Abstract
Background:
Current approved anti-HBV treatment cannot completely eliminate HBV infection, and emergence of resistant virus is an important treatment issue. Effective anti-HBV agents with different mechanisms of action on novel target sites are needed for the treatment of HBV infection and for combating the resistant virus, alone or in combination with current anti-HBV strategies. Helioxanthin analogue 8-1 displayed potent anti-HBV activity in human HBV in vitro and in animal models, with a unique antiviral mechanism. Its antiviral activity in other HBV system needs further study.
Methods:
The anti-duck hepatitis B virus (DHBV) activity of 8-1, an analogue of a natural product, helioxanthin, was studied in the DHBV inducible cell line, dstet5, in comparison to and in combination with the nucleoside analogue, lamivudine (3TC).
Results:
Helioxanthin analogue 8-1 exhibited anti-DHBV activity as demonstrated by quantification of viral DNA, RNA, covalently closed circular DNA and protein synthesis. Analogue 8-1 did not affect the stability of cellular macromolecules and did not have a sustained antiviral effect after drug removal. When DHBV replication was induced, virus-harbouring cells were more susceptible to the cytotoxicity of 8-1 than non-induced cells.
Conclusions:
8-1 exhibited effective inhibition on DHBV replication. The combination of 8-1 with 3TC resulted in additional anti-DHBV activity. Viral induced cells displayed higher susceptibility to 8-1 treatment than non-induced cells. HBV X protein might not be an essential factor in the initiation of the biological activity of 8-1, as demonstrated by its absence in DHBV. These findings warrant further development of 8-1 for the treatment of chronic hepatitis B and its associated diseases.
Introduction
Helioxanthin, a natural product isolated from the plant Taiwania cryptomerioides, exhibits broad-spectrum antiviral activity, including activity against HBV [1–4]. In order to improve the selectivity and potency against HBV, a series of helioxanthin analogues were designed and synthesized. Among them, 8-1 (Figure 1) displayed the best selective index as studied in a human HBV cell culture system [1,4]. In both wild-type and lamivudine (3TC)-resistant human HBV-producing cell lines, 8-1 effectively suppressed the production of HBV macromolecules, including DNA, RNA and proteins. Furthermore, 8-1 displayed a greater cytotoxic effect in HBV-harbouring cells than virus-free parental cells. The compound post-transcriptionally down-regulates hepatocyte nuclear factor-3 and −4, which are required for HBV RNA transcription [5]. This down-regulation is preferential in virus-harbouring cells over virus-naive cells; a finding that suggests the role that HBV plays in the action of 8-1. The HBV X protein is a non-structural viral protein that is used by the virus as a regulatory protein to modulate host and virus gene expression. Therefore, HBV X protein might be responsible for the interaction with 8-1, described by the aforementioned pathways. This possibility led us to use duck hepatitis B virus (DHBV) to study the antiviral activity of 8-1. DHBV has no X protein expression in vivo, but an unusual cryptic open reading frame exists in the genome [6,7]. Studies on the antiviral activity of 8-1 in DHBV could reveal the role of X protein through the action of 8-1 in the human HBV system. The DHBV inducible cell line dstet5 was, therefore, chosen for the examination of the antiviral activity of 8-1. This cell line supports efficient replication of DHBV in a conditional and synchronized fashion [8]. Following induction via the removal of doxycycline from cell culture medium, dstet5 cells produce DHBV DNA, RNA and proteins in a time-dependent manner. Moreover, the amplification of covalently closed circular DNA (cccDNA) in this cell line is particularly efficient because of its deficient expression of DHBV large surface protein. The DHBV large surface protein appears to suppress cccDNA amplification and deletion, and subtle substitution mutations of it increase the cccDNA copy numbers [9–11]. This cell line can, thus, also be used to study cccDNA formation and compounds antagonistic to cccDNA synthesis.
Structures of the helioxanthin analogue 8-1 and nucleoside analogue 3TC
Methods
Compounds
The helioxanthin analogue 8-1 was synthesized according to a previously published method [4]. 3TC was a gift from Triangle Pharmaceuticals (Durham, NC, USA).
Cell culture
The dstet5 cells were cultured on collagen-1-coated flasks (Becton Dickinson Laboratory, Bedford, MA, USA) at 37°C in a humidified 5% CO2/air atmosphere in Dulbecco's modified Eagle's medium/F12 supplemented with 10% (v/v) fetal bovine serum, 50 μg/ml kanamycin, 200 μg/ml G418 and 1 μg/ml of doxycycline (Sigma–Aldrich, St Louis, MO, USA). Upon confluence, cells were subcultured using pancreatin (0.5 mg/ml) followed by medium change every other day.
Determination of the replication of DHBV in dstet5 cells
A quantity of 5×105 dstet5 cells were seeded on collagen-1-coated 60 mm dishes (Becton Dickinson Laboratory) with the culture medium mentioned in Cell culture. On day 3, cells were carefully washed 3 times with phosphate-buffered saline (PBS), and replaced with doxycycline-free medium to induce DHBV expression. Tested compounds were added to the cell cultures at the same time as doxycycline removal. Medium was changed on days 3 and 6. Cells were harvested for Southern, northern or western blot analysis of viral DNA, cccDNA, RNA or protein. For Southern blot, monolayer cells were collected by detaching cells with pancreatin. Total DNA was isolated using DNeasy tissue kits (Qiagen Inc., Valencia, CA, USA) following the manufacturer instructions. DNA concentration was measured by NanoVue™ UV/Visible spectrophotometers (GE Healthcare, Piscataway, NJ, USA). An aliquot of 20 μg of DNA was resolved by electrophoresis with 1% agarose and transferred to a Hybond-N+ membrane (Amersham Biosciences, Amersham, Buckinghamshire, UK) using the capillary method with 20x saline sodium citrate (SSC) overnight. After ultravoilet cross-linking, the blot was pre-hybridized with 1 mg/ml salmon sperm DNA diluted in ExpressHyb Hybridization Solution (Clontech Laboratories, Inc., Mountain View, CA, USA) for 1 h at 65°C. Random [32P]-labelled DHBV full-length genome cut from plasmid pSP56-DHBV was added to the prehybridized reagent and the blot was hybridized overnight at 65°C. The blot was washed with 2×SSC, 0.1% SDS for 2×5 min at room temperature and 0.2×SSC, 0.1xSDS for 3×20 min at 65°C. It was then exposed to Kodak BioMax MR Film (Kodak, Rochester, NY, USA) at −70°C overnight or detected by Phosphorimager S1 (Molecular Dynamics, Sunnyvale, CA, USA). For northern blot, total RNA from harvested cells was isolated using RNeasy kits (Qiagen, Inc.) following the manufacturer instructions. After measuring the RNA concentration using the NanoVue™ UV/Visible spectrophotometer, 20 μg of total RNA was separated by electrophoresis on 1% agarose with 5% formaldehyde (Sigma–Aldrich) and 1xMOPS (American Bioanalytical, Natick, MA, USA). The RNAs were transferred to a Hybond-N+ membrane (Amersham Biosciences) as described for DNA preparation; the pre-hybridization, hybridization, washing as well as exposure were similar to the Southern blot analysis. For western blot, cells from one dish were lysed using 200 μl lysis buffer (150 mM NaCl, 50 mM Tris HCl, 5 mM EDTA, 4% SDS and proteinase inhibitors; Roche Diagonostics, Indianapolis, IN, USA) and sonicated. Protein concentration was measured using BioRad protein assay dye reagent concentrate (BioRad Laboratories, Inc. Hercules, CA, USA). Equal amounts of total protein (10 μg) from samples were mixed with fivefold protein loading buffer (250 mM Tris HCl, pH 6.8, 0.5 M DTT, 10% SDS, 0.5% triphone blue and 50% glycerol with the addition of proteinase inhibitors) and run on 10% SDS-PAGE. The proteins were then transferred to a nitrocellulose membrane (BioRad Laboratories, Inc.) with transfer buffer (20% methanol, 39mM glycine, 48 mM Tris and 0.037% SDS) using a trans-blot SD semi-dry transfer cell (Bio-Rad Laboratories) at 15 V for 20 min at room temperature. The membrane was blocked by 5% non-fat milk in PBS containing 0.2% Tween-20 (Sigma–Aldrich) for 1 h and probed by anti-DHBV core antigen antibody at 4°C overnight. After washing for 3×20 min using PBS with 0.2% Tween-20, the membrane was incubated with secondary antibody for 1 h at room temperature. After washing in the same manner as above, the blot was subjected to chemiluminescent detection following the routine procedure.
To determine the 50% inhibitory concentration (IC50), blots were scanned using a densitometer (Personal Densitometer SI; GE Healthcare) to measure the optical density values. The values of drug treatment samples were compared with that of mock treatment to obtain the percentage ratio. Next, an IC50/50% cytotoxicity concentration (CC50) calculation module based on linear regression was used to generate IC50 concentrations.
To study the stability of viral macromolecules (that is, DNA, RNA, core protein and cccDNA) under drug treatment, cells were induced for 6 days. A total of 2 μg/ml of doxycycline was then added to the cultures together with tested compounds for another 2 days. Respective viral components were then isolated and determined as above, or as described in DHBV cccDNA isolation and determination, and compared with mock treatment.
DHBV cccDNA isolation and determination
DHBV cccDNA was isolated using the Hirt [12] method with some modifications. In brief, the dstet5 cell pellet from one dish was lysed with 1 ml of lysis buffer (50 mM Tris HCl, pH 8, 10 mM EDTA, 150 mM NaCl and 1% SDS). An aliquot of 250 μl of 2.5 M KCl was added to reach a final concentration of 0.5 M. The cellular lysates were gently rotated at 4°C overnight before undergoing centrifugation at 13,000 rpm for 15 min at 4°C to precipitate the chromosomal DNA. Supernatant was collected and sequentially extracted with phenol, phenol/chloroform and chloroform. After it was precipitated by 2.5-fold of supernatant volume of ethanol with 1% NaAC in −20°C for 1 h and pelleting by centrifugation for 20 min at 13,000 rpm at 4°C, cccDNA was reconstituted in 50 μl H2O and separated by 1% agarose. Southern blot analysis was performed as previously described.
Combination index calculation
The combination index (CI) was calculated using the following equation, based on the median-effect principle of the mass–action law proposed by Chou and Rideout [13]: CI=(D1)/(Dm)1+(D2)/(Dm)2+(D1)(D2)/(Dm)1(Dm)2, where Dm is the 50% effective concentration (EC50) of the drug used alone, and D1 and D2 are the respective doses of drug 1 and 2 in combination that also inhibited 50% of virus (that is, their isoeffective dose).
Cytotoxic assays
The dstet5 cells were seeded in collagen-1-coated 24-well culture plates (Becton Dickinson Laboratory) at a density of 3×104 cells/well with or without doxycycline for 3 days. The culture medium was removed and replaced with 1 ml of the same medium with or without compounds for another 3 days. A volume of 200 μl of 5 mg/ml MTT was added to each well and the cells were further incubated for 4 h. After removal of medium, 400 μl of dimethyl sulfoxide (Sigma–Aldrich) were added to the cultures and the plates were rotated at room temperature for 5 min. The plates were analysed by a plate reader (Elx 800 UV Universal Microplate Reader; Bio-Tek Instruments, Inc., Winooski, VT, USA) under a 595 nm wavelength. The CC50 values were calculated as described above for the IC50 calculation, by comparing the values of drug treatment with those of the mock treatment samples.
Results
The helioxanthin analogue 8-1 inhibits DHBV replication in viral inducible dstet5 cells
The inhibitory activity of 8-1 against DHBV replication was examined using the DHBV inducible cell line dstet5. 3TC was used as the positive control for comparison of anti-HBV activity with 8-1. These compounds were added to cell cultures at the same time, when DHBV was induced by removing doxycycline from the medium; treatment was continued for 6 days. The synthesis of viral macromolecules, including DNA, cccDNA, RNA and protein, were then measured as described in Methods. Figure 2A shows that 8-1 inhibited DHBV DNA synthesis in the dstet5 cells. The mean ±SD IC50 value of 8-1 against viral DNA was 0.25 ±0.05 μM. Figure 2B shows that 8-1 exhibited a potent suppressive effect on viral 2.1/1.8 kb DHBV RNA species. The mean ±SD IC50 was 0.1 ±0.02 μM, indicating better inhibitory activity than that against viral DNA. In addition to its inhibitory activity on DHBV DNA and RNA, 8-1 also suppressed DHBV protein production potently, as exemplified by the inhibition of DHBV core antigen expression (Figure 2C). The IC50 against core protein was <0.3 μM and reached a mean ±SD of 0.1 ±0.1 μM in this study. After removal of doxycycline from dstet5 culture medium, the cells began to produce and accumulate cccDNA inside the nucleus. The activity of 8-1 against cccDNA production was also examined in this study and compared with 3TC. As shown in Figure 2D, cccDNA was dose-dependently inhibited by 8-1 and 3TC, but 3TC exhibited a much stronger inhibitory effect than 8-1 on cccDNA.
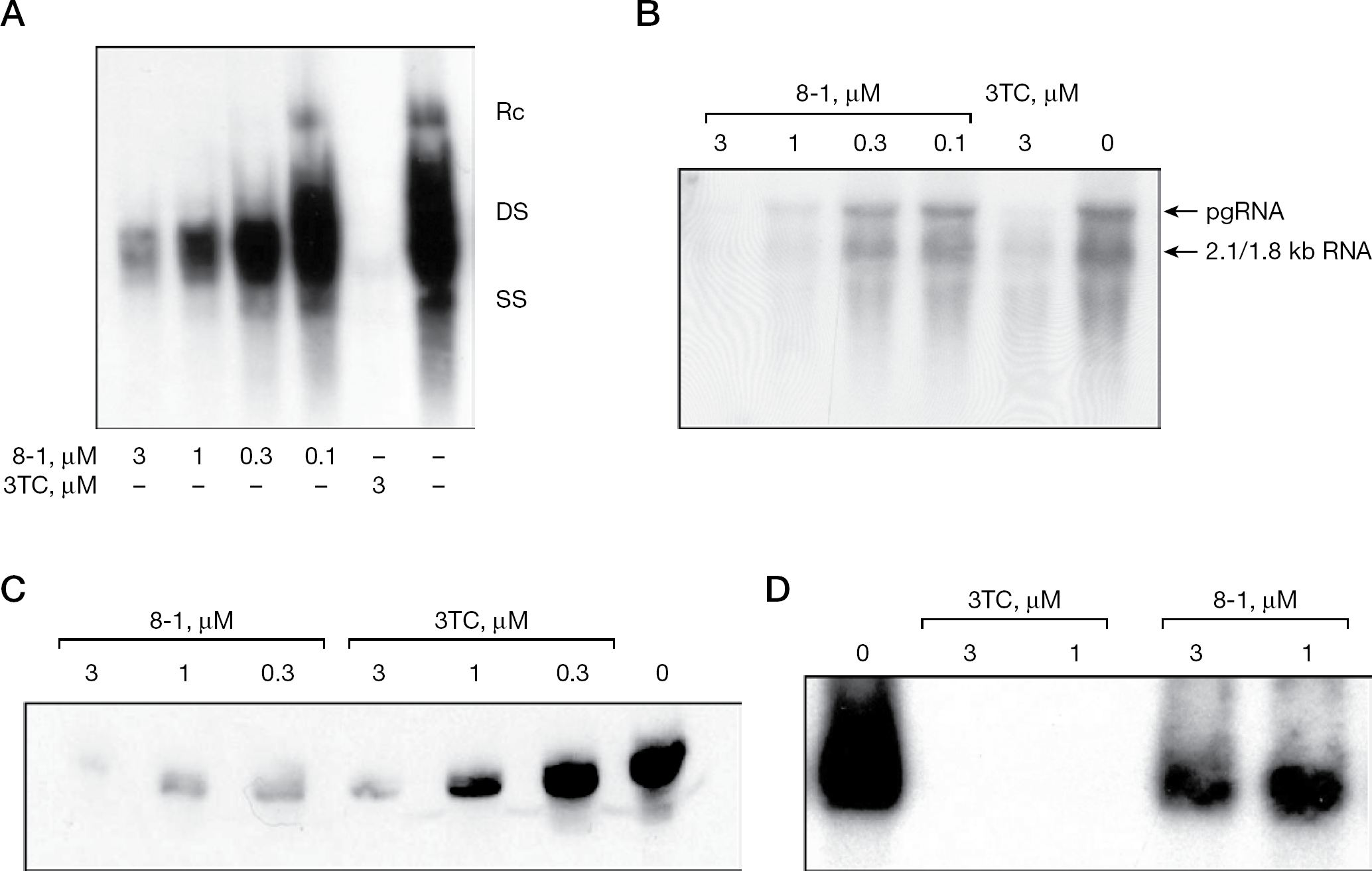
Study of 8-1 on the inhibition of DHBV replication
In order to examine whether the inhibition of viral DNA, RNA, protein and cccDNA by 8-1 resulted from the instability of these viral macromolecules – specifically, by enhanced degradation during drug incubation – cells were induced for 6 days to allow macromolecules to be highly produced. Doxycycline was added to the cultures again to block de novo production of these viral components. The accumulated macromolecules were then subjected to time-dependent degradation. An aliquot of 1 μM of 8-1 was added to the cultures for 2 days to examine whether the compound enhanced their degradation as compared with mock treatment. The result of this procedure indicated that 8-1 had no effect on DNA, RNA, protein or cccDNA stability; that is, their degradation was not affected by the presence of this compound (CY et al., data not shown).
It was previously observed that an analogue of 8-1, namely 8-1/T, exhibited long-lasting antiviral activity in our recent animal study including HBV transgenic mice. The mice were orally fed 8-1/T in 20 mg/kg twice daily for 6 days, then left for another 6 days without treatment. Viral serum DNA titres on days 6 and 12 were examined by a specific real-time PCR method and it was found at both time points that viral DNA titres were still suppressed in the 8-1/T-treated groups without viral rebound back to the original level. In that study, the control compound, 3TC, did not exhibit long-lasting antiviral activity, even with a dosage as high as 200 mg/kg following the same schedule as that of 8-1/T (CY et al., unpublished data). We therefore wanted to determine if this long-lasting effect also exists in the cell culture system. Thus, a DHBV DNA rebound experiment after cessation of 8-1 treatment was performed in dstet5 cells. Cells were treated with 1 μM 8-1 for 6 days after induction. The drug-containing medium was then removed and the cultures were carefully washed 3 times with PBS. Drug-free medium was given to cells for another 3 days before the viral DNA was examined. The 8-1 did not exhibit sustained antiviral activity in this study with this particular cell line. The viral DNA completely rebounded 3 days post-treatment (CY et al., data not shown); therefore, 8-1 does not have sustained anti-DHBV activity in the cell culture system.
Combination study of 8-1 with 3TC in inhibiting DHBV replication
The combinational antiviral activity of 8-1 with 3TC was studied in terms of its viral DNA synthesis. This experiment was designed following the median-effect principle proposed by Chou et al. [13–15] and the antiviral effect analysis was determined using the CI calculation method. Table 1 shows the doses used for the combination study and their effect in the inhibition of DHBV DNA replication. For example, when 8-1 0.1 μM and 3TC 0.03 μM were used alone, <40% of inhibition was detected; however, when 8-1 was combined with 3TC in the aforementioned concentrations, enhanced antiviral activity was observed. Approximately 50% of viral DNA was inhibited as compared with the viral control. The P-value was <0.05 in this combination.
Combination dose design for 8-1 plus 3TC anti-DHBV study a
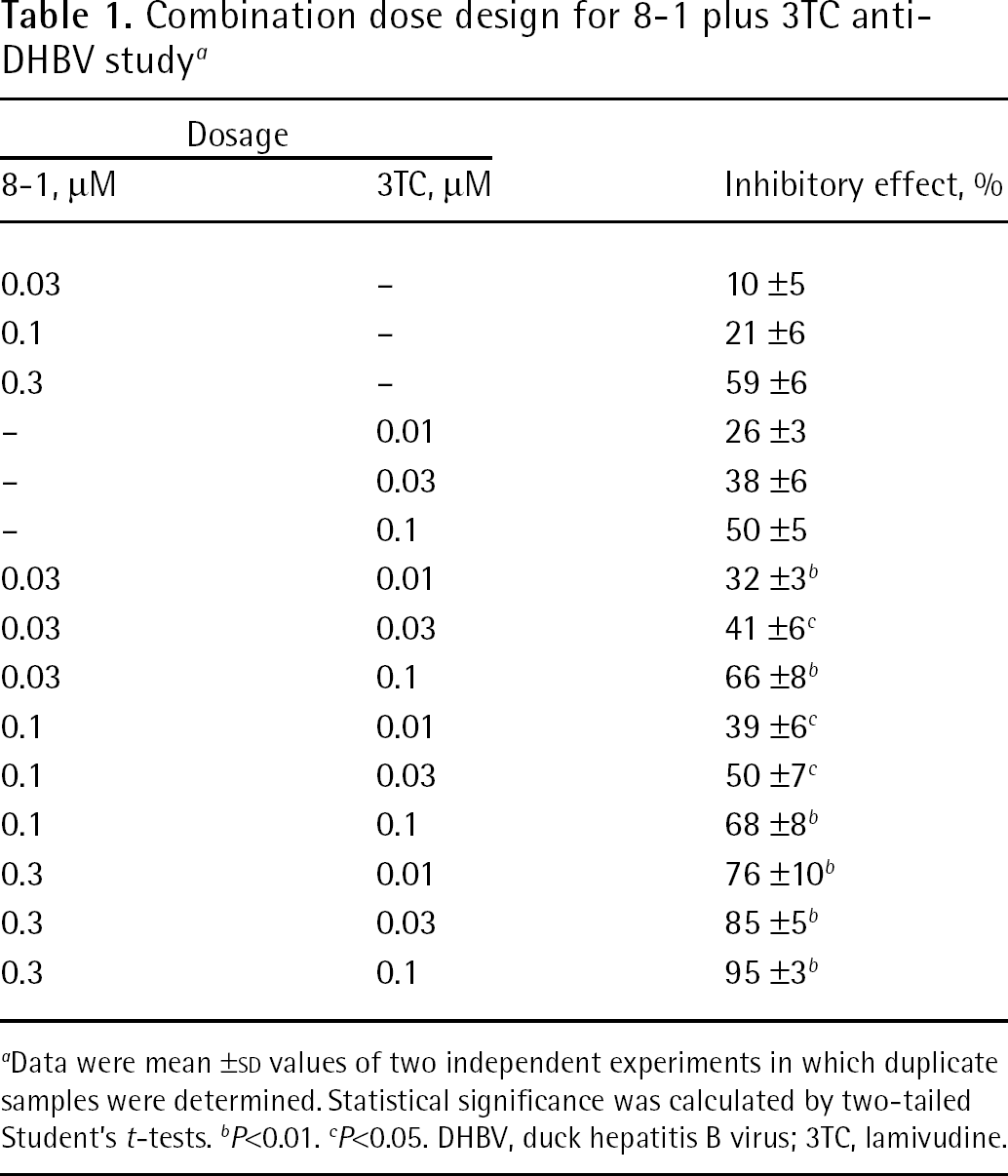
Data were mean ±SD values of two independent experiments in which duplicate samples were determined. Statistical significance was calculated by two-tailed Student's t-tests.
P<0.01.
P<0.05. DHBV, duck hepatitis B virus; 3TC, lamivudine.
Given that the EC50 of 8-1 was approximately 0.25 μM and that of 3TC was approximately 0.1 μM in the present cell culture study, CI was calculated with the equation indicated in Methods and using the values from the Table 1 as: CI=0.1/0.25+0.03/0.1+(0.1×0.03)/(0.25×0.1)=0.82
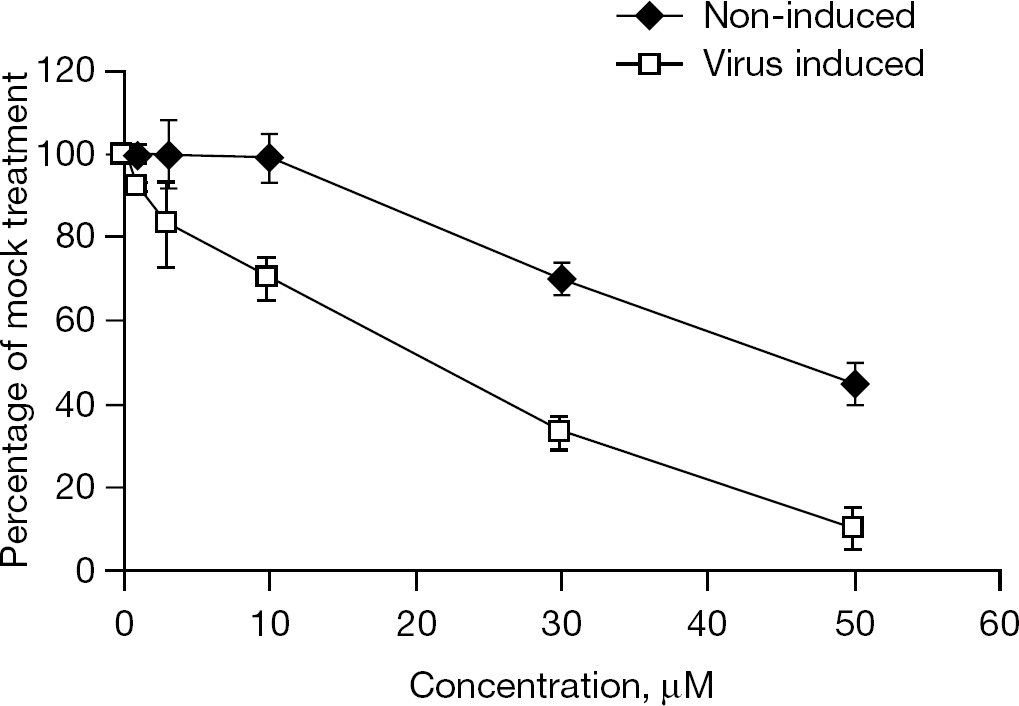
Study of 8-1 cytotoxicity
According to the theory of MEP, CI=1, CI<1 and CI>1 indicate additive, synergistic and antagonistic drug interactions, respectively; therefore, 8-1 and 3TC have the potential for causing a synergistic effect on the inhibition of DHBV that could possibly be extended to HBV replication. However, because the difference between 0.82 and 1.0 did not achieve statistical significance, 8-1 and 3TC were additive in this combination study.
Cytotoxicity study of 8-1 in induced and non-induced dstet5 cells
The cytotoxic effect of 8-1 and 3TC was determined with or without viral induction. An MTT assay was employed to examine cellular cytotoxicity. As seen in Figure 3, 8-1 exhibited an apparent cytotoxic effect in induced cells and displayed a smaller effect in non-induced cells (that is, no virus production). The mean ±SD CC50 of 8-1 was 18 ±2 μM in viral-induced cells and 45 ±3 μM in viral non-induced cells. 3TC did not exhibit any cytotoxic effect on either the viral induced and non-induced cells under the experimental conditions with a concentration as high as 200 μM (CC50>200 μM; CY et al., data not shown).
Discussion
We demonstrated that 8-1, a helioxanthin derivative, inhibited DHBV replication in cell culture using the DHBV inducible cell line dstet5. The analogue 8-1 effectively suppresses DHBV DNA, cccDNA, RNA and viral protein production. In inhibiting DHBV DNA and cccDNA synthesis, 3TC is more potent than 8-1; however, by contrast, 8-1 is more effective than 3TC at inhibiting DHBV RNA and protein synthesis. This finding indicates that the two types of compounds have different targets and mechanisms, and corroborates our former observations. According to our previous 8-1 mechanism study [5], 8-1 targets the viral RNA transcription step by decreasing viral RNA transcripts, thus rendering viral protein translation less effective, and further extending this effect to viral DNA and cccDNA synthesis. 3TC acts as a DNA chain terminator and/or inhibitor targeted at the level of viral polymerase [16], and suppresses viral DNA synthesis in both RNA reverse transcription and DNA polymerization steps (Figure 4). It was reported that during cccDNA formation, viral polymerase was required for the viral relaxed circular genome repair and gap filling [17,18]; therefore, 3TC also works at the level of viral polymerase to inhibit cccDNA formation directly. By comparison, 8-1 acts on cccDNA through an indirect step, namely RNA transcription after cccDNA formation (Figure 4). This might partially explain why 3TC is more potent in suppression of cccDNA than 8-1.
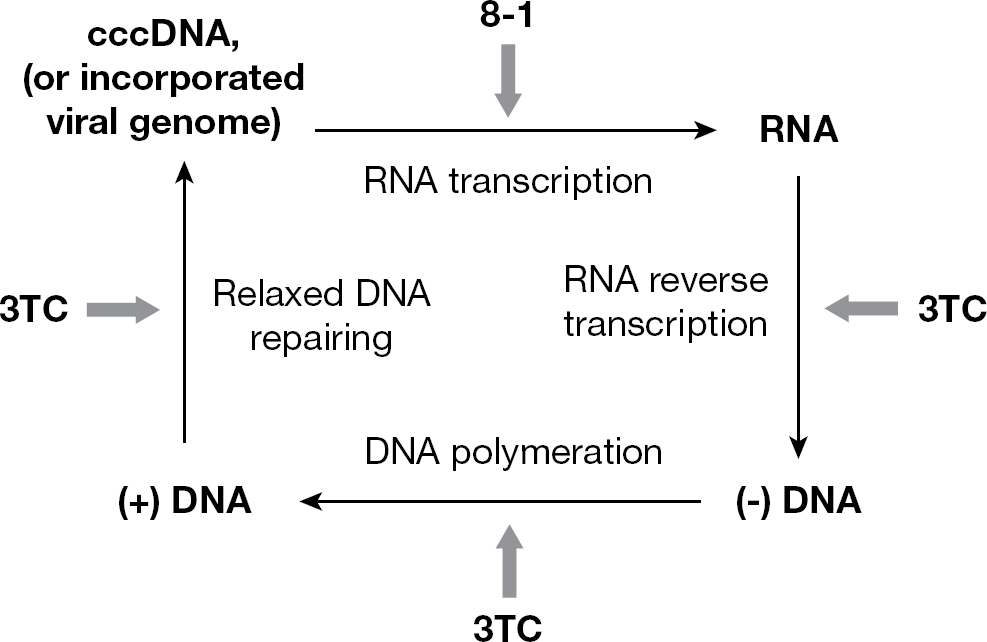
Comparison of the action of 8-1 with 3TC
Suppression of viral macromolecules could result from enhanced degradation rather than inhibition of synthesis. We explored the inducible features of the cells to study the stability of viral macromolecules under the treatment of 8-1. The degradation was not affected by the presence of 8-1 as compared with mock treatment controls. It was, therefore, concluded that the suppression of viral replication is a result of a decrease in synthesis of viral macromolecules rather than enhanced degradation by 8-1.
In this cell culture study, we did not observe any sustained antiviral activity of 8-1 that was seen in an animal study that included HBV transgenic mice treated with an 8-1 analogue, 8-1/T, which has three-fold less anti-HBV activity, as determined by viral DNA. Viral DNA rebounded after the cessation of the treatment. This implies that the sustained antiviral effect of 8-1/T might be related to factors such as immunomodulator(s) that are present in vivo, and which do not exist in the cell culture system.
In the present study, the combinational antiviral activity of 8-1 plus 3TC was examined using the median-effect principle proposed by Chou et al. [13–15]. Because 8-1 and 3TC are different drug entities – that is, mutually non-exclusive inhibitors with completely different or independent modes of action – the CI method for drug combination studies was employed for the experimental design and CI calculation. CI=1, CI<1 and CI>1 indicate additive, synergistic and antagonistic interactions of two drugs, respectively. The calculated CI value of 0.82 for 8-1 combined with 3TC demonstrated that these two mutually non-exclusive antiviral chemicals with different targets and mechanisms have a potential synergistic effect on the inhibition of DHBV replication. However, because the value 0.82 is close to 1 and 8-1 and 3TC have various mechanisms and likely do not interfere with each others action, it would be more conservative to conclude that 8-1 plus 3TC have additive antiviral activities. This result, consistent with our previous observation in human HBV where 8-1 was active against 3TC-resistant HBV [5], suggests that 8-1 and 3TC could possibly be explored in combination for enhanced suppression of HBV infections. This combination regime not only could reduce the dosage of each compound required thereby minimizing their potential cytotoxic effect, but could also restrain resistant virus emergence.
We leveraged the inducible feature of DHBV titre in dstet5 cells to study the cytotoxic effect of 8-1 in the presence and absence of viral production. The data clearly indicated that viral replication made the host cells more prone to the cytotoxic effects of 8-1. This is consistent with our previous observation in human HBV HepG2 2.2.15 and its parental HepG2 cells, and it was concluded that the presence of certain viral components contributes to the preferential cytotoxic action of 8-1 against HBV-harbouring cells.
DHBV has three open reading frames and encodes pol, C and S proteins. By contrast, the X protein, as encoded by HBV, is not expressed by DHBV. In this study, 8-1 maintained potent antiviral activity in DHBV and exerted a greater cytotoxic effect in virally induced cells. This suggests that the X protein is not an essential viral component for the initiation of the biological activity of 8-1, such as the antiviral ability and cytotoxicity observed in the human HBV system. Other viral factors, alone or in combination with X protein, might play an important role in the action of 8-1.
In summary, we have demonstrated that 8-1, a natural product analogue, effectively suppresses the production of DHBV DNA, cccDNA, RNA and protein production in an inducible DHBV replication cell line. Cells in which there was active viral replication were more susceptible to the toxic effect of the compound than other cells, suggesting that viral components play an important role in the initiation of the biological activity of 8-1; however, X protein might not be an essential factor, as demonstrated by its absence in DHBV. When combined with 3TC, 8-1 exhibits an enhanced inhibitory effect on DHBV replication. This underscores the potential of 8-1 for HBV replication treatment in combination with 3TC or other nucleuside/nucleotide analogue inhibitors in future development.
Footnotes
Acknowledgements
This work was supported by National Institutes of Health Grant R01 AI73299 (to Y-CC). Y-CC is a Fellow of the National Foundation for Cancer Research. We give our heartfelt gratitude to C Seegers and W Mason of the Fox Chase Cancer Research Institute (Philadelphia, PA, USA) for providing the dstet5 cells, DHBV full-length genome plasmid pSP56-DHBV as well as DHBV core antibody. We thank Michel Robek, Jessica Qu, Scott Bussom and Chuan-Jen Wang for very helpful discussion during this work and careful reading of the manuscript.
The authors declare no competing interests.
