Abstract
Transplantation of stem or progenitor cells is an attractive strategy for cell replacement therapy. However, poor long-term survival and insufficiently reproducible differentiation to functionally appropriate cells in vivo still present major obstacles for translation of this methodology to clinical applications. Numerous experimental studies have revealed that the expression of just a few transcription factors can be sufficient to drive stem cell differentiation toward a specific cell type, to transdifferentiate cells from one fate to another, or to dedifferentiate mature cells to pluripotent stem/progenitor cells (iPSCs). We thus propose here to apply the strategy of expressing the relevant key transcription factors to guide the differentiation of transplanted cells to the desired cell fate in vivo. To achieve this requires tools allowing us to control the expression of these genes in the transplant. Here, we describe drug-inducible systems that allow us to sequentially and timely activate gene expression from the outside, with a particular emphasis on the Tet system, which has been widely and successfully used in stem cells. These regulatory systems offer a tool for strictly limiting gene expression to the respective optimal stage after transplantation. This approach will direct the differentiation of the immature stem/progenitor cells in vivo to the desired cell type.
Keywords
Promises and Obstacles for Stem Cells in Cell Replacement Therapy
Transplantation of stem or progenitor cells is an attractive and, therefore, frequently proposed therapeutic option to replace cells lost or injured in traumatic and degenerative disorders in which the intrinsic capacity for cell renewal is limited or absent. Consequently, several phase I clinical trials have already been started or have just received approval to transplant stem cells with cell replacement as one of the objectives. A variety of stem cell sources have been used to generate cardiomyogenic cells for transplantation to patients who have suffered myocardial infarction (85), but see also (198). Muscle satellite cells or myoblasts were transplanted to patients with muscular dystrophy (179). Damaged cornea was restored by transplanting autologous stem cells derived from corneal limbal epithelium (137). Autologous human neural stem cells (NSCs) were used to replace dopaminergic (DA) neurons in Parkinson's disease (PD) (109). Clinical trials with neural stem cells are underway for chronic spinal cord injury (http://www.stemcellsinc.com/Therapeutic-Programs/Clinical-Trials.htm), amyotrophic lateral sclerosis (ALS) (115), and stroke (116). Approval has recently been granted for clinical trials with transplantation of human embryonic stem cells (hESCs) to replace photoreceptors in juvenile and age-related macular degeneration (http://www.advancedcell.com/patients/clinical-trial-information/), oligodendrocytes in spinal cord injury (116), and motor neurons in spinal muscular atrophy (SMA) (115).
Although numerous experimental studies have provided evidence that stem cell transplants can contribute to tissue repair and functional recovery in a range of animal models for injury or disease, it is still not mechanistically clear how the transplanted cells mediated these beneficial effects. In many instances, symptom amelioration appears to be mediated by cell-protective and anti-inflammatory mechanisms or by trophic bystander effects rather than through replacement of the cells lost (23, 123, 212).
However, many disorders including PD, Huntington's disease, ALS, multiple sclerosis, stroke and traumatic injuries to the central nervous system (CNS), heart disease, or the loss of β-cells in patients with type 1 and type 2 diabetes require that the cells permanently damaged or lost are replaced. Clinical trials with stem cell transplants in, for example, cardiac disease, have shown that this approach can be safe and may even have beneficial effects, but also that the long-term survival of the transplanted cells is typically poor (85). This problem is frequently observed with stem cell transplantations aimed at replacing lost cells (21); other commonly occurring problems are incomplete differentiation of the transplanted cells (70) or their differentiation to other types of cells rather than exclusively to the desired ones (57, 100, 125).
At the same time, differentiation of ESCs toward many different cell types has been achieved in vitro (20, 156), thereby providing cell sources for regenerative medicine. There are many examples when in vitro predifferentiated ESCs were successfully transplanted into the host and became functional specific cell types (90, 178, 208). Often however, these transplanted cells display unusually high and accelerated cell mortality after transplantation and, as a consequence, have inadequate therapeutic efficacy as well as the risk of inducing undesirable inflammatory reactions in the host tissue (9, 160, 164). This affects neural cells in particular, which have a very narrow time window for successful transplantation to a host (43, 186). It has been demonstrated that neural cells have to be transplanted at an early postmitotic stage to avoid damage to their fibers during the transplantation procedure. Once the cells have been transplanted, their survival and further differentiation, including incorporation to the local neuronal circuitry, becomes entirely dependent on their interaction with the host environment. It would, therefore, be of advantage if the properties of the transplanted stem cells could be influenced not only before but also after their transplantation.
In vitro stem cell differentiation is either controlled by adding specific small molecules to the culture medium (110, 111, 187, 208, 227) by ectopically expressing proteins that are essential to drive the differentiation process to the desired cell type either by transfecting expression vectors for their genes (62, 107, 120) or by introducing the recombinant proteins (93, 220) or their encoding mRNAs themselves (162).
Clearly, this small molecule-driven approach will not allow clean, clear-cut control over stem or progenitor cell fate after transplantation due to systemic interference with the other cell types in the host organism. We therefore propose an alternative approach in which we artificially control stem cell destiny from outside via heterologous expression of a limited set of genes essential for the differentiation process. Inducible gene regulation systems, exemplified by, but not limited to, the Tet system are triggered externally by administering small molecules to the recipient organism. They can, thus, provide the tools we need to help control and guide the in vivo differentiation of stem cells after their transplantation. These systems make it possible to insert cell autonomous survival or differentiation protocols into stem/progenitor cells prior to their transplantation, which can then be activated after transplantation in the cells we want for the period of time we need.
Expression of Key Transcription Factors to Guide Cell Differentiation
An important aspect in any cell replacement therapy is to generate the required cell type exclusively and efficiently in an environment that is either indifferent to or even unfavorable for stem/progenitor cell differentiation to mature cell types (26, 42, 78, 204). Since transcription factors (TFs) are downstream effectors of signaling pathways, their ectopic expression can be used instead of external cues to set up a cell autonomous program, which will then direct cell differentiation to the desired cell type. The crucial step involved in this approach will be to isolate the respective minimal set of transcription factors required to flip the switch toward the desired cell type out of the entire set of transcription factors present during developmental cell patterning and specification. One way to identify the relevant TFs for a certain differentiation pathway is to follow the normal steps taken during natural development (76). A second, alternative method, which is increasingly being applied, is to identify the key transcription factors for a specific differentiation step by transfecting or transducing the host cells with a mixture of constructs expressing different transcription factors that are suspected to be important for the differentiation process. An initially successful “TF differentiation cocktail” can then be deconvoluted to identify the minimal TF set necessary to guide the differentiation process. There are elegant examples demonstrating that a limited number of TFs is sufficient to completely change the differentiation state of the cells—the original generation of iPSCs (193) and the recent transdifferentiation of mature fibroblasts to functional neurons (201).
That transcription factors have the power to control cell fate has been demonstrated convincingly in many studies involving not just cellular reprogramming and dedifferentiation but also transdifferentiation or directed differentiation [for a minireview, see (27)]. Figure 1 shows how these three processes are connected with each other. In the original landmark reprogramming protocol, four transcription factors of 24 selected genes were sufficient to induce mature murine fibroblasts to readopt a pluripotent state (193). Careful selection of the starting cell type could even further reduce this requirement to just a single factor, octamer binding transcription factor 4 (Oct4) (95, 96). Transdifferentiation is an equally impressive example of the important role transcription factors play in deciding cell fate. Transdifferentiation describes the direct conversion of one cell type to another through simultaneous down-regulation of one genetic program and concomitant up-regulation of the new genetic program (86). For example, B-cells are converted to macrophages by ectopically expressing the transcription factors CCAAT-enhancerbinding proteins a (CeBPα) and CeBPβ (215). In adult mice, mature pancreatic exocrine cells transdifferentiate into cells closely resembling β-cells after infection with adenoviruses expressing the three transcription factors neurogenin 3 (Ngn3), pancreatic and duodenal homeobox 1 (Pdx1), and musculoaponeurotic fibrosarcoma oncogene homolog A (Mafa). These three were identified out of an initial group of nine genes, which exhibited β-cell developmental phenotypes when mutated (224). Starting from a pool of 19 candidate genes, a combination of three transcription factors, achaete-scute complex homolog 1 (Ascl1 or Mash1), brain-2 (Brn2 or OCT7), and myelin transcription factor 1-like (Myt1l), was found to transform both murine and human fibroblasts directly into functional neurons (151, 201). Human fibroblasts were then further differentiated toward dopaminergic neurons by additional expression of the two genes LIM homeobox transcription factor 1α (Lmx1a) and forkhead box A2 (FoxA2) (151). Other examples include transforming fibroblasts into cardiomyocytes (80) and cells from a human dermal subset into blood cell progenitors (189). In directed differentiation, specific cell types are generated from stem/progenitor cells by treating them either with combinations of trophic factors and/or small molecules (14, 30, 77, 79), by ectopically expressing transcription factors, or by a combination of both (19, 51, 58, 138). Panman et al. (143) induced specific neuronal subtypes effectively with single transcription factors if the neural progenitor cells were competent to differentiate to the desired cell type. But if the stem/progenitor cells had a nonpermissive regional character, then a combination of “regionalizing” regulatory factors together with the subtype-specific factor were needed to achieve efficient differentiation.

Transitions between cell types. An embryonic stem cell (top left), a neuron (top right), and fibroblasts (bottom) are arranged as a triangle. The differentiation steps that lead to their interconversion are indicated, as well as the relevant transcription factors, growth factors, and morphogens (RA, retinoic acid; SHH, sonic hedgehog; FGF8, fibroblast growth factor 8) that were used [the dedifferentiation factors are from (193), the transdifferentiation factors from (201), and those for directed differentiation were taken from (18)].
Current State of Stem Cell Transplantation as Cell Replacement Therapy for Major Disorders
There is a long list of medical conditions that are potential therapeutic targets for stem cell transplantation. Here, we limit our discussion to several disorders with huge impact on the affected individuals and on society. Based on developmental studies and in vitro differentiation protocols, we show the set of key TFs, which play an important role in generating the respective corresponding types of cells. These TFs may be activated for advanced differentiation of transplanted predifferentiated ESCs in vivo after transplantation in order to guide their further differentiation toward functional cells.
Parkinson's Disease
In PD, the dopamine-releasing neurons in the substantia nigra are gradually lost, resulting in the progressive and severely disabling motor dysfunction, which is the hallmark of this disease. Previous studies in experimental animal models of PD have shown that dopamine release can be restored and motor dysfunction reversed by transplantation of embryonic neurons into the striatum [reviewed in (114)]. Clinical trials with human embryonic DA neurons initially provided encouraging results, but later follow-up evaluations failed to demonstrate any significant improvement [reviewed in (114)].
DA neurons have been generated in vitro from iPSCs, ESCs (93), mesenchymal stem cells (MSCs), regional stem/progenitor cells, and fibroblasts. In vitro predifferentiated cells were subsequently grafted into the striatum and found to partially reverse PD-like symptoms in animal models (30, 145, 161, 169). However, the mechanism(s) responsible for symptom reversal are not fully understood, since functional improvement in the 6-hydroxy dopamine rat model of PD was shown to occur from human neural progenitor cell transplants without differentiation to DA neurons (75). More recently, using the same animal model, forced expression of the TF Lmx1a in striatal mouse ESC transplants resulted in robust graft survival and extensive striatal dopaminergic innervation by the grafted cells (51).
Several transcription factors and extrinsic factors were identified to be crucial for dopaminergic neural differentiation. Stem cells harvested from the E12.5–13.5 mouse ventral midbrain express Neurogenin2 (91, 195), there after nuclear receptor related 1 (Nurr1) (4) and subsequently Mash1 (146). The nuclear transcription factor Nurr1 is involved in the development and maintenance of the midbrain dopaminergic neuronal phenotype (82). There is a well-established correlation between olfactory symptoms and PD (16). This provides a possible link between PD and adult neural stem cells, which are continuously produced in the subventricular zone (SVZ) of the lateral ventricle and migrate to the olfactory bulb where they give rise to interneurons. Glial cell line-derived neurotrophic factor (GDNF) has been shown to protect and restore DA neurons in injury models and is being evaluated for the treatment of Parkinson's disease (214).
Generation of functional DA neurons was also achieved from hESCs in vitro (30), in vivo from neural progenitors expressing orthodenticle homeobox 1 (Otx1) (64) and after reprogramming mouse and human somatic cells by viral transduction of four transcription factors, Oct4, Krüppel-like factor 4 (Klf4), sex-determining region Y box 2 (Sox2), and myelocytomatosis viral oncogene homolog (c-Myc) (119, 147, 191). Recently, human iPSCs were efficiently generated from skin biopsies of PD patients by using doxycycline (DOX)-inducible delivery of reprogramming factors (182). These reprogrammed fibroblasts were made by cotransducing lentiviral vectors expressing a Tet-On transactivator (FUW-rtTA2S-M2) together with DOX-inducible transgene cassettes expressing either the four or just three (Oct4, Sox2, Klf4) reprogramming factors. For the subsequent directed differentiation of hESCs into DA neurons, conventional in vitro protocols were used (21, 45, 94, 149, 169). DA neurons were also recently generated from human mature fibroblasts by enforced expression of Lmx1a and FoxA2 in addition to the three neural transdifferentiating factors Ascl1, Brn2, and Myt1l (151).
Nurr1 alone is not sufficient to drive mesencephalic stem/progenitor cells to the dopaminergic phenotype (81), but differentiation to dopaminergic neurons was achieved in collaboration with paired-like homeodomain 3 (Pitx3) (180). Recently, new insights on the role of Nurr1 in PD revealed that Nurr1 appears to protect against loss of dopaminergic neurons in part by limiting the production of neurotoxic mediators by microglia and astrocytes (167). This observation highlights the importance of controlling the environmental cues to achieve optimal survival of transplanted DA neurons.
Stroke and Traumatic Brain Injury
Stroke is the third most common cause of death in the Western world and one of the main causes of long-term severe disability. One major line of research attempts to promote recovery by recruiting endogenous neural stem cells to the ischemic territory, whereas the other focuses on stem cell transplants. The brain tissues affected by stroke or trauma undergo a complex series of pathological events with spatially and temporally different requirements on stem cell transplants. Initially, neuroprotection should have the highest priority, followed by tissue repair, and finally the restoration of as much as possible of the damaged neural circuitry. The main components of neuronal networks in the brain are glutamatergic and γ-aminobutyric acid-ergic (GABAergic) neurons as well as astrocytes. These cell types have been generated in vitro primarily from ESCs or from region-specific neural precursors from embryos or adult mice and rats (65, 221). Stem/progenitor cells delivered intravascularly (177) or intracerebrally (5) to the ischemic brain are able to improve symptoms, even if the generation of functional neurons does not occur.
Thus, the positive results of stem cell transplants in experimental stroke models so far are most likely the consequence of the release of trophic factors, stimulation of intrinsic repair processes, or a combination of both.
Recent publications proposed stem cells not only as a potential source for cell replacement in central nervous system injury but also as a source for trophic molecules that might minimize damage and promote recovery (24, 113). NSCs appear to be an effective source for successful differentiation into representative cell types in parts of the brain most commonly affected by stroke, such as cortical projection neurons (46), interneurons (172), hippocampal pyramidal neurons (36, 46), motor neurons (54), astrocytes (47, 68, 210), and oligodendrocytes (154, 217). Human- and mouse-derived ESCs, including the more recently described iPSCs, are also able to differentiate into all adult cell types, including functional neurons (13, 135) and glia (22, 44, 171).
Neurons and astrocytes are derived from common multipotent neural progenitor cells, which sequentially pass through phases of expansion, neurogenesis, and astrogenesis (71, 131). If the transcription factor sequence glioma-associated oncogene family zinc finger 2 (Gli2)-Sox2-hairy and enhancer of split 5 (Hes5) maintains the undifferentiated state of neural precursor cells (194), the transcription factor paired box 6 (Pax6) controls neurogenesis and also regulates proliferation, differentiation, and migration of astrocytes in the CNS (168). The suppression of proneural basic helix-loop-helix (bHLH) genes, primarily Ngn1 and Ngn2, results in stem cell differentiation toward the glial lineage (72). Differentiation of GABAergic cortical neuron differentiation is driven by the transcription factors Nkx2.1, Mash1, and distal-less homeobox 1 (Dlx1) (34, 155) [see also (132) for a review]; glutamatergic cortical neuron development by the sequence Pax6-T-box, brain 2 (Tbr2) neuronal differentiation 1 (NeuroD), and Tbr1 (61, 69), that of cortical oligodendrocytes by a succession of Sox10, oligodendrocyte transcription factor 1/2 (Olig1/2), Mash 1, and Nkx2.2 (148, 225), whereas cortical astrocytes were either associated with a Sox9, inhibitor of DNA binding 1/3 (Id1/3), Hes1/5 expression pattern (87, 163), or with the combination of Sox10, Nkx2.2, Olig2, and Olig1 (25).
Myocardial Disease
Ischemic heart disease is the most common cause of death in the Western world. Cardiomyocytes are the contracting elements of the heart, and the intrinsic capacity of the heart to replace these cells is very limited. Transplantation of stem cells that can replace lost cardiomyocytes is therefore an attractive option to treat this condition. Cardiomyocytes have been generated in vitro from a wide range of stem/progenitor cells, including iPSCs (53, 104, 150), ESCs (15, 184), hematopoietic progenitor/stem cells, MSCs (6, 31, 56, 63), and cardiomyocyte progenitor cells (181). An additional source for repair of myocardial contractility are skeletal myoblasts (140), although they do not seem to be able to differentiate to cardiomyocytes (108, 159). Intravascular delivery or cardiac transplants of multipotent or predifferentiated cardiogenic cells from these stem cell sources have been shown to promote cardiac structural repair and functional restoration in animal models of myocardial injury (52, 66, 84, 126, 141). The encouraging results from this experimental research have prompted several clinical trials in patients with myocardial disease, using different types of progenitor/stem cells (85, 152). The results from these studies are inconsistent, but some report beneficial effects on cardiac function in the recipient patients. Differentiation of the transplanted cells to cardiomyocytes appears to be absent or minimal, their long-term survival is poor, and most importantly, the functional improvement observed does not appear to depend on their presence (103, 188).
Multipotent cardiac progenitor cells (CPCs), expressing Nkx2.5, fetal liver kinase 1 (Flk1), or insulin gene enhancer protein (Isl1), can be isolated from early embryos or generated from ESCs and give rise to cardiomyocytes as well as endothelial and smooth muscle cell lineages (32, 89, 105, 133, 213).
The discovery that heart defects in humans often result from haploinsufficiency of cardiac-specific TFs suggests that strategies to activate such developmental regulators may contribute to the differentiation of stem cells that are transplanted with the purpose of producing substituting functional heart cells. Thus, mutations in Nkx2.5 (174), T-box 5 (Tbx5) (11), Tbx1 (129, 203), or GATA4 (55), which activate the genes in the core cardiac network [reviewed in (142)], result in serious heart abnormalities. In addition, recent studies have revealed that these myogenic TFs activate the expression of a set of conserved microRNAs (miRNAs) that regulate transcriptional networks, resulting in precise cellular responses to developmental, physiological, and pathological signals [reviewed in (153, 209)]. Activation of neuregulin-1 (NRG-1) in ESCs resulted in up-regulation of the expression of the cardiac-restricted TFs Nkx2.5 and GATA-4 (205).
Insulin-Producing β-Cells
Type 1 diabetes is characterized by an autoimmune-mediated loss of insulin-producing β-cells in the pancreatic islets of Langerhans. Today, transplantation of either the entire pancreas or of isolated islets has become a treatment of choice for selected patients with diabetes mellitus (50, 165). However, long-term results after islet transplantation are disappointing with adequate graft function seen in less than 10% of the patients after 5 years (166). Furthermore, the number of patients in need of new β-cells far outnumbers the limited access to islet tissue for transplantation. Transplantation of stem cells to replace the lost β-cells is therefore an attractive therapy for long-term treatment of type 1 diabetes and also for some cases of type 2 diabetes. Insulin-producing β-cells have been generated from several sources, including ESCs (8, 49, 121, 175, 200), stem/progenitor cells from the exocrine pancreas (41, 62, 124, 139), biliary ducts (136), MSCs from various sources (88, 144, 216), and iPSCs derived from fibroblasts of patients with diabetes type 1 (117). Human ESCs have been converted to β-cells capable of synthesizing insulin through a stepwise procedure of transcriptional regulation that mimics the normal development of β-cells (37, 38). Exocrine pancreatic cells were also shown to give rise to insulin-producing β-cells by transcriptional reprogramming with a specific combination of the three transcription factors Ngn3, Pdx1, and Mafa (224).
Numerous studies have reported beneficial effects of MSC transplants in animal models of type 1 diabetes. They are presumably due to modification of inflammatory processes and to improved vascularization [(88, 144), reviewed in (202)]. Transplantation of ESCs to diabetic mice also improved glucose homeostasis indirectly by promoting endogenous β-cell neogenesis (98). Human ESCs predifferentiated to committed pancreatic endoderm developed into functional β-cells after transplantation to immune-compromised mice (101). Interestingly, treatment of the recipients with the β-cell toxin streptozotocin destroyed their endogenous β-cell population, but the grafted cells were protected and provided a functional source of insulin. Thus, although efficient and reproducible replacement of lost β-cells in type 1 diabetes has still not been fully achieved with stem cell transplants [reviewed in (197)], promising steps in this direction have been taken.
Precursor cells that give rise to the definitive endocrine pancreas emerge at E13.5 and express a series of key TFs, including Pdx1 and hepatocyte nuclear factor 6 (Hnf6), followed by Ngn-3, β2/NeuroD, and Pax4 (1).
Differentiation of β-cells/progenitors from human ESCs was achieved in vitro by activating either Pax4 (112) or Pdx1 expression (29, 106) from mouse ESCs and from progenitor cells by activation of Ngn3 expression (176, 218) and in liver progenitor cells by activation of Pdx1 (83). In already differentiated cells, ectopic expression of Pax4 lead to redifferentiation of endocrine precursor cells (35), whereas in liver cells activation of Pdx1 and Ngn3 was required (219). Adult pancreatic exocrine cells were reprogrammed with Ngn3, Pdx1, and Mafa (224).
Drug-Inducible Regulation of Transcription Factor Expression in Cell Differentiation
General Remarks
In the previous two sections, we provided several examples, out of many possible, that enforced expression of only a few key transcription factors, which are normally present during development, can induce cell differentiation to a desired cell type. Major efforts in current stem cell research focus on in vitro methods to induce commitment to a defined cell type in the donor cells prior to their transplantation. This valuable approach resulted in the successfully continued differentiation of neural stem cells transplanted to a developing nervous system (208). However, to guide differentiation of the transplanted cells to the right cell (sub)type out of the large number of different cell types they can diversify to and to have them establish the correct connections after grafting are still major challenges for successful neural stem cell transplantation. To meet these challenges, inducible gene regulation offers a tool to control differentiation of stem/progenitor cells after their transplantation to the damaged or diseased tissue.
Inducible Gene Regulation Systems
Inducible gene regulation systems have several properties that predestine them for in vivo control of gene expression: (i) they allow transgene expression to be turned on or off at any desired time-point (60), (ii) they also allow transgene expression to be turned on or off repeatedly (128), and (iii) the transgene expression level can be modulated by titrating the effector molecule's concentration (170). This allows flexible adjustment of transgene expression to the specific needs of the differentiation protocol previously established in vitro [reviewed in (33)]. The importance of conditional regulation systems for inducing cell fate changes becomes clear from studying the large amount of published data on the generation of iPSCs from partially or fully differentiated cell types, which has clearly shown that (i) some cell types require longer expression of the reprogramming factors than others (118); (ii) expression of the reprogramming factors is needed only transiently, as evidenced from the down-regulation of their expression as a hallmark of true iPSCs (28) or the successful generation of iPSCs using either adenoviral vectors (183) or recombinant transducing proteins (92, 223); (iii) the expression levels of the four reprogramming factors required for dedifferentiation vary from cell type to cell type (207); and (iv) careful selection of the cell type used to initiate reprogramming can reduce the number of transcription factors needed down to a single one, Oct-4 (96). Taken together, the properties of inducible regulation systems nicely fulfill the requirements needed for controlling cell fate.
In most cases, the TFs needed to guide cell differentiation are expressed only for brief periods of time (151). It is therefore advisable to use conditional expression systems that require the presence of external ligands to induce target gene expression. Such regulatory systems are typically termed “ON” and two different implementations will be presented in the next section. The use of “OFF”-type regulatory systems, in which the presence of an effector molecule turns transgene expression off, would require this effector molecule to be applied for the remainder of the organism's life to prevent complications—which is clearly not acceptable in a clinical setting.
The Tet System
The Tet system is the most widely used conditional system for inducible control of transgenes in cell lines and organisms [reviewed in (17, 59)]. Its use in vivo and in gene therapy applications has been extensively reviewed recently (185, 196, 206), so that we are only going to focus on several, in our opinion, key aspects of Tet regulation with respect to cell differentiation in vivo. Tet system regulation is derived from bacterial tetracycline resistance determinants (17) and functions as an artificial transcription unit (Fig. 2). One component is a chimeric transcription factor, generated by fusion of a eukaryotic regulatory domain to the bacterial tetracycline repressor TetR. The TetR portion mediates sequence-specific DNA-binding and effector response, the regulatory domain transcriptional readout. The second component is a hybrid promoter formed by positioning multiple repeats of the specific TetR binding site tetO upstream of a TATA-box containing minimal promoter (59).
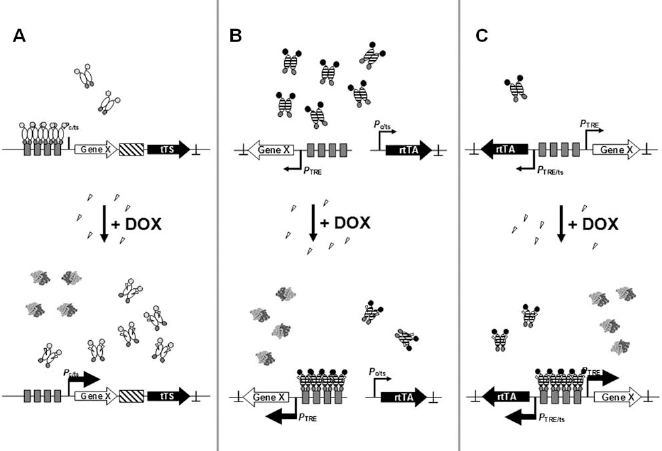
Regulatory principles of different Tet-On systems. (A) Repression by tetracycline-dependent transsilencer (tTS): In the absence of effector, tTS down-regulates expression by the promoter. In the presence of doxycycline (DOX), tTS dissociates from the promoter and both genes are expressed. (B) Regulation by a constitutively expressed reverse tetracycline-dependent transactivator (rtTA): rtTA is expressed independent of the presence of effector. In the presence of DOX, it binds to the promoter activating expression of the target gene. (C) Regulation by an autoregulatory rtTA loop: In the absence of effector, only small amounts of rtTA are made. After addition of DOX, rtTA binds to the promoter and activates expression of its own gene and of the transgene. The Tet-transregulators are symbolized by gray circles as DNA-binding domains, hatched ellipses representing rtTA and open ellipses TetR. A black circle symbolizes an activation domain, while a gray stop sign represents a repression domain. The transregulator-encoding genes are depicted as black arrows with inverse print, the target genes by an open arrow. Promoters are displayed as broken arrows with “TRE” representing a Tet-responsive element, “c” a constitutive, and “ts” a tissue-specific promoter. tTS- and rtTA-binding sites (tetO) are shown as gray boxes, an internal ribosome entry sequence as a hatched box, and polyA sites as ⊥. DOX is depicted as open triangles.
Tet-On type regulatory setups, in which transgene expression is initiated by addition of an effector, can be realized by different strategies. One is to fuse a silencing domain to unmodified TetR (Fig. 2A). The resulting transregulators are termed tetracycline-dependent trans silencers (tTS). In most applications, the silencing domain fused to TetR is the Krüppel-associated box (KRAB) domain from the human zinc finger transcription factor Kox1. It represses transcription from constitutive or cell-type-specific promoters effectively, but its presence can lead to irreversible silencing of target gene expression by introducing heterochromatin formation (7, 190, 211). A typical expression vector for tTS-mediated regulation is shown in Figure 3A. Its advantages are its small size that allows it to be introduced into lentiviral or adeno-associated virus (AAV) vectors. Its disadvantage is that any irreversible transgene silencing mediated by the KRAB domain should not already occur during the transduction and selection phase, prior to cell transplantation. Here, transgene expression has to be shut off to prevent premature differentiation of the stem cells. An effective alternative repression domain based on a “Pro-Leu-Asp-Leu-Ser” C-terminal binding protein (CtBP)-recruiting domain with reduced tendency to introduce irreversible transgene silencing was recently introduced (39, 67).
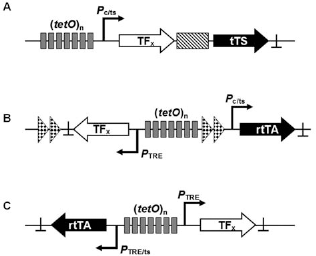
Typical designs for one vector Tet-On type regulatory systems (A) based on tTS-mediated regulation, (B) based on rtTA-mediated regulation, and (C) based on autoregulatory rtTA-mediated regulation. The Tet-transregulators are symbolized by a black arrow with inverse print, target genes by an arrow. Promoters are displayed as broken arrows with “TRE” representing a Tet-responsive element, “c” a constitutive, and “ts” a tissue-specific promoter. tTS- and rtTA-binding sites (tetO) are shown as gray boxes, an internal ribosome entry sequence as a hatched box and polyA sites as ⊥. Insulators derived from the chicken HS4 element are depicted as arrow heads.
The second approach is based on using TetR mutants that absolutely require tetracyclines, like doxycycline, to bind to their operators (Fig. 2B, C). Once fused to activation domains, they are designated as reverse tetracycline-dependent transactivators (rtTA). Second-(199) and third-generation rtTA variants (226) have been generated by random mutagenesis and screening in yeast or by viral evolution. They display no intrinsic leakiness and high sensitivity toward their effector, DOX. This is of great advantage, because the increased effector sensitivity should permit their usage in applications regulating gene expression in the central nervous system. This has been difficult with Tet-On systems in the past, due to their low sensitivity toward DOX (130). rtTA can either be expressed from constitutive or tissue-specific promoters (Fig. 3B) or in an autoregulatory setup (Fig. 3C).
With active polymerase II (Pol II) promoters, the transgene-expressing unit in a one-vector system should be insulated (3, 157). We routinely use 250 bp core sequences derived from the chicken HS4 insulator (158), but other insulators are also available like the sns5 (40) and arylsufatase elements (134) from sea urchins. They were shown to be functional in erythroid cells (40) and in a mouse quadriceps muscle transduction model (134). Insulators shield the transgene expressing unit from position effects due to activating or silencing elements at the vector integration site and from the nearby promoter driving rtTA expression. This efficiently reduces the integration site-dependent leakiness that is often observed with inducible system. They also increase the yield of regulating clones and improve the homogeneity of transgene expression between different clones (3, 157). The size of the expression cassette allows its insertion into AAV and lentiviral vectors. Autoregulated, bidirectional retroviral vectors combine the advantages of small size, allowing single-step transduction with strict regulation of the gene of interest in the infected target cells. Their disadvantage is that they lose the rheostat-type graded response to intermediate concentrations of the effector molecule. Instead, auto-regulated expression leads to a bimodal, On/Off expression response (12, 127).
A third source of leakiness is intrinsic to the minimal promoter used. Here, careful selection can help to avoid leaky transgene expression. A recent example of several Tet-controlled minimal promoters in a T-cell line demonstrated the mouse mammary tumor virus long terminal repeat (LTR) promoter-derived minimal promoter and the ClonTech-distributed minimal promoter TREtight showed no intrinsic leakiness when flanked by chromatin insulators and driven by a second-generation reverse transactivator (39).
Multiple Gene Expression Systems
Many of the examples mentioned in the third section require more than one TF to be expressed before full differentiation can be achieved. Expression of two genes in an independent manner can be obtained with the Tet system, but only if one of the transregulators is of the Tet-Off type, that is, requiring the constant presence of effectors to prevent transgene expression (10, 102). This is not advisable for reasons mentioned above. We therefore recommend the combined use of several of the many published artificial regulatory systems to achieve independent multigene expression control. Streptogramin-(PIP-ON) and macrolide-responsive (E-ON) expression units utilize a KRAB repression domain to control gene expression [reviewed in (206)]. They will therefore have the same advantages and disadvantages as tTS-regulated systems. Other On-type inducible gene expression units that have been used in vivo and in gene therapy applications include (i) the dimerizer system (97), which exploits chemically induced heterodimer formation to activate gene expression; (ii) modified steroid receptors and synthetic steroid analogs like mifepristone and tamoxifen (173, 192); and (iii) gene switches employing the insect-specific ecdysone receptor (48).
Taken together, the tools for guiding differentiation of stem/progenitor cells after their transplantation are available. These tools are likely to be particularly useful for stem cell-based cell replacement therapy in conditions where current protocols for postgrafting survival and predictable differentiation to desired cell types are unsatisfactory.
Application of the Tet System in Stem Cell Research
The Tet system has been used successfully for the inducible expression of transgenes in many cell types, with more than 150 publications describing its application in different types of stem cells, thereby impressively demonstrating its potency as an artificial regulatory system. Within the scope of this review, we therefore cannot provide an extensive analysis, but will rather give a brief overview, focusing on a few important aspects, of Tet system regulation.
In reprogramming (73, 74, 118, 182, 207) and transdifferentiation (151, 201) experiments, DOX-inducible lentiviral vectors were employed to alter cell fate. This approach uses a sole Tet-transactivator to drive all transcription factor expressing units, so their expression is not independent and cannot be used to further dissect the individual expression profile for each factor.
Inducible expression of either GFP or the phosphatidylinositol-glycan complementation class A gene (PIG-A), a gene involved in the synthesis of glycosylphosphatidylinositol-anchored proteins (GPI-APs) on the cell surface, was introduced in hESCs by a tTS-based transsilencing approach similar to the one presented in Figure 2A (222). The authors used a two-vector system to regulate expression of the reporter or the cellular gene. They demonstrated reversible induction of transgene expression in the hESC lines H1 and H9. They also noted that transgene expression was not fully restored after DOX addition to tTS-silenced cells. Both the number of cells responding to DOX treatment (frequently ~50%) and the level of transgene expression were affected, in agreement with the concerns voiced above against using a KRAB-based tTS variant.
Recent publications, in vitro (176) and in vivo (2, 99), demonstrate how the Tet system can be used in directed differentiation protocols using single transcription factors. DOX-induced expression of Ngn3 in ESCs that had been chemically predifferentiated to pancreas progenitors greatly enhanced their potential to differentiate into bona fide endocrine pancreas cells, although, to achieve terminal differentiation, the restoration of normal glucose levels, withdrawal of bone morphogenetic protein 4 (BMP4) and fibroblast growth factor 10 (Fgf10), and inclusion of nicotinamide were still required (176). This highlights several important points for in vivo guided differentiation. First, genetically modified ESCs can be chemically differentiated to a still fairly undifferentiated progenitor population. Second, Ngn3 expression greatly improved the further differentiation to pancreatic endocrine cells but was still not sufficient. One or maybe more transcription factors are still needed to achieve a fully cell autonomous differentiation from the progenitor stage. The in vivo proof of principle for such an approach was recently demonstrated by the successful generation of a specific type of sensory neurons—nociceptors—guided by Tet-controlled expression of the diversification factor runt-related transcription factor 1b (Runx1b) specifically in Sox10-expressing neural crest stem cells transplanted to the dorsal root ganglion cavity. Forced expression of Runx1b strongly increased transplant survival in the enriched neurotrophic environment of the dorsal root ganglion cavity and was sufficient to guide differentiation of boundary neural crest stem cells (bNCSCs) toward a nonpeptidergic nociceptive sensory neuron phenotype both in vitro and in vivo after transplantation (2).
To establish protocols for controlled differentiation of stem cells after transplantation (Fig. 4), it is first advisable to identify in vitro the optimal combination of transcription factors that, after induced activation, will drive un- or predifferentiated donor cells to fully functional cells once they have been equipped with drug-inducible regulatory systems controlling the factors previously identified. In vivo the protocol derived from these experiments will have to be tested and, most likely, modified following transplantation to small and large animals models for the relevant disorder(s) as we cannot predict all possible interactions in and with the host organism. But the prior in vitro validation is still necessary, because our approach aims at uncoupling stem or progenitor cell differentiation from the influence of extrinsic factors present in the host's cellular environment at the transplantation site. It is therefore essential to recapitulate the differentiation process in vitro both in terms of the specific transcription factors expressed and their temporal relationship, as well as the levels of their expression, since their quantities may influence their impact on the differentiation process (122).

Flowchart for setting up an in vivo directed cell differentiation study. The individual steps to be taken are indicated from top to bottom including the environment in which they should be undertaken. A shortcut from in vitro to in vivo work that can be started in parallel is highlighted on the right by an arrow. Adapted and extended from (27).
Conclusion
Several recent publications have shown that differentiation to the desired type of cells can be accomplished by expressing a limited number of TFs. Thus, the differentiation of dopaminergic neurons was successful after induced expression of Lmx1a (51), β-cell differentiation was achieved by expressing Ngn3, Pdx1, and Mafa (224), differentiation of neural crest stem cells toward nociceptor neurons was realized by forced expression of Runx1b in Sox10-expressing cells (2) and motor neurons were generated from human spinal cord derived stem/progenitor cells by ectopic expression of Runx1 (99). A consensus is beginning to emerge from several different fields of research that using regulated gene expression with drug-inducible systems is a promising experimental approach to differentiate stem cells in vivo after their transplantation. The practical utilization of this method in cell replacement therapy requires on the one hand the development of controlled guidance protocols for directed stem cell differentiation and on the other hand efficient methods of gene transfer into the stem cells and for delivery of the transfected material into the host. Once these goals have been met, this method can become an indispensable tool in regenerative medicine.
Footnotes
Acknowledgments
This review was funded by the Swedish Research Council, project 20716, Stiftelsen Olle Engkvist Byggare and Signhild Engkvist Stiftelse and German Research Foundation (DFG) for support through SFB473/D6. The authors declare no conflicts of interest.
