were found to be susceptible to acyclovir (half maximal effective concentration = 1.5 μM) by plaque reduction assay.
Conclusions
These two novel amino acid changes could be therefore considered as natural polymorphisms, a phenomenon widely associated with the HSV-UL23 gene.
Introduction
According to a recent prevalence study, the global burden of herpes simplex virus (HSV) infection is estimated to be 67% for herpes simplex virus type 1 (HSV-1) and 11.3% for herpes simplex virus type 2 (HSV-2), among people aged between 0–49 years and 15–49 years, respectively, reflecting the colossal burden of the disease [1,2]. Although a number of therapeutic agents such as acyclovir (ACV), valacyclovir, famciclovir, penciclovir, foscarnet and cidofovir may be used for the treatment of HSV infections [3], simultaneously the problem of drug resistance has also emerged, especially in the immunocompromised population [4].
Phenotypically, ACV resistance in HSV strains may develop through four mechanisms, which are complete deficiency in viral thymidine kinase (TK) activity, decreased activity or production of low amounts of viral TK, viral TK protein with altered substrate specificity or viral DNA polymerase gene (DNA pol) with altered substrate specificity [5,6]. Mutations in the TK and DNA pol genes are responsible for mediating drug resistance, and the TK gene mutation alone accounts for 95% of drug resistance. A total of 60–80% of TK gene mutations are due to nucleotide insertions or deletions occurring frequently in G or C homopolymer repeats, considered as the hot spots for point mutations [7,8]. Nucleotide substitutions, occurring mostly in the active or conserved sites of the TK gene, contribute up to 40% of TK gene mutations [8]. The worldwide prevalence of ACV-resistant HSV infections is nearly 5% in various immunocompromised patients but may reach 14–30%, particularly in transplant recipients [9,10].
There is a dearth of background data on the drug susceptibility status of HSV strains circulating in India [11]. To date, the prevalence studies from India reveal 3% ACV resistance among HSV-1 strains and 7.8% among HSV-2 strains in immunocompromised patients [12]. The present study aimed at characterizing the TK gene of HSV-1 and −2 clinical samples for the presence of mutations and their effects on in vitro acyclovir susceptibility. The information derived thereof is expected to update the existing knowledge on TK gene polymorphisms resulting in susceptibility or resistance of HSV- 1 and −2.
Methods
Clinical samples
A laboratory-based retrospective study was designed on real-time PCR positive cases of HSV (CT value below 30). A total of 13 samples comprising 5 HSV-1 and 8 HSV-2, archived at Manipal Centre for Virus Research, during the period of January 2012 to December 2016, from south-west Indian states, were included in the study.
Genotypic assay
Viral DNA extraction
Viral DNA was extracted from the clinical sample by using QIAamp® DNA Mini Kit (QIAGEN®, Hilden, Germany), according to the manufacturer's instructions. The extracted nucleic acids were stored at −20°C for further use.
TK Gene Amplification
The full-length TK (UL23; 1131bp) gene of HSV-1 and 2 was amplified following a previously described protocol with slight modifications [4]. The selected primers bound at a few hundred base pairs distant to either side of the TK gene, to yield a final product of size 1622bp for HSV-1 and 1595bp for HSV-2. The amplification and sequencing were carried out using HSV specific primers, forward primers 5'-TAACCCCCACGAAC-CATAAA-3’ (HSV-1) and 5'-GTACCCACGGCCCAAA-GAG-3’ (HSV-2), and reverse primer 5'-CGAATTC-GAACACGCAGAT-3’ (HSV-1 and −2).
The TK gene was amplified using 25 μl of reaction mixture containing 5 μl of the extracted nucleic acid, 4.5 μl of nuclease free water, 13.5 μl of buffer mix (ThermoFisher Scientific, Waltham, MA, USA), 0.5 μl (10 μM) of each primer (Merck, Darmstadt, Germany) and 1.0 μl of DNA polymerase enzyme (ThermoFisher Scientific). Amplification was carried out using ProFlex PCR System thermal cycler (ThermoFisher Scientific). Cycling conditions involved an initial denaturation of 5 min at 94°C, followed by 40 cycles of 1 min at 94°C (denaturation), 1 min at 60°C (annealing) and 2 min at 72°C (extension) with a final extension of 10 min at 72°C.
TK Gene Sequencing
The amplified products were purified using GenElute™ Gel Extraction Kit (Merck) and sequenced using the 3500xL Genetic Analyzer (Thermo Fisher Scientific). The sequences were analysed using Sequencher 5.4.6 software. All nucleotide and amino acid sequences were compared using MEGA 7.0.18 software with published sequences of the reference strain 17 (HSV-1) and HG52 (HSV-2); GenBank accession numbers X14112 and Z86099, respectively.
Phenotypic assay
HSV samples with novel amino acid changes, detected by the genotypic method, were further analysed using phenotypic assay. The assay was performed using the standard protocol with slight modifications [13]. Half maximum (50%) effective concentrations (EC50) of acyclovir (Merck) was calculated using GraphPad Prism 5.
Ethical approval
The study was approved by the Institutional Ethics Committee, Manipal Academy of Higher Education, Manipal (Ref No. MUEC/014/2017).
Results
Description of the study population
Among the five HSV-1 cases, two (40%) were males, whereas, among the eight HSV-2 cases, six (75%) were males. The mean age of HSV-1 cases was 22 years and for HSV-2 was 39 years. The clinical specimens included oral vesicle lesion swabs (HSV-1 and −2) and genital vesicle swabs (HSV-2). No information on prior antiviral therapy was available.
Mutations in TK gene
Nucleotide substitutions identified within the 1131bp coding sequence, for HSV-1 and HSV-2 strains, were 15 and 4, respectively (Table 1).
HSV TK mutations detected in this study

Bold and underlined font: novel amino acid mutations. HSV, herpes simplex virus; TK, thymidine kinase.
Overall, the number of amino acid changes was nine for HSV-1 and four for HSV-2 (Table 1), representing 2.4% and 1.1% of the total codons of the protein, respectively. Among the nine amino acid changes identified in the HSV-1 samples, the mutations N23S, K36E and A265T were frequently observed, whereas, R89Q was detected only in three samples. The mutations, C6G, L42P, V348I (sample number 1), H283N (sample number 3) and A37V (sample number 5) were unique to their respective samples. Among the HSV-2 samples, four amino acid changes were identified within the TK gene, of which G39E was observed in all samples, whereas N78D was present in only five samples. The mutations, A27T and R220K were unique to sample number 11.
Almost all the amino acid changes were located in the non-conserved domains of the viral protein encoded by both the HSV-1 and −2 TK genes (Figure 1), except the mutation R220K in HSV-2, which was located in the conserved domain.
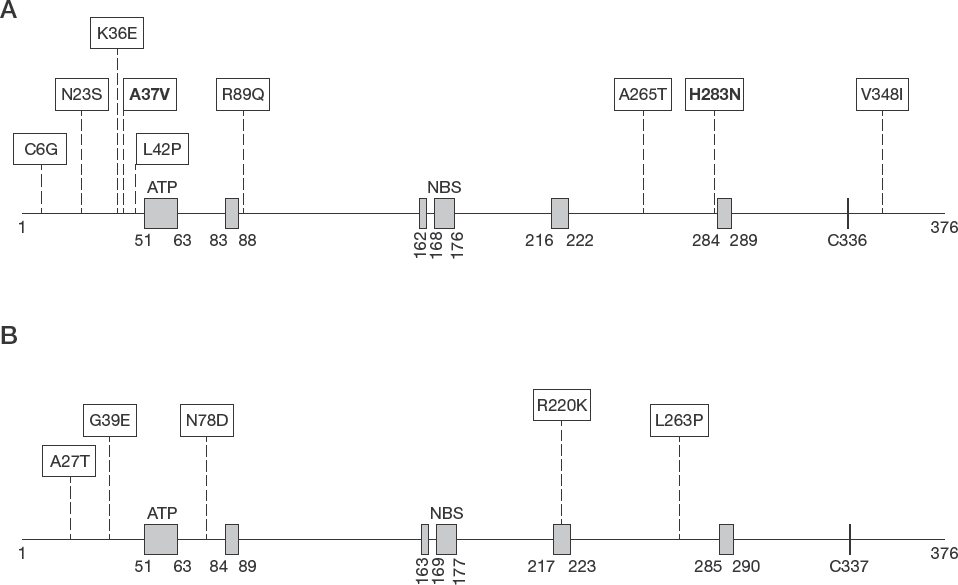
Location of identified amino acid substitutions in HSV thymidine kinase.
Phenotypic ACV susceptibility of HSV isolates
Two samples of HSV-1, identified with novel amino acid changes, were found susceptible to acyclovir as observed from the EC50 values (1.54 μM [r 2 =0.97] and 1.51 μM [r 2 =0.98] for sample numbers 3 and 5, respectively).
Discussion
Consistent with previous studies, our data indicated a high prevalence of amino acid polymorphisms in the TK gene of HSV-1 (nine amino acid polymorphisms) compared to HSV-2 (four amino acid polymorphisms) [14]. The amino acid substitutions, C6G, N23S, K36E, L42P, R89Q, A265T and V348I, observed in HSV-1 samples (n=5), are documented as natural polymorphisms associated with acyclovir susceptible phenotypes [14].
Two new mutations, at codons 37 (A37V) and 283 (H283N), identified by genotypic assay, had no reported association with either ACV-sensitive or -resistant HSV-1 strains. To predict the impact of these mutations on TK gene function, we determined their location with respect to the highly conserved and functional domains of the TK gene. Both the new mutations, A37V and H283N, were located outside the active/conserved site of the enzyme. Although most polymorphism-associated mutations are located outside, and most of the resistance-related mutations are situated inside active or conserved gene centre, natural polymorphisms can also involve conserved domains and mutations of resistance can be found in non-conserved regions [15].
TK gene sequencing helped in the rapid identification of the HSV mutants. However, once the mutants were identified, the phenotypic assay was performed to evaluate the role of the novel amino acid changes in ACV-resistance/susceptibility. From the plaque reduction assay, it was found that both the HSV-1 samples, with the new amino acid mutations, were susceptible to acyclovir (EC50: 1.51 μM). The EC50 cutoff values for resistance to ACV is 6.5 μM for HSV-1 [8,16]. Therefore, it may be inferred that the mutations, A37V and H283N in HSV-1 samples, do not affect the enzymatic activity of TK and probably contribute to natural polymorphism, a phenomenon largely associated with the HSV TK gene [9,17].
In HSV-2 samples, four amino acid polymorphisms, G39E, N78D, A27T and R220K were detected. The mutations, N78D and A27T, are well documented natural polymorphisms with no association of acyclovir resistance [14]. The mutation, R220K, though present in the conserved region of HSV-2 TK, has been also associated with an ACV-susceptible strain [18]. Regarding the mutation G39E, it had been previously associated with acyclovir-resistant phenotype in combination with other mutations [19]. However, in a recent study, the role of G39E as a natural polymorphism has been clearly demonstrated [20]. Since no new mutations were observed, we did not further evaluate the HSV-2 samples for phenotypic characterization.
In this study, we limited our focus on sequencing the TK gene. Nevertheless, a combined sequence analysis of both the viral TK and DNA polymerase genes can be considered in order to document the role of any new mutations that may lead to acyclovir resistance among HSV-1 and HSV-2 strains. Moreover, a smaller sample size and the lack of information on the immune status of the samples could be considered as other limitations of the present study.
Footnotes
The authors declare no competing interests.
Acknowledgements
This work was funded by ICMR Virology Network Laboratory Grade-I; file. No:5/8/7/15/2010/ECD-1, Manipal Academy of Higher Education, Manipal for providing the necessary infrastructural support.
