Abstract
Background:
Ribavirin is a broad-spectrum antiviral agent that derives some of its activity from inhibition of cellular inosine monophosphate dehydrogenase (IMPDH), resulting in lower guanosine triphosphate (GTP) levels. Here we report the biological activities of three ribavirin analogues.
Methods:
Antiviral activities of test compounds were performed by in vitro cytopathic effect inhibition assays against influenza A (H1N1, H3N2 and H5N1), influenza B, measles, parainfluenza type 3 (PIV-3) and respiratory syncytial viruses. Compounds were modelled into the ribavirin 5‘-monophosphate binding site of the crystallographic structure of the human type II IMPDH (hIMPDH2) ternary complex. Effects of compounds on intracellular GTP levels were performed by strong anion exchange HPLC analysis.
Results:
Of the three compounds evaluated, the 5-ethynyl nucleoside (ETCAR) exhibited virus-inhibitory activities (at 1.2–20 μM, depending upon the virus) against most of the viruses, except for weak activity against PIV-3 (62 μM). Antiviral activity of ETCAR was similar to ribavirin; however, cytotoxicity of ETCAR was greater than ribavirin. Replacing the 5-ethynyl group with a 5-propynyl or bromo substituent (BrCAR) considerably reduced antiviral activity. Computational studies of ternary complexes of hIMPDH2 enzyme with 5‘-monophosphates of the compounds helped rationalize the observed differences in biological activity. All compounds suppressed GTP levels in cells; additionally, BrCAR suppressed adenosine triphosphate and elevated uridine triphosphate levels.
Conclusions:
Three compounds related to ribavirin inhibited IMPDH and had weak to moderate antiviral activity. Cytotoxicity adversely affected the antiviral selectivity of ETCAR. As with ribavirin, reduction in intracellular GTP may play a role in virus inhibition.
Introduction
For many years the antiviral agent ribavirin (1-β-D-ribofuranosyl-1,2,4-triazole-3-carboxamide,
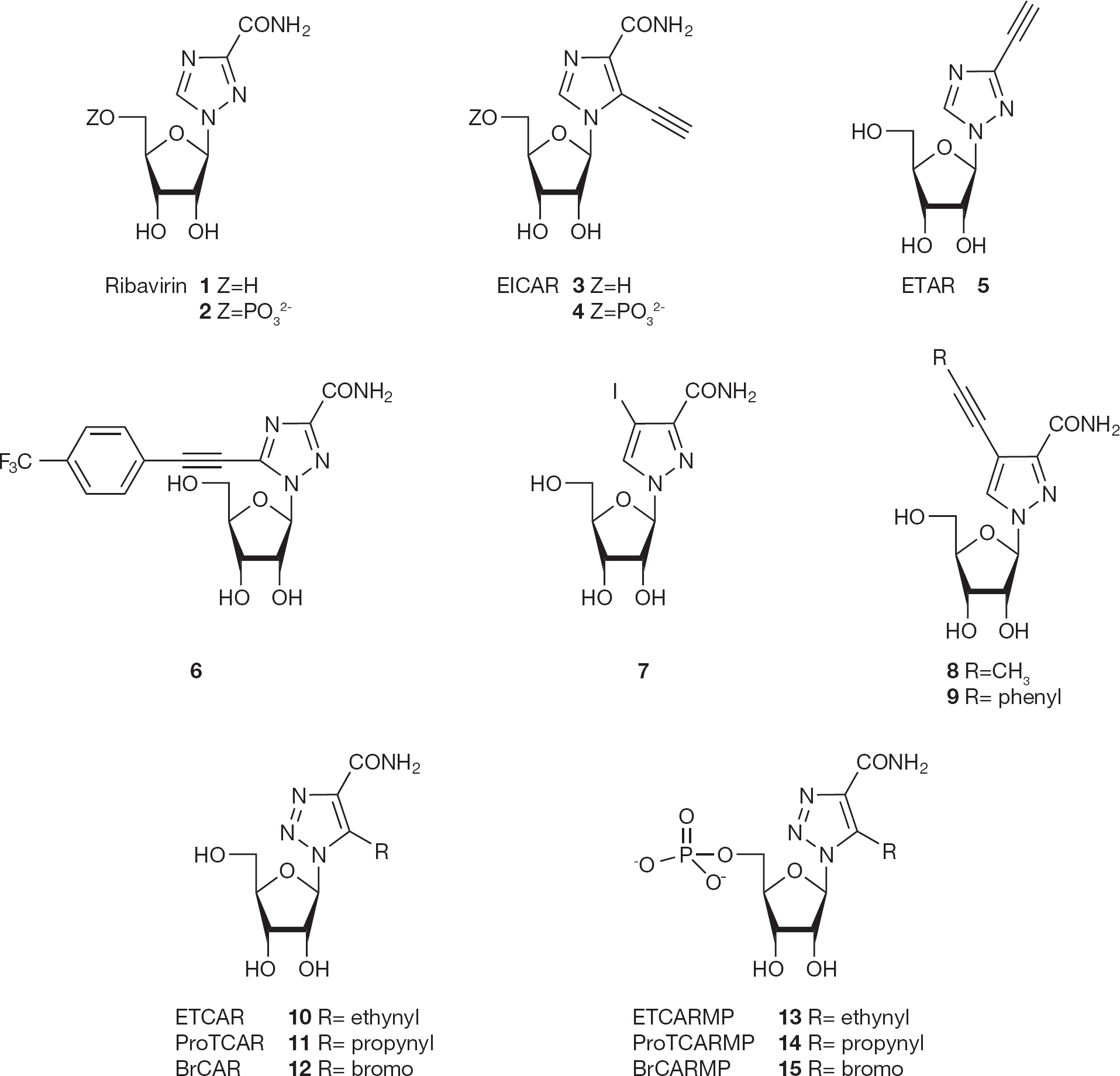
Chemical structures of triazole compounds related to ribavirin and ribavirin 5′-monophosphate
Along with its application as a drug, ribavirin has become a lead compound for many other nucleoside analogues. One of them, 5-ethynyl-1-β-D-ribofuranosylimidazole-4-carboxamide (EICAR;
Materials and methods
Chemistry
General methods and techniques were applied to the synthesis of ETCAR (
Synthesis of 5-bromo-1-(ß-D-ribofuranosyl)-1H-[1,2,3]triazole-4-carboxamide, 12
To a suspension of 1-(5-O-acetyl-2,3-O-isopropylidene-β-D-ribofuranosyl)-5-amino-1H-[1–3]triazole-4-carboxylic acid amide [29] (790 mg, 2.31 mmol, dried in vacuo over P2O5, at 90°C, for 4 h) in bromoform (34.4 g, 136 mmol) was added isoamyl nitrite (949 mg, 8.1 mmol). The mixture was stirred at 100°C for 2 h, then it was applied onto a silica gel column. The column was eluted with CH2Cl2 (in order to collect the bromoform) followed by CH2Cl2-MeOH (95:5) to give crude product. It was next re-chromatographed using ethyl acetate/hexane (1:1→2:1) to afford 537 mg (55% yield) of 5-bromo-1-(5-O-acetyl-2,3-O-isopropylidene-β-D-ribofuranosyl)-1H-[1–3]triazole-4-carboxylic acid amide: 1H NMR (DMSO-d6) δ 7.98 and 7.65 (2xs, 2H, CO-NH2), 6.22 (s, 1H, 1′-H), 5.63 (d, J2′,3′ =6.0 Hz, 1H, 2′-H), 5.04 (dd, J3′,2′ =6.0Hz, J3′,4′ =2.0 Hz, 1H, 3′-H), 4.47 (m, 1H, 4′-H), 4.00 and 3.81 (2×dd, J5′,5′ =11.6 Hz, J5′,4′ =7.2 Hz, 2H, 5′-H), 1.94 (s, 3H, OAc), 1.53 and 1.36 (2×s, 6H, 2×CH3). 13C NMR (DMSO-d6) δ 169.84 (CO), 160.40 (CO-NH2), 139.55 (C-4), 115.19 (C-5), 112.93 (CH3)2 C, 91.60 (C-1′), 85.90 (C-4′), 83.37 (C-2′), 81.40 (C-3′), 63.10 (C-5′), 26.64, 24.97 (CH3)2 C, 20.38 (CO-CH3). HRMS [M+Na]+ calcd for C13H17BrN4NaO6: 427.0224; found: 427.0219.
Deacetylation of the compound (435 mg, 1.04 mmol) was performed with sodium methoxide (83 mmol) in anhydrous methanol (13 ml) at 0°C. After the reaction was complete (3 h), neutralization with Dowex H+ (100 mg) prewashed with anhydrous methanol followed. The filtrate was evaporated and the residual oily product was dried in vacuo to give 370 mg (98% yield) of 5-bromo-1-(2,3-O-isopropylidene-β-D-ribofuranosyl)-1H-[1–3]triazole-4-carboxylic acid amide. 1H NMR (DMSO-d6) δ 7.97 and 7.65 (2xs, 2H, CO-NH2), 6.17 (s, 1H, 1′-H), 5.62 (d, J2′,3′ =6.0 Hz, 1H, 2′-H), 4.98 (dd, J3′,2′ =6.0 Hz, J3′,4′ =1.6 Hz, 1H, 3′-H). 4.91 (t, J5′OH,5′ =5.6 Hz, 1H, 5′-OH), 4.21 (dt, J4′,5′ =6.0 Hz, J4′,3′ =1.6 Hz, 1H, 4′-H), 3.18 (m, 2H, 5′-H), 1.52 and 1.36 (2xs, 6H, 2xCH3). 13C NMR (DMSO-d6) δ 160.45 (CO-NH2), 139.52 (C-4), 115.16 (C-5), 112.62 (CH3)2 C, 91.57 (C-1′), 89.04 (C-4′), 83.19 (C-2), 81.78 (C-3′), 60.86 (C-5′), 26.65, 24.95 (CH3)2 C. HRMS [M+Na]+ calcd for C11H15BrN4 NaO5: 385.0118; found: 385. 0126.
Isopropylidene blockade was cleaved with an ice-cooled 80% aqueous trifluoroacetic acid (4 ml/1 mmol of substrate). After the reaction was complete (0°C, 2 h), the solution was co-evaporated several times with EtOH, then with anhydrous toluene. The oily residue was dissolved in EtOAc/MeOH (10:1) and evaporated slowly to enforce precipitation of
In vitro antiviral evaluations Viruses and cells
Influenza A/California/07/2009 (H1N1), A/Perth/16/2009 (H3N2) and B/Florida/4/2006 viruses were obtained from the Centers for Disease Control and Prevention (Atlanta, GA, USA). Influenza A/Duck/MN/1525/81(H5N1), a low pathogenic avian virus isolated in the United States, was kindly provided by Robert Webster (St Jude Children's Research Hospital, Memphis, TN, USA). Experiments with the above influenza virus strains were carried out in Madin–Darby canine kidney (MDCK) cells. Parainfluenza type 3 (PIV3) virus strain 14702, RSV strain A2 and measles virus strain CC were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA). Experiments with PIV-3 and RSV were performed in African green monkey kidney (MA-104) cells, whereas measles virus was assayed in the Vero 76 line of African green monkey kidney cells. The cell lines were all obtained from ATCC.
Antiviral assays
Inhibition of viral cytopathic effect assays were performed as previously described using confluent monolayers of cells in 96-well microplates [30], with cytotoxicity determined in parallel in uninfected cells. Trypsin (10 units/ml; Sigma–Aldrich, St Louis, MO, USA) in serum-free medium was required to facilitate the replication and cytopathology of influenza A and B viruses. Assays with the other viruses were performed in medium containing 2% fetal bovine serum. Quantitation of percent cytopathic effect was accomplished by treating cells with 0.011% neutral red dye for 2 h, then determining dye uptake into viable cells [31]. Unincorporated dye was removed from cells by aspiration and rinsing with phosphate-buffered saline, then the absorbed dye was eluted from the cells for 30 min with 0.1 ml of 50% Sörensen's citrate buffer (pH 4.2)/50% ethanol. The plates were read for optical density determinations at 540 nm with 405 nm reference. Readings were converted to percentages of uninfected control and normalized to the virus control. The 50% virus inhibitory concentration (IC50) values and 50% cytotoxic concentration (CC50) values were determined by linear regression analysis. Selectivity index (SI) values were calculated as CC50/IC50.
Computational methods with IMPDH
Analysed 5′-monophosphates of 1,2,3-triazole nucleosides:
All water molecules were removed and hydrogen atoms were added. While coordinates of all heavy atoms of the enzyme were kept frozen, hydrogen atoms positions were optimized with the use of adopted basis Newton-Raphson minimization algorithm (max steps =500, RMS gradient =0.01, distance-dependent dielectrics as implicit solvent model). Compounds
For the docking procedure the dock ligands (CDOCKER) protocol, an implementation of the CDOCKER algorithm [33] within the Discovery Studio 3.0 package was used. Random ligand conformations were generated from the initial ligand structure through high temperature molecular dynamics at 700 K. The binding site was defined from the volume of a co-crystallized ligand (RMP). The random conformations were refined by grid-based simulated annealing and a final full force field minimization. In all calculations for proteins and ligands, CHARM forcefield with MMFF94 partial charge rules was used. To account for protein flexibility, the ligand binding site of each model was further refined by restraint energy minimization.
HPLC analysis of intracellular nucleotide pools
Confluent MDCK cell cultures in 6-well microplates were treated with compounds in serum-free medium for 24 h. The medium was aspirated from each well, followed by precipitation and breaking open of the cells with 3.5% perchloric acid and neutralization of the acid with 1 N KOH/1 M imidazole. The supernates were collected from each well and frozen at −80°C until analysed by HPLC.
HPLC analysis of the samples was conducted as previously published [34]. Briefly, chromatographic separations were performed using an HPLC apparatus (Waters Corp., Milford, MA, USA) fitted with a 10×250 mm column packed with Whatman partisil SAX resin (Phenomenex, Torrance, CA, USA). A linear gradient from 10 mM to 1 M potassium phosphate (Sigma–Aldrich) at pH 5.0 was run over 30 min at 1 ml/min. The 1 M buffer ran an additional 2 min before re-equilibrating the column in low salt buffer. Detection of nucleotides was made at 260 nm with a Waters LC spectrophotometer and the peaks analysed using an integrator (Shimadzu Scientific Instruments, Columbia, MD, USA). Relative peak areas of D282-treated cells were compared to untreated cells and expressed as percentages of control levels.
Results
Synthesis of compounds
Alkynyl-1,2,3-triazole nucleosides
In vitro antiviral and cytotoxic activities
Primary in vitro screening tests were conducted against a broad panel of 12 RNA viruses. Further assays allowed us to select compounds
Antiviral activities of triazole compounds and ribavirin against certain respiratory viruses in cell culture
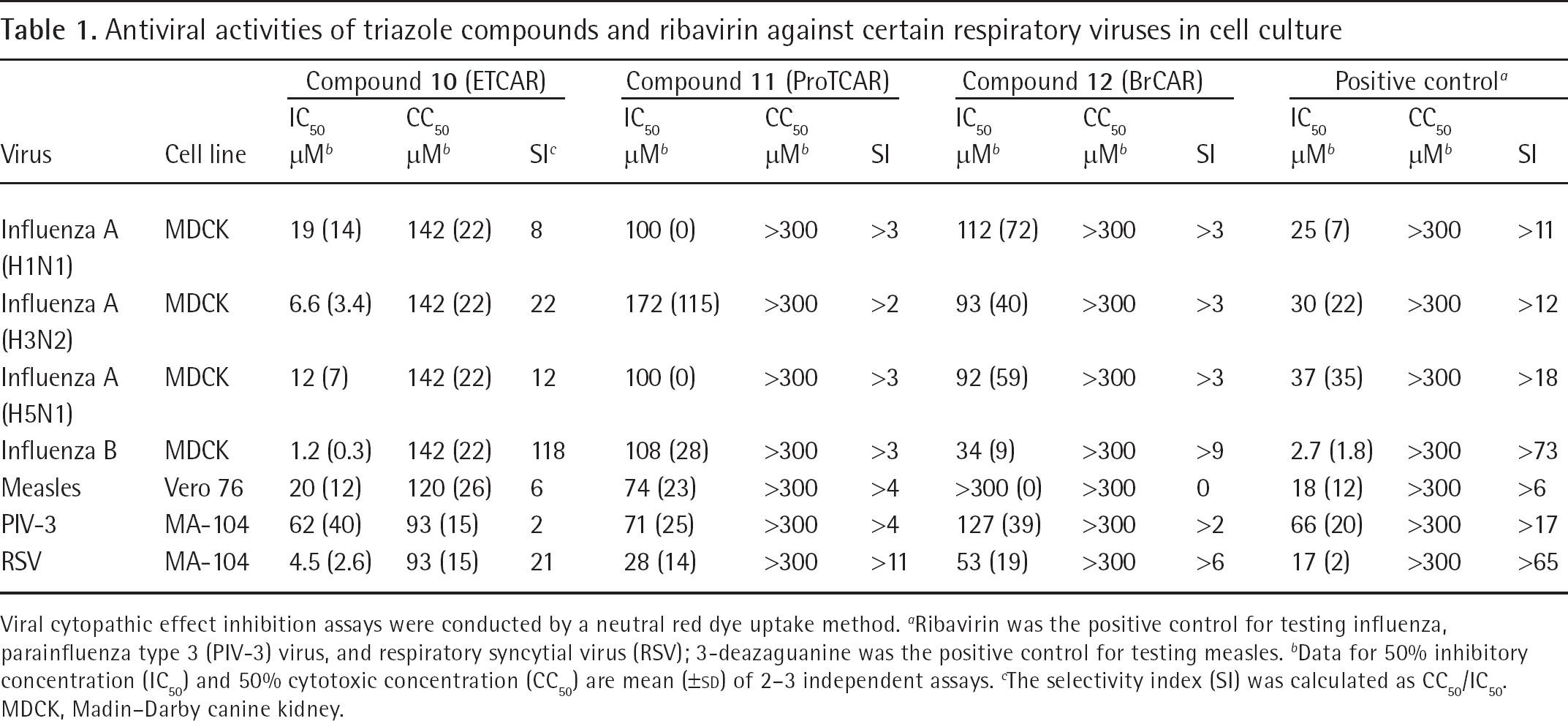
Viral cytopathic effect inhibition assays were conducted by a neutral red dye uptake method.
Ribavirin was the positive control for testing influenza, parainfluenza type 3 (PIV-3) virus, and respiratory syncytial virus (RSV); 3-deazaguanine was the positive control for testing measles.
Data for 50% inhibitory concentration (IC50) and 50% cytotoxic concentration (CC50) are mean (±SD) of 2–3 independent assays.
The selectivity index (SI) was calculated as CC50/IC50. MDCK, Madin–Darby canine kidney.
Interaction of compounds with IMPDH
The structural resemblances of
Compound
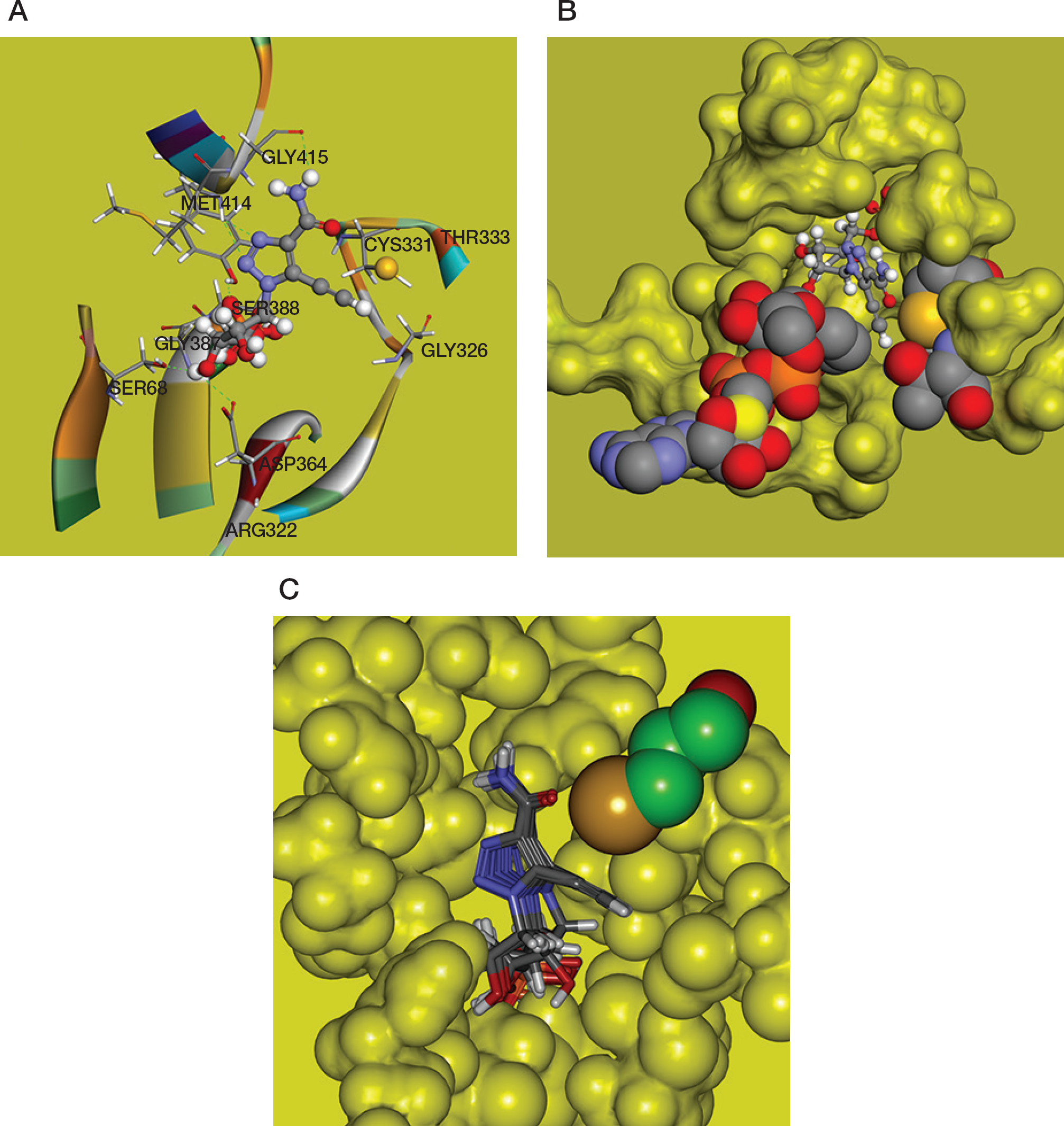
Computational modelling and docking of the compound
The 4,5-disubstituted 1,2,3-triazole ring is well accommodated in the substrate binding site of the hIMPDH2 enzyme and makes three hydrogen bonds with the active site flap: N(2) and N(3) of triazole form bifurcated hydrogen bonds with the Met414 backbone amide NH group, N(6)H atoms of 4-carboxamide group form a hydrogen bond with Gly415 backbone carbonyl. The heterocyclic N(3) and amide N(6) atoms are on the same side of the triazole moiety. In this configuration N(6) and O(6) atoms are spatially similar to the N(6) and N(1) atoms of adenosine, which can be of great importance for the biological activity of ETCARMP (
By contrast, examination of the X-ray structures of the RMP complexes with IMPDH revealed that in all of them ribavirin (
Similar to EICARMP (
CDOCKER docking results are shown in Figure 2C. One can see that dominant poses of ETCARMP are similarly located at the IMP binding pocket, as it was observed in the crystal structures of IMPDH complexes with IMP, XMP* or XMP covalently bound to the catalytic cysteine residue.
Compound
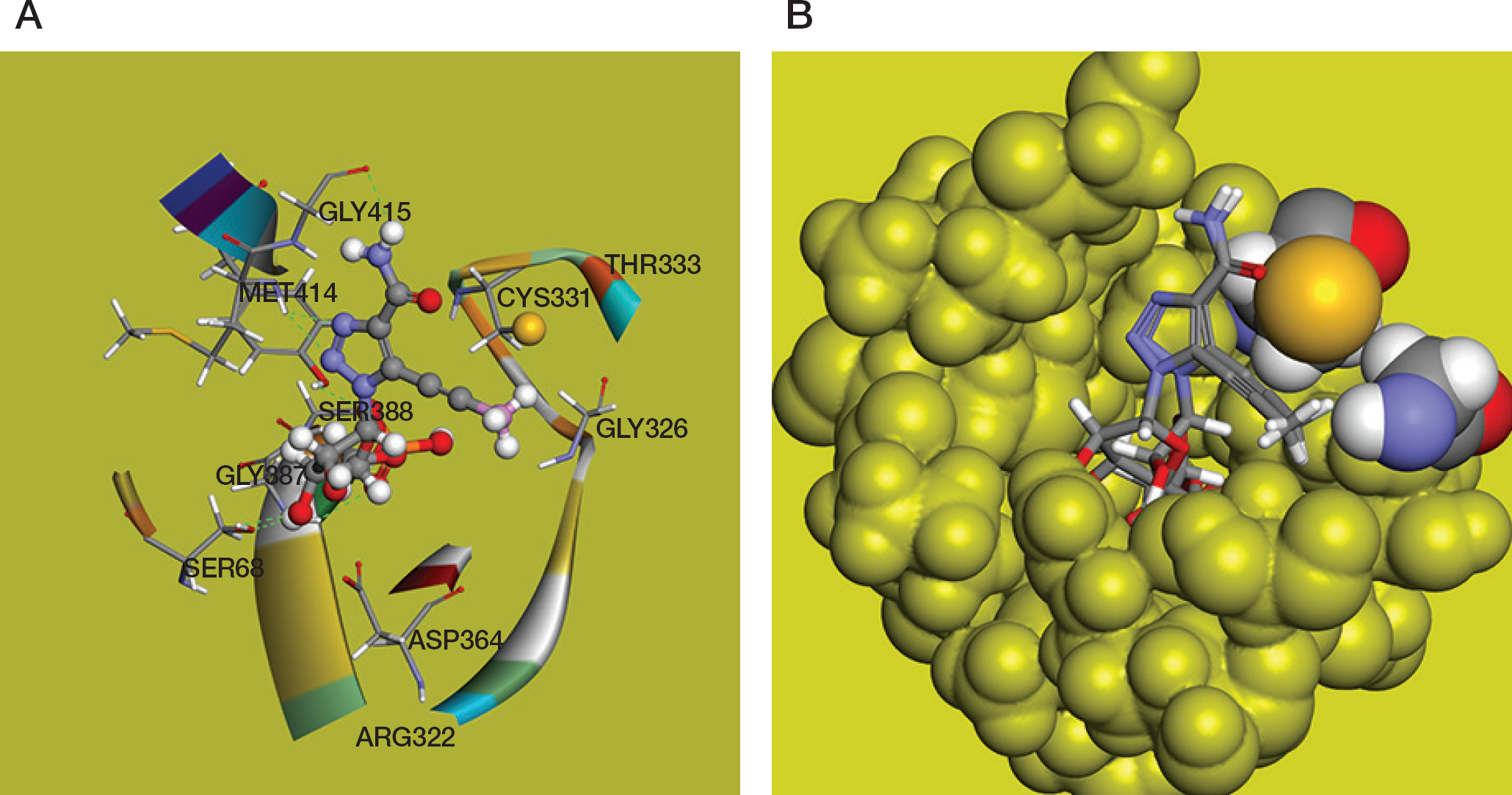
Computational modelling and docking of the compound
Figure 3B presents dominant poses of ProTCARMP obtained in the CDOCKER docking procedure. The orientation of ProTCARMP in the substrate binding site is like ETCARMP with the reorientation of the 5-propynyl group shifting away from the Gly326 residue.
The bulky 5-bromo substituent in compound
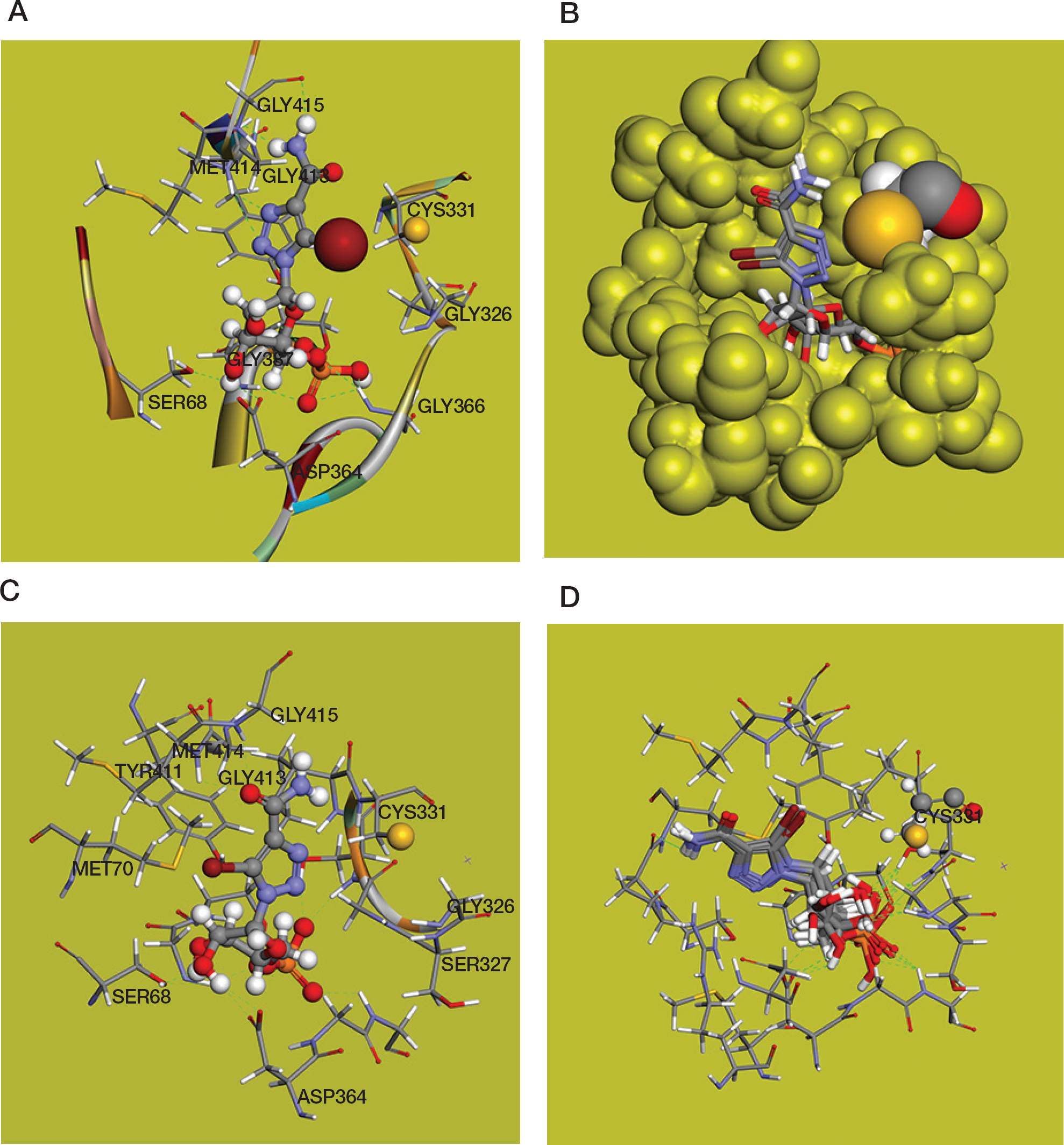
Computational modelling and docking of the BrCAR with a 5′-bromo substituent modelled into the substrate binding site of hIMPDH2 enzyme
Effects of compounds on intracellular nucleotide pools
The IMPDH assays described above do not directly prove whether treatment of cells with the compounds leads to reductions in intracellular GTP concentrations as a result of inhibition of IMPDH. For a direct demonstration we treated MDCK cells with compounds
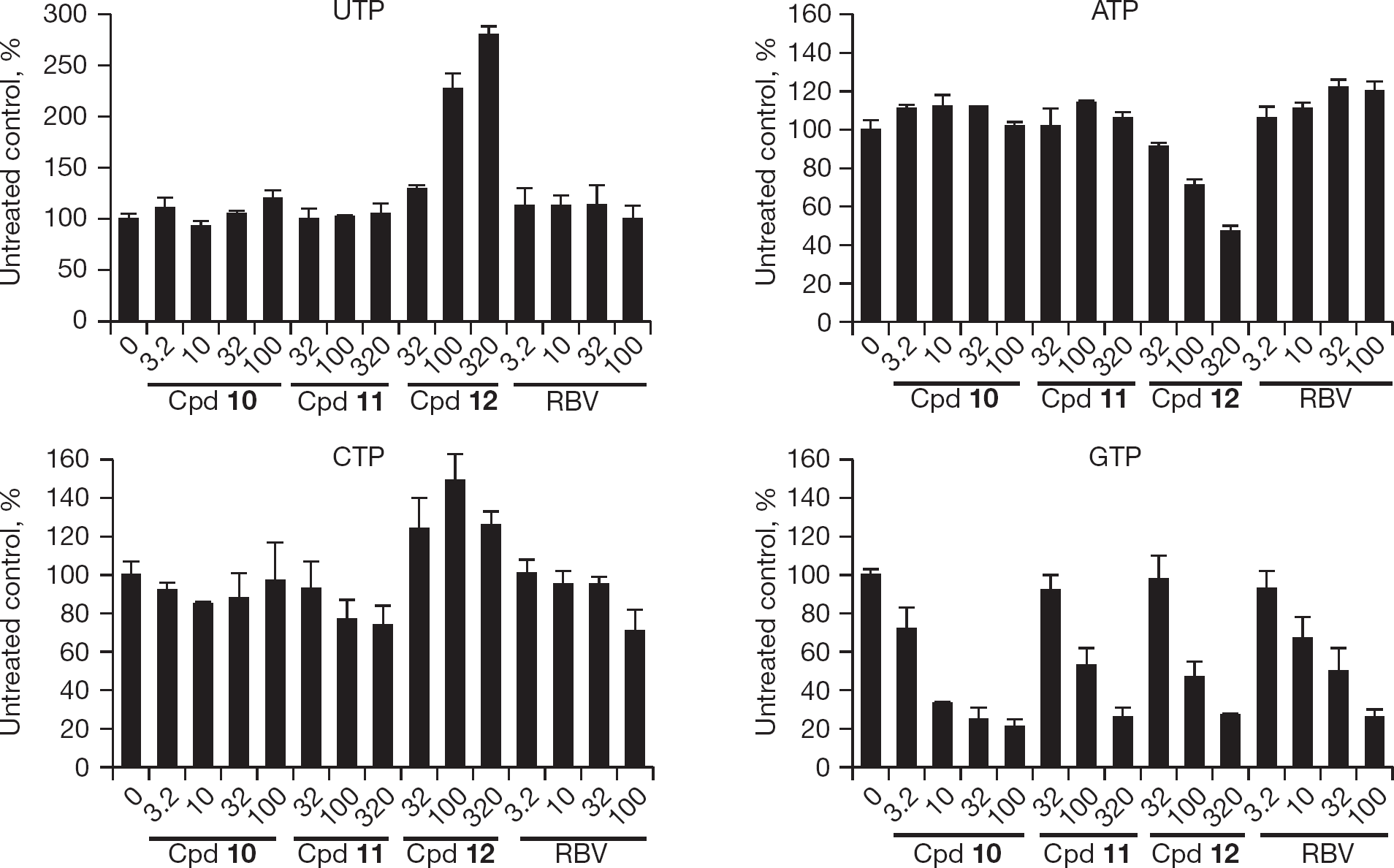
Nucleotide pool changes in Madin–Darby canine kidney cells treated with varying micromolar concentrations of compound
Discussion
In these studies, we demonstrated how small or profound changes at the C-5 position of 1,2,3-triazole-4-carboxamide, a pseudobase isomeric to that of ribavirin, modulate the antiviral activity of nucleosides
Results of studying effects of compounds on intracellular nucleotide pools revealed a more complex picture than simple reduction in GTP levels. Compounds
It has been pointed out that a drug combination therapy of influenza virus infections would be feasible to prevent an emergence of new drug-resistant strains of the virus [51]. Antiviral drugs used in combination should exert their activities by different modes of action [52]. Thus, ribavirin or a selection of its analogues could be important components of such combinations.
Footnotes
Acknowledgements
Financial support from the Polish Ministry of Science and Higher Education (project numbers N N405 2516 33 and N N519 4050 37) is gratefully acknowledged. Cell culture studies were funded by contract N01-AI-15435 from the Virology Branch, Division of Microbiology and Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, USA.
The authors declare no competing interests.
