Abstract
Background:
We have shown that the K65R resistance mutation in HIV type-1 (HIV-1) reverse transcriptase (RT) is selected more rapidly in subtype C than subtype B HIV-1 in biochemical, cell culture and clinical studies. Template-usage experiments demonstrated that subtype C nucleotide coding sequences caused RT to preferentially pause, leading to K65R acquisition. This new study now further establishes the basis for differential occurrence of both K65R and thymidine analogue mutations (TAMs) between subtypes.
Methods:
Gel-based nucleotide extension assays were used to study the homopolymeric sequence surrounding K65.
Results:
When positive double-stranded DNA synthesis was evaluated from a negative single-stranded DNA template, pausing at the 67 region, which is linked to occurrence of TAMs, was alleviated with both subtype B and C templates at high dCTP concentrations, but this alleviation was more pronounced with the subtype C template. By contrast, pausing at the 65 region on the subtype C but not subtype B template always occurred and was not alleviated at high levels of nucleotide triphosphates or by other means. Furthermore, templates containing repeats of the homopolymeric sequence spanning codons 64–66 of pol showed corresponding pausing repeats at the 65 region with the subtype C template only. Inverted RNA and DNA templates both displayed pausing at position K65 for the subtype C template and a ladder of pausing events culminating at codon 67 for the subtype B templates.
Conclusions:
These results further establish a mechanistic basis for the exclusion of both K65R and TAMs on single templates as well as the preferential acquisition of K65R in subtype C viruses.
Introduction
HIV type-1 (HIV-1) is thought to have been introduced into the human population as early as 80 years ago [1] and today approximately 40 million people are infected with the virus [2]. The virus has also evolved into three distinct groups: M, N and O, the largest of which is group M that is further subdivided into distinct subtypes: A1, A2, A3, B, C, D, F1, F2, G, H, J, K and N, as well as many circulating recombinant forms [3]. HIV-1 and HIV type-2, a related virus, share approximately 50% sequence homology. The M, N and O groups, which are the main HIV-1 groups, share up to 70% sequence homology. Furthermore, each subtype subdivision of the individual HIV-1 groups can possess up to 90% homology with other corresponding subtypes [4–6]. The three most prevalent forms of the virus are subtype A1 (representing approximately 12% of the infected population globally and found mainly in north Asian countries), subtype B (representing approximately 10% of infections worldwide and pre dominating in North America, South America, Europe and Australia), and subtype C (causing >50% of all infections in the world and found mostly in sub-Saharan Africa and India) [7]. The different subtypes differ on the basis of their nucleotide sequences by 10–12% and by 5–6% in terms of amino acid sequence [8]. Understandably, most HIV-1 research has focused upon subtype B, seeing as it is the most common circulating form in high-income countries in which the research has been conducted and in which antiretroviral drugs have been available [2,9].
Antiretroviral therapy has successfully reduced HIV-1-related mortality [10–12], and responsiveness to therapy has been noted regardless of viral subtype. However, all antiretroviral drugs to date have been prone to the emergence of drug resistance that can lead to treatment failure. Although resistance mutations have been well characterized, the vast majority of such studies have been limited to subtype B HIV-1 [13–15].
Recently, our laboratory reported that HIV-1 subtype C develops the K65R resistance mutation in reverse transcriptase (RT) more quickly than subtype B viruses in cell culture [16]. Clinical data also reveal a higher prevalence of K65R in subtype-C-infected patients in Botswana, Malawi and South Africa [17–19]. K65R, even if present as a minority species in a drug-naive patient, is outcompeted by wild-type viruses. The fitness constraints of the K65R mutation are too great for it to exist on its own without selective drug pressure.
K65R confers broad high-level resistance against most nucleoside/nucleotide reverse transcriptase inhibitors with the exception of zidovudine [20–23]. In addition, it has been well established that HIV-1 RT experiences difficulty synthesizing homopolymeric nucleotide stretches, resulting in recombination, template–primer slippage, misincorporation and strand transfer events that can lead to the development of mutations [24–27]. To that effect, we have shown that the preferential development of K65R in subtype C is based on the nucleotide polymorphisms that exist between subtype B and C template sequences (Figure 1) [28]. Specific differences in these sequences make subtype C viruses more prone to RT pausing, which we hypothesize as being the event that allows for a correct nucleotide incorporation to occur but into a misaligned template primer, which is subsequently realigned at the specific nucleotide position, thereby causing the misincorporation responsible for the K65R mutation [28]. A correct nucleotide can thereby become incorporated into a misaligned template–primer sequence and is likely to give rise to the AAG to AGG transition of K65R in the context of subtype C HIV-1 [28].
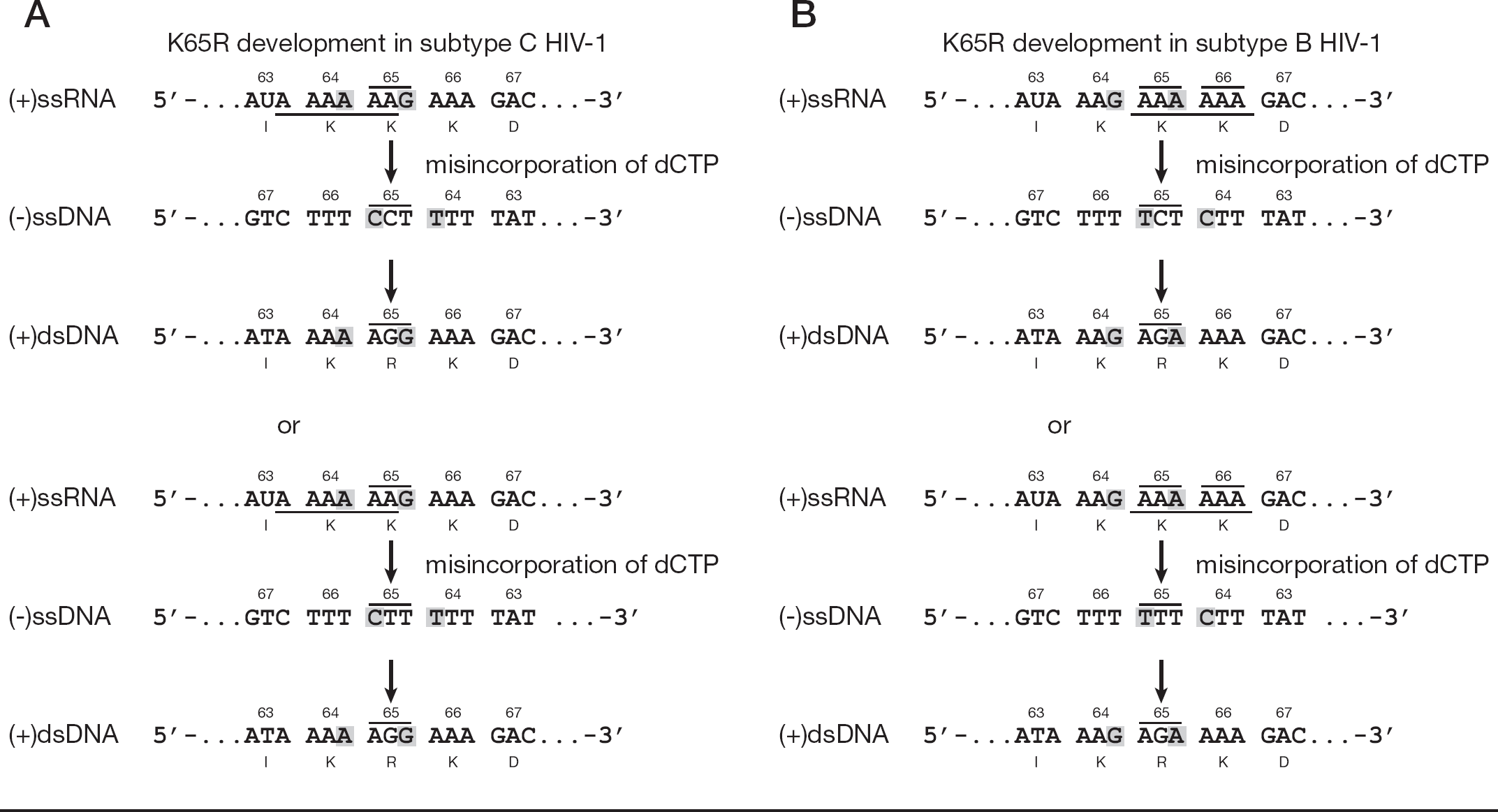
Comparison of genomic sequences and of K65R development in subtype B and C reverse transcriptase
The RT enzyme is characterized by its ability to synthesize negative (−) single-stranded (ss)DNA from the virally packaged genomic RNA positive (+)ssRNA, and to then produce a transcript that becomes integrated into the host genome (+) double-stranded (ds)DNA. We have shown that there is little enzymatic slippage or pausing in the K65 region of the pol gene of HIV-1 during (−)ssDNA synthesis in the case of both subtype B and C viruses. However, when (+)dsDNA synthesis was studied using (−)ssDNA templates, subtype-specific patterns of DNA synthesis were observed [28]. In brief, the use of subtype B DNA with either subtype B or C RT to synthesize the (+)dsDNA yielded a ladder of pausing events spanning codons 65, 66 and 67. By contrast, when subtype C DNA was used with either subtype B or C RT to synthesize (+)dsDNA, DNA synthesis stopped at the exact nucleotide position responsible for the K65R mutation, potentially resulting in higher mutational rates and in faster K65R development. Because the subtype C sequence is not present in the subtype B template, K65R resistance is thought to be delayed and, in most cases, even entirely avoided in the latter case [28]. We have now investigated more closely the role of the above-mentioned sequence particularities that distinguish subtypes B and C of HIV-1 with respect to factors that might participate in accentuating or alleviating pausing at the K65 position, as determining factors in the development of drug resistance. The factors that we studied include the role of correctly-oriented and inversely-oriented templates spanning the K65 region of pol in the presence of variable dNTP concentrations as well as use of templates containing repeats of the 64, 65 and 66 regions of pol.
Methods
Templates and primers for RT assays
Templates and primers were purchased in desalted purity from Invitrogen (Carlsbad, CA, USA) or Integrated DNA Technologies (Coralville, IA, USA); subtype B wild-type templates and subtype C wild-type templates were derived from the NL4–3 and MJ4 infectious viral clones respectively unless otherwise mentioned [28]. The templates used were specifically designed to study (−)ssDNA synthesis using a (+)ssRNA template as well as (+)dsDNA synthesis using a (−) ssDNA template derived from the K65 region of the pol gene as described previously [28]. All template and primer sequences employed in the experimental protocols are summarized in Table 1. The templates included in these assays are the following: subtype B wild-type (−) ssDNA: 5′-AGT ACT GTC TTT TTT CTT TAT GGC AAA
Subtype B and C HIV type-1 templates and primers used to evaluate DNA synthesis in various experimental protocols

Bold sequences denote primer binding regions. ss, single-stranded; (−), negative; (+), positive.
Templates containing repeats of the 64, 65 and 66 region of pol include the following: subtype B wild-type inverted (+)strand DNA with repeats: 5′-GT TTA AAA AAA GAA AAA AAA GAA AAA AAA GAA
Enzyme preparation and purification
Heterodimeric recombinant subtype B and C (p66/p51) RT enzymes were expressed in Escherichia coli cells and purified as previously reported [29].
DNA synthesis from the DNA and RNA templates
The DNA synthesis reactions were conducted using 100 nM of the 5′-labelled 19D primer that was annealed with 300 nM of template at 95°C for 2 min, 72°C for 20 min and at 37°C for 20 min. The annealed RNA/DNA or DNA/DNA hybrids were incubated with 300 nM of HIV-1 RT in a buffer containing 50 mM Tris-HCl, pH 7.8, 50 mM NaCl, 6 mM MgCl2 and 10 μM each of the dNTPs. When RNA was used, RNase inhibitors were added to the reaction. The reaction was allowed to proceed at 37°C and stopped with the addition of 100 μl of 100% isopropyl alcohol containing 300 mM ammonium acetate and 10 ng/μl bulk yeast RNA into 20 μl of the reaction mix. Measurements were taken at 0, 2, 4, 6, 8, 10, 12 and 15 min. After completing the reaction, the DNA was resuspended in 10 μl of 100% formamide containing bromphenol blue and xylene cyanol. The samples were then resolved on 8% polyacrylamide 7 M urea gels containing 50 mM Tris-borate pH 8 and 1 mM EDTA. Analysis of the results was possible by exposing radioactive reaction products to phosphor screens and phosphorimaging. For the nucleotide competition experiments, different individual dNTP concentrations including 0, 1, 5, 10, 50 and 100 μM were used and reactions were terminated after 15 min with the addition of 100 μl of 100% isopropyl alcohol containing 300 mM ammonium acetate and 10 ng/μl bulk yeast RNA into 20 μl of the reaction mix as described earlier.
Results
(−)ssDNA synthesis from a (+)ssRNA template with variable nucleotide concentrations
To investigate whether specific nucleotides, when present in excess or at lower concentrations, are favoured for incorporation, we synthesized (−)ssDNA from a (+)ssRNA template in the presence of varying amounts of dATP, dCTP, dGTP or dTTP (Figure 2). No pausing was observed in any of the reactions, and increasing the individual nucleotide concentration up to 100 μM did not result in any pausing at codons 63 through 67 (positions +7 to +21). This was true for both subtype B (Figure 2A) and C (Figure 2B) templates. Therefore, an incorrect incorporation event during synthesis of (−)ssDNA from (+)ssRNA that would result in the development of the K65R mutation is not likely to occur.
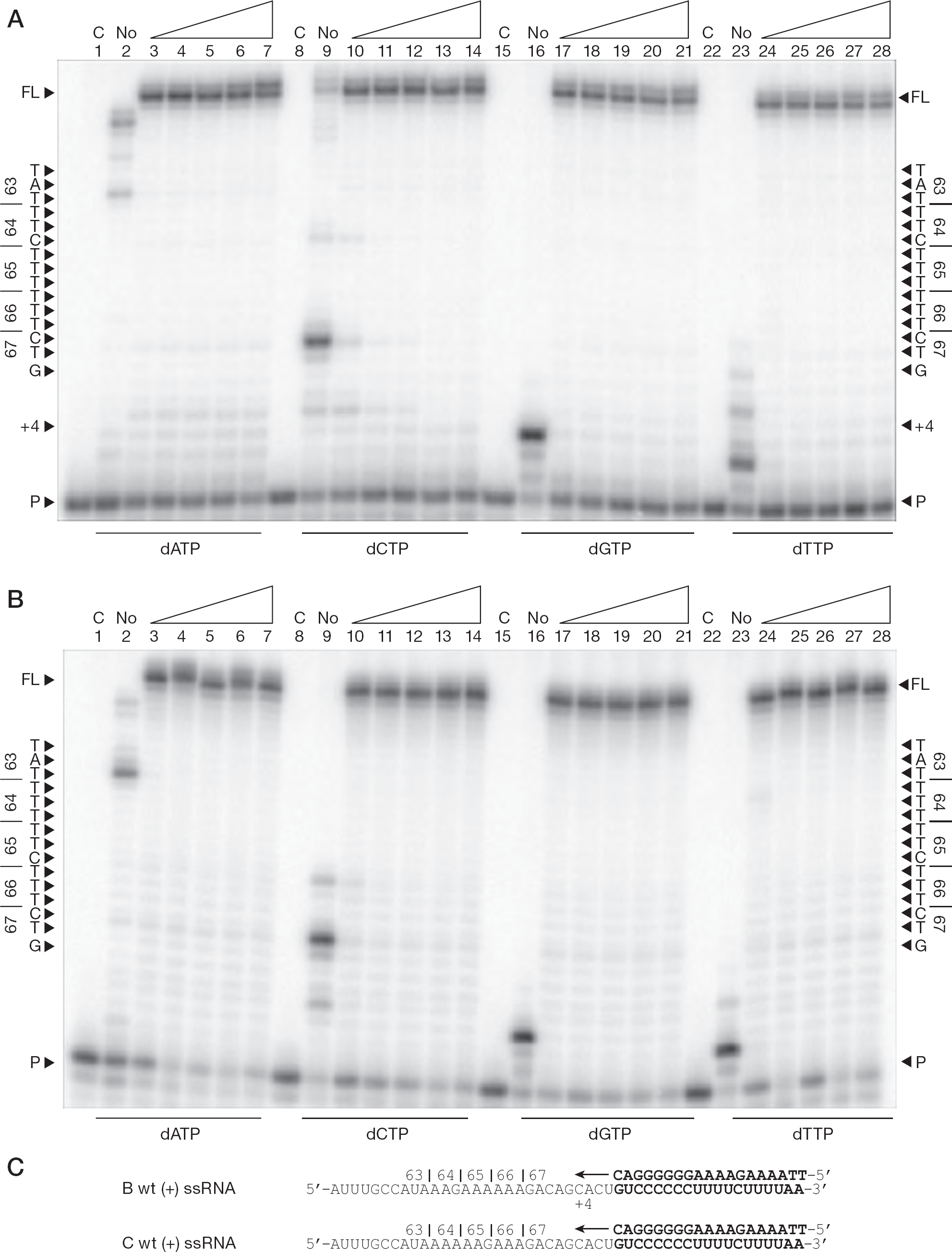
(−)ssDNA synthesis from the 65 region of pol using variable dNTP concentrations
In addition, different rates of incorrect nucleotide incorporation that resulted in residual primer extension occurred in reactions that did not contain any of a particular dNTP (Figure 2A and 2B, lanes 2, 9, 16 and 23). When employing subtype B RT on a subtype B RNA template (Figure 2A, lane 2), one would anticipate that DNA synthesis should have stopped before the +1 position opposite the first UTP of the RNA template sequence. Surprisingly, however, it did not; rather, synthesis continued to almost complete the full-length DNA product. In lane 23 (Figure 2A and 2B), when no dTTP was present in the reaction, an incorrect incorporation event occurred at the +3 position and allowed for DNA synthesis to extend beyond that point. In the total absence of dCTP (Figure 2A, lane 9), DNA synthesis did not stop before the +4 position prior to the first GTP of the template, but continued beyond subsequent GTP positions on the subtype B template (Figure 2C), leading to full-length DNA synthesis in approximately 15% of transcripts despite the absence of dCTP in the reaction.
Similarly, when DNA synthesis was evaluated in the complete absence of dATP on the (+)ssRNA templates (Figure 2A, lane 2), it proceeded beyond the +1 position, which is the location of the first UTP of the template for both subtypes B and C. In the case of subtype B (Figure 2B, lane 2), DNA synthesis extended beyond the +20 position of the second UTP of the template and stopped before a series of three UTPs at the +24 position. By contrast, DNA synthesis on the subtype C template extended beyond the +1 position and stopped strongly at the +19 position before the second UTP. Only a small amount of DNA product was observed before the series of UTPs at position +24. These observations also suggest that RT, in the absence of dATP, had incorrectly incorporated other nucleotides at template positions opposite UTP in order for DNA synthesis to proceed to an almost full-length product.
(+)dsDNA synthesis from a (−)ssDNA template with variable nucleotide concentrations
When (+)dsDNA synthesis was evaluated using the (−) ssDNA template and varying nucleotide concentrations, the characteristic ladder of pausing events was observed with the subtype B template (positions +12 to +20), and pausing was at the exact nucleotide position responsible for the K65R mutation (position +14) as in the case of the subtype C template (Figure 3), consistent with our previously reported findings [28]. Interestingly, for both subtype B and C templates, increasing the dCTP concentration above 5 μM alleviated pausing at the D67 (+20) position at the very end of the ladder and more full-length product was formed (Figure 3A, lane 12; Figure 3B, lane 12) representing a decrease in pausing from approximately 15% to <5% at the highest concentrations of dCTP.
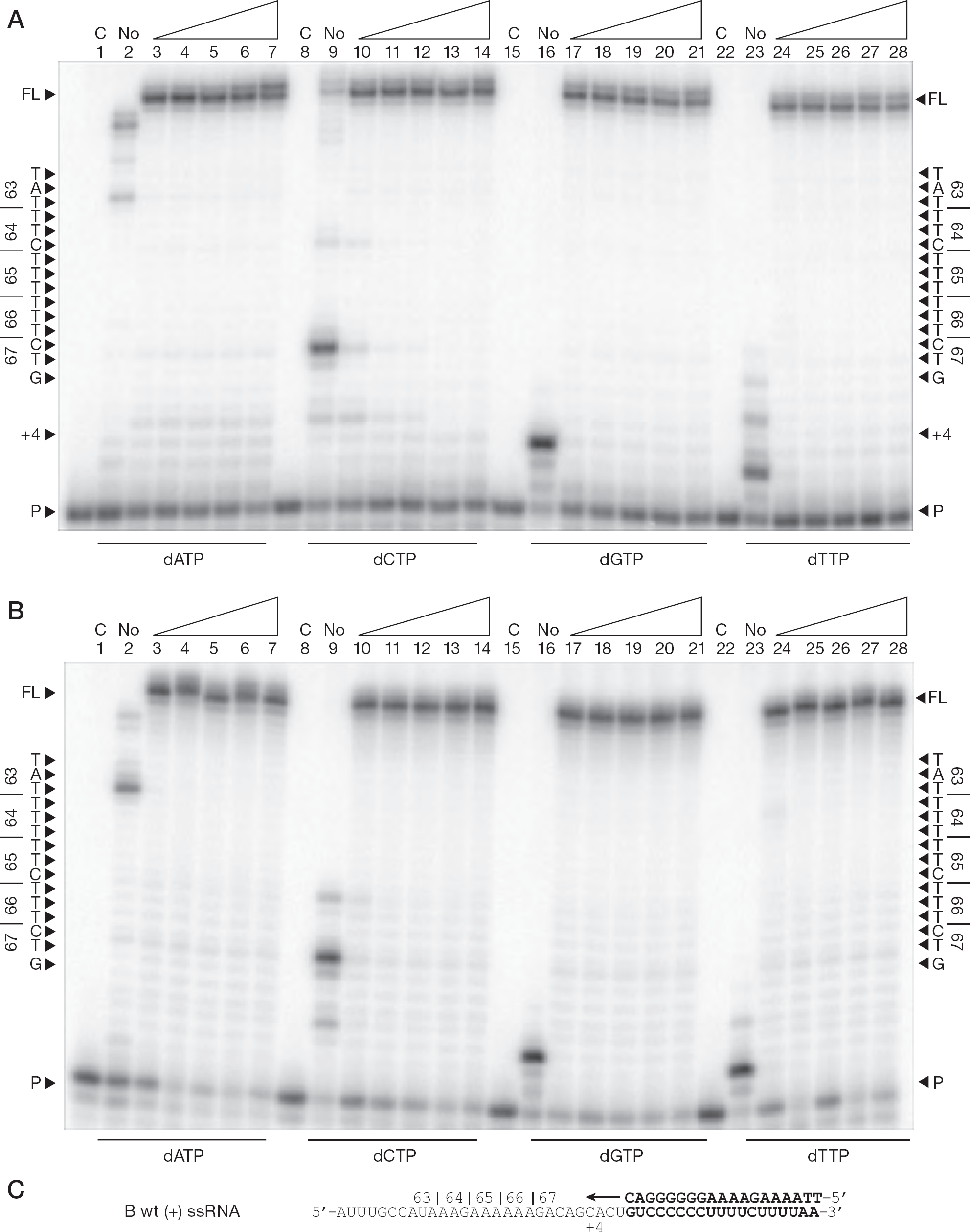
(+)dsDNA synthesis from the 65 region of pol using variable single dNTP concentrations
Interestingly, when (+)dsDNA synthesis was evaluated using the subtype B (−)ssDNA template at the lowest concentration of dATP (Figure 3A, lane 3), increased pausing was seen at the K65 codon, which also corresponds to the beginning of the ladder of pausing events between codons K65 and D67. As soon as the concentration of dATP was increased above 5 μM, this weak pausing disappeared, but the ladder of pausing events culminating at the D67 position remained. This ladder of pausing events is characteristic of DNA synthesis only with subtype B template.
For the subtype C sequence, the major pausing event at the K65 (+14) position remained unchanged with varying concentrations of dCTP (Figure 3B) but, as with the subtype B template, the slight pausing seen at the D67 (+20) position was alleviated with increasing dCTP concentrations (Figure 3A).
Quantification of (−)ssDNA and (+)dsDNA synthesis results
A quantitative analysis of the results shown in Figure 2 and Figure 3, each of which was performed on at least several occasions with similar results, is depicted in Figure 4. In addition to position +4, positions +8, +15 and +23 are the other sites that occur immediately before a GTP on the template at which pausing is observed (Figure 4A). In reactions performed with subtype B wild-type RT and subtype B (+)ssRNA template, the strongest pausing was seen at position +8, suggesting that RT incorporated an incorrect nucleotide at position +4 only to stop in 45% of the transcripts at position +8 just prior to a homopolymer tract of adenines on the subtype B template. Weak wobble base pair formation or nucleotide misincorporation might account for some residual incorporation beyond the point where DNA synthesis should have terminated in the absence of a particular dNTP, but is unlikely to account for intersubtype differences.
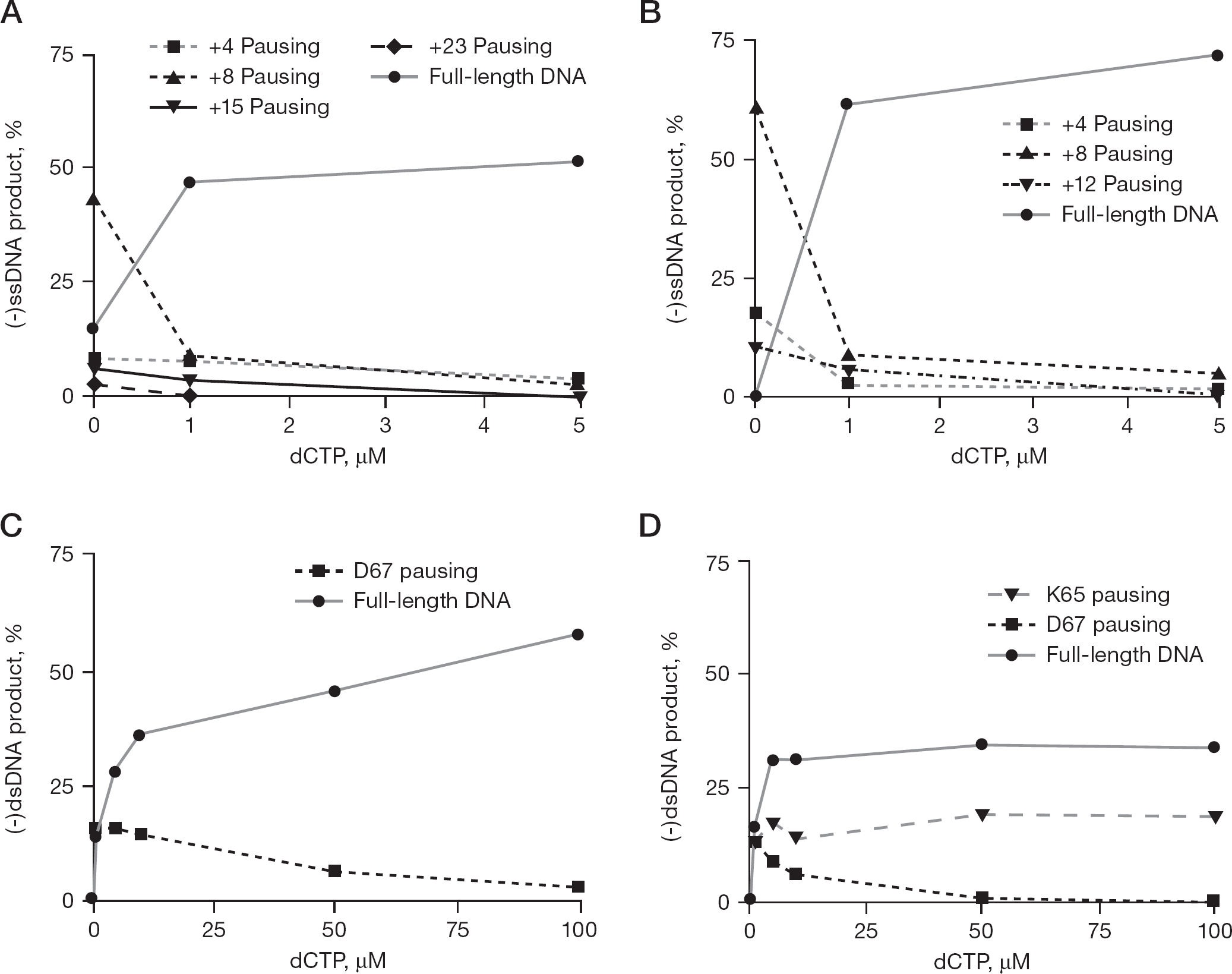
Effect of dCTP variability on pausing in synthesis of (−)ssDNA and (+)dsDNA
Although both subtypes exhibited similar DNA synthesis patterns with variable nucleotide concentrations, full-length DNA products were much more prevalent with subtype B as opposed to subtype C RT. When subtype C RT was used to synthesize (−)ssDNA from a (+)ssRNA template, in the absence of dCTP (Figure 2B), DNA synthesis stopped in 60% of the transcripts at the +8 position immediately before the homopolymer tract of adenines characteristic of the K65 region of the pol gene (Figure 4B). In addition, both subtypes were relieved from pausing at the positions before GTP (+4, +8, +15 and +23 on the subtype B template; Figure 4A; and +4, +8 and +12 on the subtype C template; Figure 4B) when the employed dCTP concentrations exceeded 1 μM. Figure 4A and 4B show dCTP concentrations up to 5 μM because they focus on nucleotide misincorporation events opposite the first G bases on the RNA templates used. Because DNA synthesis stops when no dCTP is included in the reaction, DNA synthesis resumes as soon as dCTP is provided at the lowest concentration. As a result, concentrations >5 μM would not provide insight into the misincorporation events that occur in the complete absence of dCTP. Furthermore, when (−)ssDNA synthesis was assessed using a particular subtype-specific (+)ssRNA template, similar results were obtained regardless of whether subtype B or C RT was used.
An alleviation of pausing at D67 (+20) occurred slower with the subtype B template (50% at 50 μM of dCTP; Figure 4C) than with the subtype C template (90% at 50 μM of dCTP; Figure 4D). With the subtype B template, the full-length DNA product increased by approximately 20%, whereas pausing at the D67 (+20) position decreased by 15% (Figure 4C). With the subtype C template, the full-length DNA product did not increase and remained constant at 32% of the amount of product produced in a wild-type reaction; this was true for all dCTP concentrations used and occurred not-withstanding the fact that increased dCTP concentrations affected the amount of pausing at the D67 (+20) position (Figure 4D). Pausing at the K65 (+14) position with the subtype C template remained constant at about 18%, irrespective of increases in concentrations of any of the nucleotides used (Figure 4D).
Figure 4C and 4D show dCTP concentrations of up to 100 μM because they focus on the pausing events located at positions K65 and D67. Because these positions are in the template after the very first GTP, higher nucleotide concentrations are required to observe the pausing alleviation that occurs at the D67 position, while emphasizing the maintenance of the pausing at the K65 position. The data indicate that pausing at the K65 position on the subtype C template is difficult to alleviate by increasing dNTP concentrations during DNA synthesis. By contrast, pausing at the D67 position is alleviated more readily by increasing dCTP concentrations on the subtype C template, whereas increasing dCTP concentrations on the subtype B template allows for slower pausing alleviation.
It is unlikely that this mechanism operates inside cells because the concentrations of nucleotides are approximately equivalent. Moreover, the use of varying concentrations of MgCl2 altered the observed pausing at varying sites with all the templates used. In addition, when (+) dsDNA synthesis was assessed using a particular subtype-specific (−)ssDNA template, similar results were obtained regardless of whether subtype B or C RT was used.
DNA synthesis from DNA templates containing repeats of codons 64, 65 and 66 of pol
Synthetic DNA templates containing three inverted repeats of the 64, 65 and 66 codons of pol were employed to investigate how the pausing reactions in the region would vary if they were inverted and repeated (Figure 5). When subtype C RT was used along with the subtype C template containing the repeats to synthesize DNA, the strong pausing reaction seen at the K65 codon was repeated three times and was strongest at the third repeat at later time points (Figure 5A, lanes 1 to 6). Therefore, repeating the homopolymeric region multiple times produced the same strong pausing reaction at each segment at which RT encounters the final A base just prior to the G at codon 65 (Figure 5B). By contrast, when subtype B RT was employed along with the subtype B template containing the repeats to synthesize DNA, the ladder of pausing events that spanned all three codons was again seen and, as expected, no distinct pausing at the 65 region was observed (Figure 5A, lanes 7 to 12). Moreover, the pausing remained at the same position regardless whether the template used contained a homopolymeric stretch of thymine or, as in the case of the inverted DNA templates, a homopolymeric stretch of adenines.
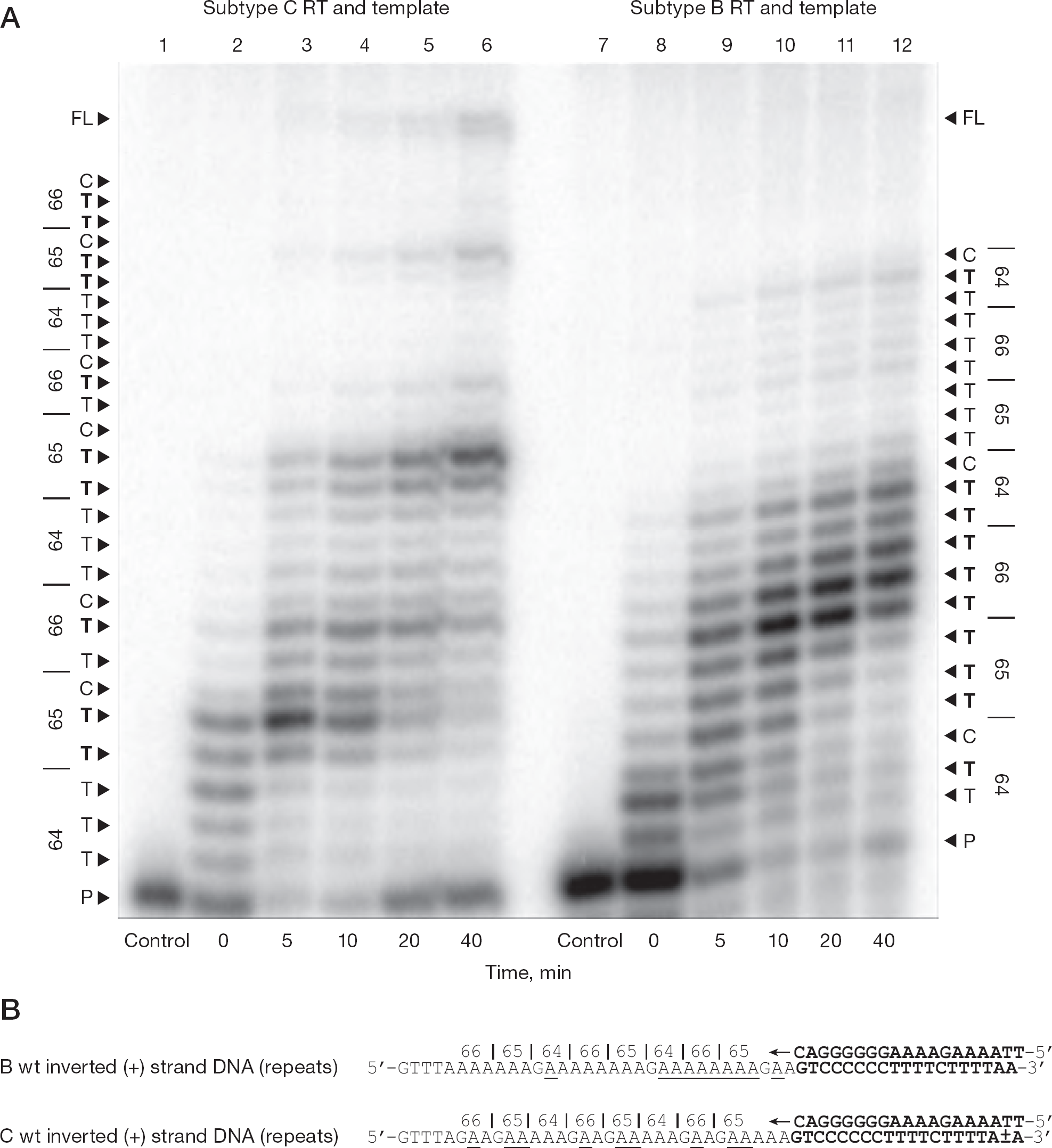
DNA synthesis from DNA templates containing repeats of codons 64, 65 and 66 of pol
DNA synthesis from inverted RNA and DNA templates
To further explore the pausing patterns observed with different templates and nucleotide concentrations, inverted (+)ssRNA templates were used (Figure 6). Inverted template sequences, although not readily encountered during the replication cycle of HIV-1, could provide insight on the importance of the orientation of the homopolymeric sequences with respect to pausing and possible resistance development. The data show no detectable pausing with the subtype B template, whereas use of subtype C template resulted in pausing at the two last positions of the thymidine stretch at the K65 (+15) position when employing the inverted (+) RNA templates (Figure 6A). The time course experiments revealed that pausing at position +15 in the K65 region can be spontaneously alleviated in certain reactions extending beyond 40 min using the inverted templates possibly because of the orientation of the homopolymeric tract. However, as shown in Figure 6B, pausing was much less intense than in (+)dsDNA synthesis from the subtype C (−)ssDNA template. With increasing dCTP concentrations >5 μM, pausing at the last two positions of the thymidine stretch in the K65 (+15) region was again alleviated, and the amount of full-length product was increased.
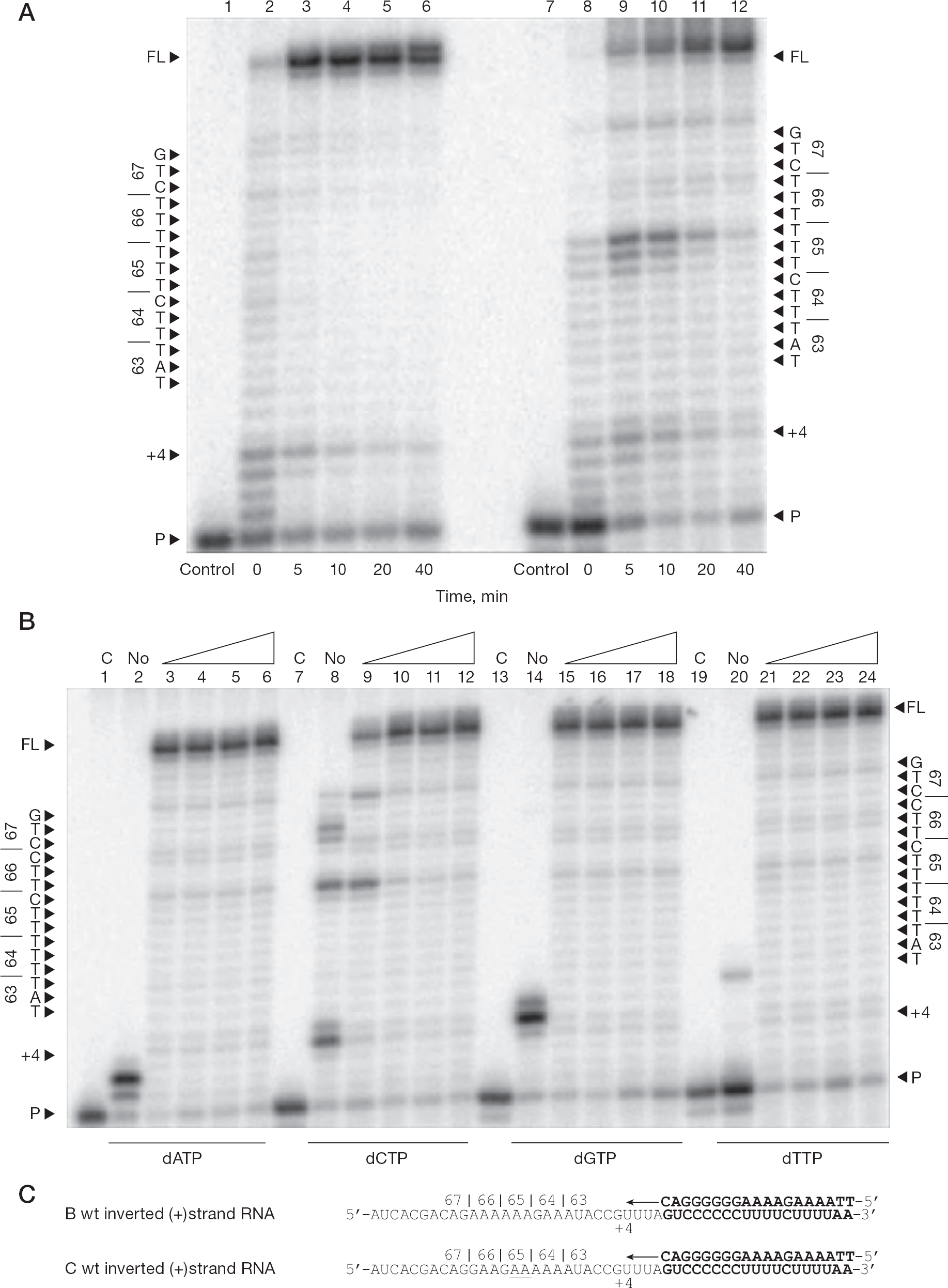

DNA synthesis from inverted 65 region of pol using variable single dNTP concentrations
These findings once again demonstrate the template-specific nature of the pausing observations. When inverted DNA templates were used to synthesize (+) dsDNA, no significant differences were observed with varying nucleotide concentrations (Figure 7). The subtype B template revealed the characteristic ladder of pausing events that spanned codons 64, 65 and 66 (positions +12 to +19; Figure 7A) whereas the subtype C template yielded the characteristic and more intense pausing at the K65 (+15) position (Figure 7B). The results using the inverted templates differ from those with the natural viral templates, suggesting that the orientation of the homopolymeric region might not have a major effect regarding the location of the pausing patterns observed, but does appear to be involved in the alleviation of pausing at the positions examined in our study.
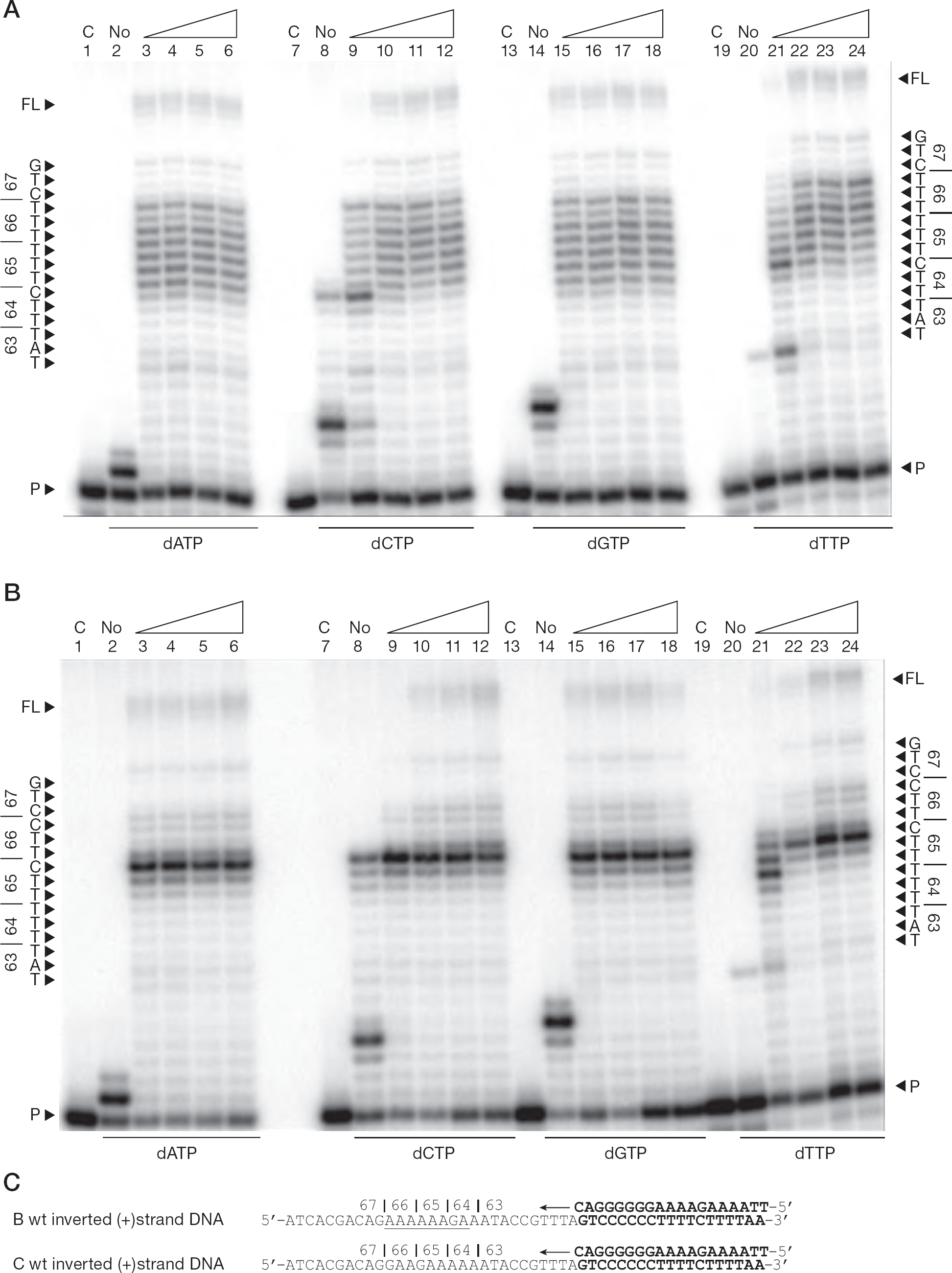
DNA synthesis from inverted (+)strand DNA templates using variable single dNTP concentrations
Discussion
The mechanism that we describe is based on increased rates of incorrect nucleotide incorporation at pausing sites of subtype-specific nucleotide sequences of HIV-1. Because the pol gene of HIV-1 has a high adenine content of approximately 35%, including many homopolymers of thymidine and adenine, RT has difficulty synthesizing DNA products on such sequences [30]. The pausing produced by the RT enzyme at these homopolymeric sites can then allow for misalignment, misincorporation, strand transfer, insertions, deletions and recombinations, all of which can lead to the development of drug resistance [24–27,31–34]. Such differential rates of incorrect nucleotide incorporation in the context of the homopolymeric subtype C coding sequences of the pol gene result in the more rapid acquisition of the K65R mutation in subtype C HIV-1 [28]. It is important to note that the incorrect incorporation event that allows for subtype C HIV-1 to develop K65R more readily than subtype B occurs in the replication step that uses the (−)ssDNA template to produce the (+)dsDNA which subsequently becomes integrated into the host genome. This replicative step requires the erroneous incorporation of dGTP instead of dATP to yield the AGG of subtype C that encodes K65R (Figure 1).
Secondary structure is known to play an important role in mutagenesis for many viruses. Because mutagenesis occurred in our study during synthesis of (+) dsDNA from the (−)ssDNA intermediate and not during the first step of the replication cycle from the (+)ssRNA template, the compromised RNA secondary structure is unlikely to have affected our findings.
In addition, the sequences of the other subtypes are known. Sequencing of codons 60 through 70 of subtypes A, D, F1, F2, G, J and K offers similar results to that of subtype B, whereas the sequences of subtypes H, N and O resemble that of subtype C. The NL4–3 sequence is highly conserved compared with the vast majority of subtype B clinical isolates with the exception of two silent polymorphisms, that is, an A62 that can be encoded by GCT instead of GCC and a K66 that can be encoded by AAG instead of AAA. These silent polymorphisms are located outside and at the end of the homopolymeric region and do not affect our results. The MJ4 sequence differs from a proportion of subtype C clinical isolates in that K66 is encoded by AAG instead of AAA. Because this silent polymorphism is located after the end of the homopolymeric region, it does not affect our observations that show that pausing is always present at the K65 position of subtype C templates regardless whether an AAG or AAA codon is present at position 66.
During the replicative step that uses (+)ssRNA to generate (−)ssDNA, no pausing is seen, suggesting that the probability of a mutation to occur at this stage is low. However, repetition of this experiment with variable dNTP concentrations yielded interesting findings. When dCTP was absent from reactions, DNA synthesis resumed and generated full-length DNA that reflected four independent incorrect dNTP incorporation events on the templates employed (Figure 2). The full-length DNA product was only achieved with subtype B and not subtype C template, when no dCTP was present. In addition to the possibility of the formation of a weak wobble-base-pair that might have provided a limited amount of DNA extension in the absence of dCTP, synthesis with both subtypes proceeded past the +4 pausing site. Once past the +4 position, DNA synthesis then stopped strongly at the +8 pausing site, immediately before the homopolymer tract of adenines on both templates. This might be a result of the difficulty that RT experiences in synthesizing DNA on polyadenine or polythymidine stretches on the template, thereby predisposing it to pause at these locations. Because the homopolymeric tract ends at the exact nucleotide position responsible for the K65R mutation, adding an additional G at that position (as in the case of K65R occurrence) would shorten the tract; however, pausing would still occur one base earlier on the DNA template.
Also, when subtype B RT was employed to synthesize DNA on a subtype C template, similar pausing profiles were obtained as when subtype C RT was used to synthesize DNA on a subtype C template [27]. Moreover, when (−)ssDNA synthesis was assessed using subtype-specific (+)ssRNA templates or when (+)dsDNA synthesis was assessed using subtype-specific (−)ssDNA templates, similar results were obtained regardless of whether subtype B or C RT was used.
DNA synthesis by subtype C RT appears to be less efficient at the single nucleotide position responsible for the K65R mutation (Figures 2 and 3). However, the biochemical and enzymologic profiles of subtype B and C RTs are similar and, in general, patterns of DNA synthesis by both enzymes are also similar [35,36]. As a result, one would not expect to see differences in clinical disease caused by subtype C RT alone. Although a new study suggests that Nef sequences of different subtypes could play a role in the differential development of neurodegenterative disease [37], long-term follow-up of patients is still needed.
In general, these results further strengthen the notion that the mechanistic basis of the observations lies in the template sequences themselves, rather than in the RT enzymes, thereby confirming the template-specific and enzyme-independent nature of the observations.
To study the specificity of the pausing reaction, we employed inverted DNA templates containing a homopolymeric stretch of adenines which mimics that of the 65 region of pol, and which was repeated several times. The data show that pausing was still present at codon 65 but that the pausing reaction was repeated only in the context of the subtype C template. By contrast, the ladder of pausing events observed with the subtype B template remained intact despite the inverted sequence, and became progressively shifted with time toward the later repeats. These findings confirm that the pausing reaction can proceed with templates containing homopolymeric regions of either adenine or thymine and that the pausing only occurs at the end of such sequences regardless of their orientation.
Moreover, when (+)dsDNA synthesis was evaluated using the subtype B (−)ssDNA template, higher concentrations of dCTP promoted full-length DNA synthesis and alleviated pausing at the end of the ladder (Figure 4), which culminated at the D67 (+20) position. The D67 codon is an important site for the development of thymidine analogue mutations (TAMs) that also confer high levels of resistance against many drugs used in antiretroviral therapy. The pausing at position D67 (+20) was alleviated faster with use of subtype C than with the subtype B template, suggesting that subtype B viruses might have a higher mutational propensity at the D67 (+20) position than subtype C viruses. Interestingly, because the D67N mutation results from a GAC to AAC mutation, one would expect that increasing dGTP (the correct nucleotide) or dATP (the nucleotide that gives rise to D67N) and not increasing dCTP levels would provide pausing alleviation and subsequent full-length DNA synthesis. The mechanism by which dCTP produces pausing alleviation remains unclear; however, it is likely that the homopolymeric sequence that ends in this region on the subtype B template plays a role similar to that of the homopolymeric sequence that ends at the 65 position on the subtype C template. These findings are again suggestive of the sequence-specific and enzyme-independent nature of the mechanism. By contrast, pausing at the K65 (+14) position of the subtype C template was not affected by altering the concentration of dCTP or any other nucleotide during DNA synthesis, perhaps because of the antagonism that exists between the K65R and TAM mutational pathways [38–41]. Differences in intensities and maintenance of pausing at position K65 (+14) are therefore likely responsible for the greater propensity of subtype C viruses to enter the K65R pathway, whereas subtype B viruses might favour the TAM pathway, based on template coding sequences that promote pausing and, as a result, possible mutational events leading to drug resistance. However, because pausing is observed at the D67 (+20) position of the subtype C template, K65R development might be delayed because of the antagonism that exists between TAMs and K65R. Confirmation of these hypotheses does, however, still require long-term follow-up of patients infected with subtype C HIV-1 receiving combination antiretroviral therapy.
In summary, we have proven biochemically that subtype C viruses prefer to enter a K65R mutational pathway whereas subtype B viruses might be more likely to generate TAMs because of the nucleotide differences between their respective templates. In addition, dCTP, when present at high concentrations, can contribute to alleviation of pausing at the D67 position on both subtype B and C templates. By contrast, pausing at the K65 region was readily maintained on the subtype C template and was not alleviated by altering its orientation or by repeating the homopolymeric sequence surrounding it, nor was the pausing alleviated by using higher concentrations of dNTPs. These findings help to explain clinical, tissue culture and biochemical observations suggesting that subtype C viruses might generate the multi-nucleoside resistance K65R mutation more frequently than subtype B viruses.
Footnotes
Acknowledgements
This research was supported by grants to MAW from the Canadian Institutes of Health Research (CIHR) and Gilead Sciences, Inc. DC is the recipient of a CIHR MD/PhD fellowship award. Work by DC was performed in partial fulfillment of the requirements for a PhD degree from the Faculty of Graduate Studies and Research, McGill University, Montréal, QC, Canada. We thank Daniela Moisi, Maureen Oliveira, Susan P Colby-Germinario, Cesar Collazos and Shalom Spira for technical assistance.
The authors declare no competing interests.
