Abstract
Wikidata is a massive Knowledge Graph (KG), including more than 100 million data items and nearly 1.5 billion statements, covering a wide range of topics such as geography, history, scholarly articles, and life science data. The large volume of Wikidata is difficult to handle for research purposes; many researchers cannot afford the costs of hosting 100 GB of data. While Wikidata provides a public SPARQL endpoint, it can only be used for short-running queries. Often, researchers only require a limited range of data from Wikidata focusing on a particular topic for their use case. Subsetting is the process of defining and extracting the required data range from the KG; this process has received increasing attention in recent years. Specific tools and several approaches have been developed for subsetting, which have not been evaluated yet. In this paper, we survey the available subsetting approaches, introducing their general strengths and weaknesses, and evaluate four practical tools specific for Wikidata subsetting – WDSub, KGTK, WDumper, and WDF – in terms of execution performance, extraction accuracy, and flexibility in defining the subsets. Results show that all four tools have a minimum of 99.96% accuracy in extracting defined items and 99.25% in extracting statements. The fastest tool in extraction is WDF, while the most flexible tool is WDSub. During the experiments, multiple subset use cases have been defined and the extracted subsets have been analyzed, obtaining valuable information about the variety and quality of Wikidata, which would otherwise not be possible through the public Wikidata SPARQL endpoint.
Introduction
Wikidata [41] is a collaborative and open knowledge graph founded by the Wikimedia Foundation on 29 October 2012. The initial purpose of Wikidata is to provide reliable structured data to feed other Wikimedia projects such as Wikipedia. Wikidata contains 101,449,901 data items and more than 1.4 billion statements as of 19 January 2023. Wikidata and its RDF and JSON dumps are licensed under Creative Commons Zero v1.0 ,1
Wikidata is a key player in Linked Open Data and provides a massive amount of linked information about items in a wide range of topics. The topical coverage of Wikidata spans from scientific research and historical events to cultural heritage and everyday facts. With its ability to integrate data from multiple sources, Wikidata serves as a powerful tool for knowledge management and data integration. Its structured format and rich linking capabilities make it an ideal resource for machine learning and artificial intelligence applications. Although there is this massive data, most research and industrial use cases need a
In its broadest sense, subsetting refers to extracting the relevant parts from a KG. Considering a KG (regardless of semantics) as a collection of nodes, edges and an associated ontology, a subset can be an arbitrary number of combinations of these three. Thus, in a broad definition, any query graph pattern can be considered a subset, but subsets can include more general cases. Including repetitive graph algorithms such as shortest paths and connectivity [37]. To the best of our knowledge, there is no precise formal definition for submitting accepted by the community [5].
The input of the subsetting process is generally a KG. Over the KG,  queries on the endpoints of a triplestore. This method is suitable for simple and small subsets but has limitations for large and complex subsets. SPARQL endpoints are usually slow and have run-time restrictions. Moreover, recursive data models are not supported in standard SPARQL implementations [20].
queries on the endpoints of a triplestore. This method is suitable for simple and small subsets but has limitations for large and complex subsets. SPARQL endpoints are usually slow and have run-time restrictions. Moreover, recursive data models are not supported in standard SPARQL implementations [20].
The significance of subsets
Having subsets of KGs has many benefits, widening the span from avoiding massive size and computational power issues to data reuse and benchmarking purposes.
See this post: Estimated by Google Cloud Pricing Calculator:
BioHackathon Europe 2021, Project 21: Handling Knowledge Graphs Subsets (group discussions). Notes:
This research aims to collect all available Wikidata subsetting approaches and tools, test their capabilities, and analyse their advantages and disadvantages. The scope is individual, independent, local and arbitrary subsetting, i.e., use cases where users can subset Wikidata locally over any subsetting filters they desire without relying on publicly available servers or datasets. The main reason is that public servers usually apply limitations on the type and run-time of applications. The contributions of this paper are:
A survey of emerging practical knowledge graph subsetting tools (Section 3);
Performance analysis of practical Wikidata subsetting tools (Section 4);
Discussion of the flexibility of practical subsetting tools through tangible Life Science subsetting use cases (Section 5).
This paper first reviews the Wikidata RDF model and the terminology used in the paper in Section 2. In Section 3, a survey of the available methods for subsetting will be presented in detail.
In Section 4, the paper investigates the performance (run-time and extraction statistics) and accuracy (what has been extracted and excluded) of the state-of-the-art subsetting tools. In Section 5, a discussion of the flexibility of the practical tools will be given by going through three Life Science subsetting use cases. Finally, the paper will be concluded in Section 6.
Wikidata RDF model
Core format
The fundamental components of Wikidata are
Underloying software stack: Wikibase and Blazegraph
Wikidata is powered by the
Wikidata uses reification based on intermediate nodes to store contextual metadata, known as  . To access qualifiers, references, and the rank of the statement, the intermediate ‘Statement Node’ must be used, represented with a
. To access qualifiers, references, and the rank of the statement, the intermediate ‘Statement Node’ must be used, represented with a  prefix. This intermediate node can be accessed by the
prefix. This intermediate node can be accessed by the  combined with the same statement property identifier. From the statement node, qualifiers are accessible by the
combined with the same statement property identifier. From the statement node, qualifiers are accessible by the  , references by the
, references by the  , ranks by the
, ranks by the  , the default value-unit with
, the default value-unit with  , and the conversion to the default IRI mapping using
, and the conversion to the default IRI mapping using  . Note that in Wikidata, values can be simple literals (i.e. text or values), IRIs, or complex data types called a
. Note that in Wikidata, values can be simple literals (i.e. text or values), IRIs, or complex data types called a
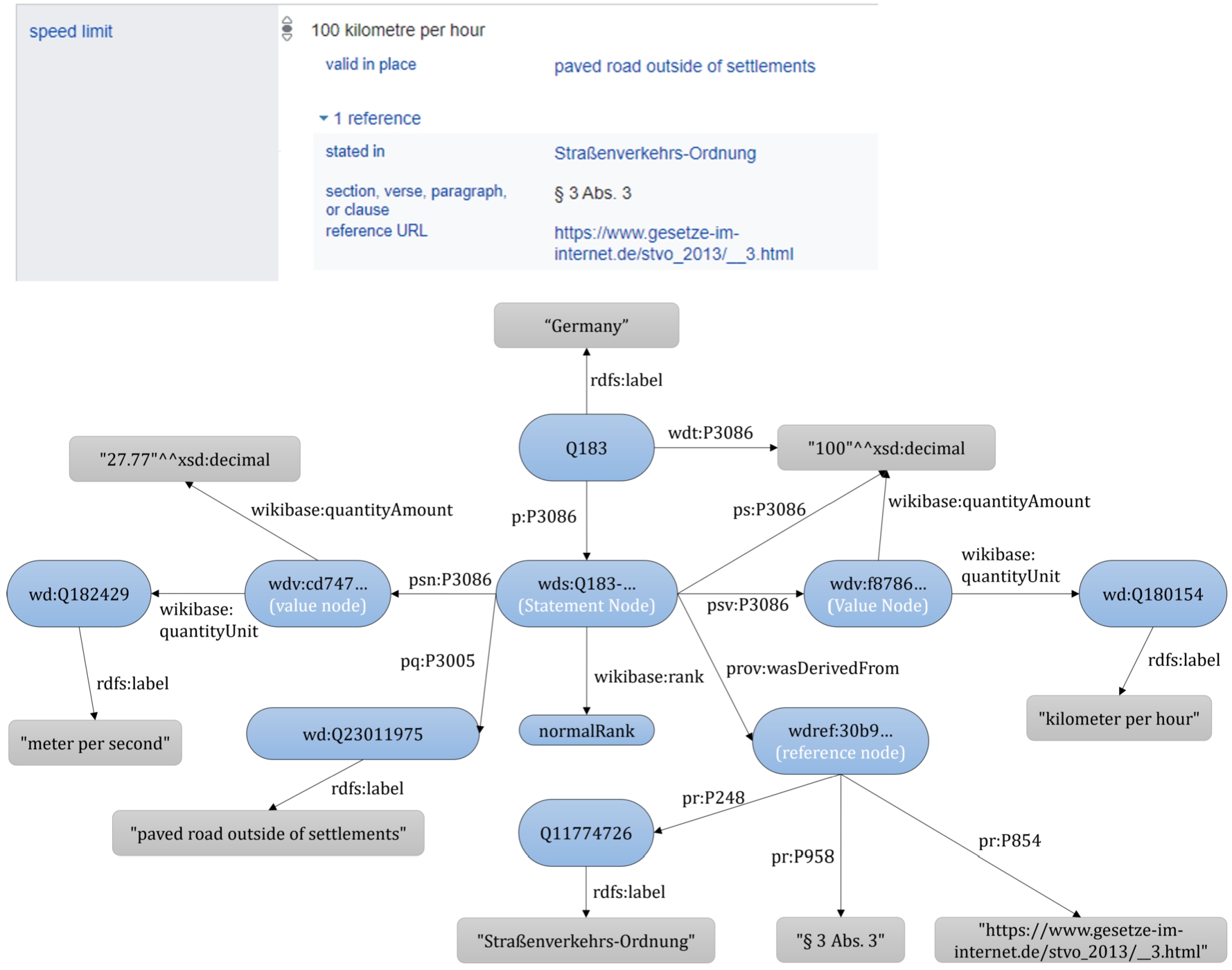
Top: one of the
Subsetting is a recent research problem in KGs. To the best of our knowledge, the early demand for creating a biomedical subset of Wikidata was in 2017 [29], the subsetting discussions in the Wikidata biomedical community were concretely started at the 12th international SWAT4HCLS conference in 2019 by Andra Waagmeester
Matsumoto GitHub:
Mimouni
Henselmann and Harth [15] developed an algorithm for creating on-demand subsets around a given topic from Wikidata, starting from a seed set of nodes and performing multiple SPARQL queries to obtain the desired triples. Their approach can be used to create subsets around topics. However, the authors do not provide use cases or evaluation of their algorithm, thus it is more a theoretical approach than a practical tool. The proposed algorithm and its SPARQL queries are also not compatible with references. Aghaei
Shape Expressions (ShEx) [28] is a structural schema language allowing validation, traversal and transformation of RDF graphs. There are several ShEx validator implementations, e.g., shex.js [35] and PyShex [39], which receive a ShEx schema as the input and validate an RDF graph over it. These validators can keep track of the triples traversed during validation and return the matched triples out (called ‘slurping’), which can be used to define data schemata which could result in extracting a subset. ShEx is a language for validating RDF data, and its evaluators are for checking the shape of the graph against a schema, not for extracting. Although the language has the most flexible way to define subsets, its evaluators’ slurping capabilities are limited as they can not handle the massive size of Wikidata.
WDumper15 Demo:
The flexibility of the ShEx language motivated researchers to develop a specific subsetting tool for Wikidata based on this language. WDSub [22] is a subsetting tool implemented in Scala that accepts ShEx schemata and extracts a subset corresponding to the defined schema from a local Wikidata JSON dump. The extractor part of the WDSub is similar to WDumper, i.e., the WDTK java library. In addition to traditional ShEx schemata in ShExC format, WDSub has its own subsetting language, WDShEx [21], which is a shape expression language based on ShEx and optimized for Wikidata RDF data model. WDSub can produce both RDF and Wikibase-like JSON outputs.
Knowledge Graph Toolkit (KGTK) [16,40] is a collection of libraries and programs to manipulate KGs. KGTK is designed to make working with knowledge graphs easier, both for populating new KGs or developing applications on top of KGs. It is implemented in Python, including a command-line tool for multiple utilities such as importing and exporting Knowledge from various formats (e.g., RDF, CSV, JSON), merging and combining KG data, creating KGs from unstructured sources, querying and analyzing KG data, etc. The fundamental operations in KGTK are importing and querying. KGTK imports massive KGs, converts the data to TSV files, and uses a Cypher-inspired language (called Kypher) to query from these TSV files. In the context of Wikidata, KGTK has been deployed in multiple quality and population-related studies (such as [17,38]). However, its main limitation in Wikibase-driven datasets is not to support indexing of referencing metadata.
Wikibase Dump Filter (WDF) [31] is a Node.js tool to filter and process the JSON data dumps Wikibase, developed and maintained by the Wikimedia Foundation. Similar to WDumper, WDF is an item-based filtering tool, i.e., it applies different filters on items, claims, qualifiers and other Wikibase JSON dump components to create a new dump of
The summary of subsetting tools capabilities. WIP stands for work in progress

Table 1 summarises the capabilities of tools in the following columns:
The first column lists the name of the tool.
The second column lists the output format the tool generates.
The third column shows the required data input format for the tool.
The fourth column lists the language/format used to define a subset.
The fifth column reflects the average hardware and software infrastructure required for using the tool to extract a subset.
The sixth column reflects whether or not a full Wikidata dump download is required for submission.
The seventh column indicates whether or not the tool runs on live data.
The eighth column reflects whether or not the tool is scalable for large subsets or extract subsets of Massive KG.
The ninth column indicates whether or not the tool supports extracting qualifiers.
The tenth column indicates whether or not the tool supports extracting references.
The eleventh column indicates whether or not the tool provides support for graph traversal (i.e. exploring paths between nodes, including cycles to define a subset).
The twelfth column indicates whether or not it is possible to use the tool to perform additional data transformations (e.g., RDF format conversion) without third-party tools once the subset is extracted. The thirteenth column reflects whether the tool provides analytics about the content of the extracted subset, e.g., the number of triples and items. Note that approaches such as the Context Graph or Aghaei et al. are not listed as those are one-purpose and cannot be reused for arbitrary subsetting.
The table shows that no tool provides all positive functionalities. Most of the tools can be run on PCs except SparkWDSub, which is designed for scalability purposes. WDumper is considered a tool that requires no access to the local dump as it is available from an online demo which uses the latest Wikidata JSON dump. None of the practical tools is capable of live subsetting. Instead, they can deal with massive dumps, where ShEx slurping and SPARQL queries fail. Supporting graph traversal is also a challenging feature available in KGTK and SparkWDSub amongst the practical tools (however, SparkWDSub is in the early development stages).
The SPARQL  queries can be considered the most available approach for subsetting Wikidata and other KGs, while regarding independent and local subsetting, they have limitations that exclude them from being a practical approach. The first limitation is defining a subset with
queries can be considered the most available approach for subsetting Wikidata and other KGs, while regarding independent and local subsetting, they have limitations that exclude them from being a practical approach. The first limitation is defining a subset with  queries is time-consuming, as the end-user needs to write the entire graph shape they want to extract. For example, if the end-user defines a subset of Genes, (in addition to the select filters) they should explicitly define what statements, labels, qualifiers, and references should be in the output. Once the scope of the subset gets complicated, specifying the connectivity of the output graph is even more challenging. Such detailed graph patterns can also be outdated very fast as the RDF data is schema-independent; therefore, users should constantly review and modify their queries. Another limitation is the query endpoint. Public endpoints usually apply concrete run-time and query-type limitations, which reduces the capabilities of
queries is time-consuming, as the end-user needs to write the entire graph shape they want to extract. For example, if the end-user defines a subset of Genes, (in addition to the select filters) they should explicitly define what statements, labels, qualifiers, and references should be in the output. Once the scope of the subset gets complicated, specifying the connectivity of the output graph is even more challenging. Such detailed graph patterns can also be outdated very fast as the RDF data is schema-independent; therefore, users should constantly review and modify their queries. Another limitation is the query endpoint. Public endpoints usually apply concrete run-time and query-type limitations, which reduces the capabilities of  queries (for example, users can not extract
queries (for example, users can not extract  triples or write heavy queries with more classes included), and raising a local endpoint (on a Blazegraph instance or other triplestores) returns us to the cost limitations again. Another limitation to using SPARQL
triples or write heavy queries with more classes included), and raising a local endpoint (on a Blazegraph instance or other triplestores) returns us to the cost limitations again. Another limitation to using SPARQL  queries to describe the subsets is the lack of support for recursion so it would not be able to handle the definition of subsets with cyclic data models. While SPARQL
queries to describe the subsets is the lack of support for recursion so it would not be able to handle the definition of subsets with cyclic data models. While SPARQL  queries are a tangible approach for small subsetting use cases, we don’t consider them a practical solution for subsetting.
queries are a tangible approach for small subsetting use cases, we don’t consider them a practical solution for subsetting.
This section is dedicated to an evaluation experiment on the performance and accuracy of the four practical tools: WDSub, WDumper, WDF, and KGTK. Considering the size of Wikidata, the subsetting tools need to extract data in a feasible time. A fast extraction can reduce processing costs and pave the way for regular subset updates and live subset generation. Subsetting tools should also create accurate outputs. Accuracy in this context means the output of a subsetting tool should include all desired statements and exclude any other data. To assess the performance and the accuracy of the practical Wikidata subsetting tools, a unified test on each subsetting tool is performed and the extraction time and the content of the output are reported. The scripts, schemas, and SPARQL queries of this experiment can be found in the GitHub repository of the paper18
In addition to the size of Wikidata, there are other factors contributing to the speed of subset extraction: (i) the number and complexity of filters applied to the input, (ii) the type of the output data (RDF, JSON, etc.), and (iii) the internal operations of the tool. By keeping the input dump and the desired filters fixed, the internal operations run-time is calculated.
Input dump
The Wikidata JSON dump of 3 January 2022 [45] is used as the input to the four subsetting tools. Table 2 shows the details of the input dump. The input dump was downloaded from the Wikidata Database Download page [46].
Details of the 3 January 2022 Wikidata dump used as the data input for experiments
Details of the 3 January 2022 Wikidata dump used as the data input for experiments
The experiment considers a life-science subset of Wikidata as the test use case with the following conditions.
The subset includes all and only ‘ The subset does not include the instances of subclasses. For example, if the tools extract the instances of The subset includes all statements about the items but does not require qualifiers or references.
To measure the accuracy, after finishing the extraction and recording the execution time and the raw volume of the output, we perform the following set of queries on the input (Wikidata dump) and the output of each tool:
Comparing the results of Condition 1 and Condition 2 in the input dump and the output of each tool is a measure of how well the tools extract what they are supposed to fetch. Condition 3 checks the existence of two subclass instances (Operon as a subclass of Gene, and Acid as a subclass of Chemical Compound), aiming to avoid including false positives. The Operon and Acid are arbitrary subclasses; however, operons have an extra semantic relation to genes (an operon is a functioning unit of DNA containing a cluster of genes) and proteins, while acids do not have such extra relation to chemical compounds. In that way, the two subclassing relations can be further compared and the misconfiguration of the tools can be found.
Output format
In this experiment, the output type of WDumper and WDSub is RDF. WDumper creates GZip NTriple files. WDSub creates GZip Turtle files. WDF produces NDJSON files. The output type of KGTK is a TSV file. There are also differences in the size of different RDF formats. The type and the format of the outputs are reported; however, the difference should be kept in mind when comparing the results. The calculated time includes serialization to RDF and the time required to write to disk.
Experimental setup
Host machine
The experiments were performed on a multi-core server powered by 2 AMD EPYC 7302 CPUs (16 cores and 32 threads per CPU), 320 GB of memory, and 2 hard disks: a 256 GB SSD that runs the operating system (CentOS 7 kernel 3.10.0-1160.81.1.el7.x86_64 amd64) and a 6TB HDD that is used for the extraction steps.
Software versions
Table 3 shows the versions of subsetting tools and software used for compiling. All versions were available on 12 November 2022. WDumper has no released version; therefore, the used commit ID is mentioned. All tools except WDF have Docker containers; however, all mentioned versions are cloned and compiled with no need to have root permissions. For KGTK, the repository-recommended binary package in Conda is installed, using pip. To the best of our knowledge, WDSub and KGTK are being upgraded regularly.
Software versions and compiler/interpreter used
Software versions and compiler/interpreter used
A Python script20 It is worth reporting that KGTK v1.5.3 over 32 threads and avoiding deprecated statements has been run and the tool was unsuccessful to return an output after three days of processing.
Note that while each Wikidata JSON dump has an RDF pair dump, these two different serializations are not identical [34]. Therefore the JSON dump is queried directly using the Python parallel program.
The results of running the four practical tools: size and type of the output, the average (Avg.) and standard deviation (STD) of the extraction time, the number of items and the number of statements
The results of running the four practical tools: size and type of the output, the average (Avg.) and standard deviation (STD) of the extraction time, the number of items and the number of statements
Table 4 shows the output detail, results of extraction time, and the total number of distinct items and statements in the output of each tool. The output of WDSub and WDumper is significantly smaller due to compression. The KGTK output is not compressed; however, it is still as small as WDSub and WDumper. It is because other tools extract the entire metadata of the matched item, including labels, descriptions, qualifiers, etc., while KGTK extracts the statement triples only. In its TSV output, KGTK keeps the Q-IDs only and omits any prefixes, which results in light and fast-writing outputs. Note that KGTK can be set to extract other metadata; however, performing this requires additional conditions and filters, which are not necessary for the experimental scenario (see Section 4.1.2) and increases its extraction time. As well, WDumper, WDF, and WDSub can be set not to extract metadata; however, applying such filters enforces unnecessary overhead in their extraction time.
The extraction times show that WDF is the fastest tool. Part of that is because JavaScript is efficient in reading JSON files. The WDF filters are also basic, and parsing the conditions can be done straightforwardly. KGTK is the second fastest tool which benefits from multithreading, providing a high variance of extraction time. KGTK extraction includes two stages: importing Wikidata and the query itself. In these experiments, 40% of the KGTK run-time was spent importing the Wikidata JSON dump and converting it into three TSV files corresponding to nodes, edges, and qualifiers. The rest 60% of the run-time was spent on the query. KGTK creates a graph cache in SQLite format from the edges TSV file once the first query is performed, which significantly speeds up subsequent queries to at most one hour. Thus, most of the query run-time is spent creating the graph cache for the first time. With such a feature, KGTK can be used to compute the graph cache once. Then the graph cache can be shared by Wikimedia or third-party associates for queries. However, in the context of this investigation, since the paper considers autonomous and arbitrary subsetting (and not publicly available servers), the graph cache processing in the run-time is included. Although WDF and WDumper traverse the JSON dump similarly line by line, and WDumper is a compiled tool, WDumper is slower. A part of this slowness is because WDumper serializes the matched JSON blobs to RDF. Also, WDumper can accept more complex filters that create a level of overhead in extraction (regardless of having a simple specification input). The same is true for WDSub. The RDF serializer in WDumper and WDSub is the same; however, the WDSub filtering system (based on ShEx) can parse quite complex filters at the SPARQL level, which creates a massive overhead. WDumper also has a better level of multithreading than WDSub.
Comparing the number of extracted items and statements shows that KGTK has the least number. The reason behind the higher ratio of missed items and statements in KGTK output is not clear, but it can be hypothesized to be due to the greater complexity of indexing and query procedures in KGTK compared to other tools, a higher likelihood exists for skipping more blobs during intermediate steps due to their un-parsability. The number of extracted items and statements in WDF and WDumper is identical, although this identicality is coincidental as these numbers are the distinct add-up of four different classes. The disaggregated statistics, as discussed in Section 4.4, show that these tools extract a different number of instances in each class.
Accuracy test results of the four tools
Accuracy test results of the four tools
Table 5 shows the result of accuracy test queries on the input dump and each tool separately. In the Condition 1 column, the number of instances of each class can be seen. Compared to the input dump, all tools missed extracting some Q-IDs except WDF. The WDF filter matching process is the simplest among the available tools. It involves scanning the input dump line by line, with each line containing a JSON blob corresponding to a Wikidata item. The filters provided are then applied to the values within each JSON blob, and if a successful match is found, the entire blob is returned. Moreover, the number of extracted statements matches the input dump, highlighting the exceptional accuracy of WDF compared to other tools. The ratio of the missing items in other tools is less than 0.05%, and the ratio of missing statements is less than 0.75%. From 1,196,532 gene instances in the input dump, WDSub did not extract 44, and WDumper and KGTK did not extract 29 gene instances. Although the rate is acceptable, a 100% accuracy is expected for this task. Reviewing the gene instances items that are present in the input dump but are not in the outputs of tools shows that the 29 missed items in KGTK27 List of missed gene instance Q-IDs: List of missed gene instance Q-IDs: List of missed gene instance Q-IDs:
Analysis of the JSON blobs for some missed instances, such as See Line 111 of file ‘item-Q418553-found.json’ in [6] and Line 40 of file ‘item-Q29718370-found.json’ in [6] – accessed 8 Jun 2023. See Line 11 of file ‘item-Q29685684-found.json’ in [6] and Line 11 of file ‘item-Q29678017-found.json’ in [6] – accessed 8 Jun 2023. List of duplicated gene instance Q-IDs: See Lines 407–481 of file ‘item-Q418553-found.json’ in [6] – accessed 10 June 2023.
Choosing amongst the available subsetting approaches depends on the task at hand. The methods introduced in Section 3.1 are single-purpose and usually cannot be reused to create any arbitrary subset. Amongst the practical tools (Section 3.2), the performance and accuracy evaluation showed that WDF has the fastest and most accurate performance; however, this tool is not flexible in defining subsets. This problem also exists in WDumper. In these two tools the inclusion and exclusion of items, statements, and contextual metadata can be defined, there is no possibility to make a connection between these conditions. For example, disease instances and chemical compound instances can be extracted together; however, if only the chemical compounds related to the extracted diseases are needed, this joined KG cannot be extracted with these tools.
KGTK and WDSub offer much higher flexibility due to their subset-defining structure derived from graph query languages. KGTK extracts data after a round of indexing relatively fast; however, in the context of Wikidata lacks indexing references, which is a major drawback. WDSub has the most flexible subset-defining structure in the Wikidata ecosystem and is reasonably accurate; however, response time is slow and still in its early stages of development (as of June 2023).
Flexibility evaluation
The extent to which each tool supports common subsetting workflows is crucial. While Section 4 focuses on the performance and accuracy in a single subsetting scenario, it should be noted that tools offer varying degrees of support for various subsetting tasks, depending on their functionalities and features. The flexibility experiments showcase the range of potential applications and highlight the appropriateness of each tool for specific subsetting requirements. This section investigates more diverse subsetting tasks involving different parts of the Wikidata data model supported by each evaluated tool, thereby providing a more comprehensive understanding of their practical applicability. The first use case is the Gene Wiki project evolution from 2015 to 2022, the second is genes names and descriptions in four languages, and the third is instances of chemical compounds that are referenced with
Gene wiki evolution
The Gene Wiki Project [47] focuses on populating and maintaining Wikidata as a central hub of linked knowledge on genes, proteins, diseases, drugs, and related Life Science items. This project is one of the most active WikiProjects in terms of human and bot contribution [4]. The project is initiated based on a class-level diagram of the Wikidata knowledge graph for biomedical entities, which specifies 17 main classes [8]. The Wikidata WikiProject has extended the classes into 24 item classes.
The Gene Wiki evolution experiment aims to (i) capture a subsetting schema where the participating classes have connectivity to each other, and (ii) show the change in the amount of data instances from the early years of Wikidata. WDSub is deployed to extract the Gene Wiki subsets containing instances of the 20 classes pictured in [47] UML class diagram. The steps are:
Creating a ShEx schema that represents the data model depicted in [47]. The ShExC format of the defined shapes is in Appendix A; Downloading the Wikidata JSON dumps from 2015 to 2022 (exact dates are in Appendix B) which are available at Internet Archive;34 Deploying WDSub to create a subset from each dump.
A SPARQL query script is then run that counts the number of each item for each shape (class) and each link between shapes. Table 6 shows the number of instances for each class.
The number of instances for each gene wiki class from 2015 to 2022 and the number of instances on the live Wikidata query service (queried on 22 December 2022)
The first attention-drawing point is the variation in the number of instances in different classes. The taxon, gene, protein and chemical compound classes have the highest number of items, such that more than 97% of the items in all the investigated dumps are instances of these four classes. Part of this heterogeneity is due to the nature of the abundance of classes. For example, the number of genes should be more than diseases, but it is not clear why in some classes the number of instances is so low, e.g., the number of anatomical structure instances seems less than expected. The number of instances in all classes except the biological process, cellular component, disease, molecular function, sequence variant, and symptom has increased continuously from 2015 to 2022. In addition to having the largest amount of data in all dumps, the data growth acceleration in the taxon, gene, protein, and chemical compound classes is also more than the other classes from 2015 and 2022. In all exceptional classes above, the peak point belongs to dump 2020. Then, the number of instances decreases in 2021 and 2022, reaching the previous 2020 level in 2023, where the Wikidata SPARQL endpoint has been queried. The reason for this behaviour is not clear. It has been hypothesized that the number of instances was raised due to inaccurate bot activities in 2020, which was restored during human curations in the following two years and reached the same level again due to more accurate bots. Another observation is the low number of genes, proteins and chemical compound instances before 2017. The Gene Wiki WikiProject started and began populating data in 2015. These classes are the main focuses of the Gene Wiki community data population. It is found that the low number of instances in the 2015 and 2016 dumps is not due to the lack of A-Boxes, but due to the lack of
The extracted subsets in this experiment can also be constructed by other practical tools of Section 3.2. The definition of these subsets in WDSub is based on writing a shape corresponding to each class containing the properties defined in the class diagram [47]. Such filters can be implemented by all other tools as well. In WDumper and WDF, one can simply write the corresponding filters based on the value of the parameters of the mentioned properties (the properties inside a Shape will be logical AND together). However, in some definitions, WDumper and WDF can not imitate the WDSub definition exactly. The reason for this is that in WDSub any number of relationships amongst shapes can be defined. For example, the  class in Appendix A is related to the form
class in Appendix A is related to the form 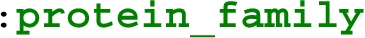 class via
class via  property. Now suppose the
property. Now suppose the  operator in line 34 is replaced with a
operator in line 34 is replaced with a  . At extraction time, WDSub will not extract any active site instances that are not connected to at least one instance of a protein family. Unfortunately, such filtering and connections are not possible in WDumper and WDF. In these two tools, only one specific value can be defined for a property filter; it is not possible for the value to be of a specific class or related to other conditions (in WDumper, there is a possibility to define a condition saying a value should have existed, whatever that value is). KGTK can establish any relationship between conditions as its Kypher definition system is based on Cypher query language and has definition flexibility similar to ShEx. The extracted subsets can be found on Zenodo [24].
. At extraction time, WDSub will not extract any active site instances that are not connected to at least one instance of a protein family. Unfortunately, such filtering and connections are not possible in WDumper and WDF. In these two tools, only one specific value can be defined for a property filter; it is not possible for the value to be of a specific class or related to other conditions (in WDumper, there is a possibility to define a condition saying a value should have existed, whatever that value is). KGTK can establish any relationship between conditions as its Kypher definition system is based on Cypher query language and has definition flexibility similar to ShEx. The extracted subsets can be found on Zenodo [24].
Using the ShEx schema in Appendix C, a subset of Genes and Taxons instances from 2015 to 2022 is created, considering instances which have both labels and descriptions in English, Dutch, Farsi, and Spanish. Item instances that do not have a label or description in one of these four languages should not be extracted (aliases condition is considered with a  operator, which means that instances with zero aliases in the four languages can be in the subset).
operator, which means that instances with zero aliases in the four languages can be in the subset).
The total and language seperated number of instances for gene and taxon class from 2015 to 2022 and the number of instances on the live Wikidata query service (queried on 14 February 2023)
The total and language seperated number of instances for gene and taxon class from 2015 to 2022 and the number of instances on the live Wikidata query service (queried on 14 February 2023)
Table 7 shows the number of instances separated by label, description, and alias languages along with the total number of extracted items. The difference between the number of labels, descriptions and aliases can also be seen. In general, English aliases are more than labels, which shows that on average each item has more than one English alias. By comparing between languages, it can be seen that the amount of labels, descriptions, and aliases in Farsi is lower than in other languages. This is more obvious in Genes compared to Taxons. In Spanish and Dutch, the number of labels and descriptions are close, which shows that wherever there is a label for this language, a description has also been added (note that labels and descriptions are usually added once for each language while aliases are more than one). While having fewer Farsi labels and aliases can be justified by the lack of proper translation, having fewer descriptions is due to the fewer Farsi-speaking participants (or their limited activity in Genes and Taxons). The low amount of data in Genes before 2017 which is explained in Section 5.1, can be seen here again. As the table shows, counting the number on Wikidata Query Service has been timed out in multiple taxon queries.
Subsetting on labels and comments can also be done by KGTK. KGTK and WDSub can define conditions even on the values of the label, e.g. define a shape (in KGTK a Kypher term) with a label condition the value specified to  and extract all entities with the name John Smith from Wikidata. Filtering labels and comments is not possible with this flexibility in WDF and WDumper. In both WDF and WDumper, users can choose whether to skip labels and textual metadata (such as descriptions) along with the selected item. It is also possible to extract labels and comments in their specified languages (and not all languages). However, these options are considered post-filters, i.e., items are first selected based on property-based conditions, and then textual metadata can be ignored or kept on the selected items. Another limitation is that this option can be deployed either on all extracted items or none of them, e.g., it is not possible to extract a group of items with English labels and another group with Farsi labels. Initial selection based on language or value of a label/comment is not doable in WDF and WDumper. The extracted subsets can be found on Zenodo [25].
and extract all entities with the name John Smith from Wikidata. Filtering labels and comments is not possible with this flexibility in WDF and WDumper. In both WDF and WDumper, users can choose whether to skip labels and textual metadata (such as descriptions) along with the selected item. It is also possible to extract labels and comments in their specified languages (and not all languages). However, these options are considered post-filters, i.e., items are first selected based on property-based conditions, and then textual metadata can be ignored or kept on the selected items. Another limitation is that this option can be deployed either on all extracted items or none of them, e.g., it is not possible to extract a group of items with English labels and another group with Farsi labels. Initial selection based on language or value of a label/comment is not doable in WDF and WDumper. The extracted subsets can be found on Zenodo [25].
This section deploys references as filters and extracts those chemical compound instances that their This schema is designed to extract all This schema extracts all Counts those chemical compound instances that their Counts the number of chemical compound instances in general.36
The property
Table 8 shows the number of chemical compound instances obtained from performing the two queries on the 2015 to 2022 subsets. In the last column, it can be seen the number of referenced and not referenced chemical compound instances on Wikidata. As the results show, WDSub accurately excludes not-referenced chemical compound instances in extraction. In all dumps, the number of referenced instances fetched by the referenced query (Query1) in the general subset (extracted using Schema 2) is equal to the total number of instances (fetched by the general query, Query2) in the referenced subset (extracted using Schema 1). The only inconsistency is in the column of dump 2021, where there are 17 referenced instances in the subset extracted by the general schema, while there are 16 instances in the subset extracted by the referenced schema. In other words, there is one referenced instance in the input dump which is not extracted by WDsub. However, this is not an unexpected missing item. The missed item is
 qualifier to open the
qualifier to open the  to Line 11 of the Schema D.1 solves this inconsistency. Overall, the number of instances referenced by the
to Line 11 of the Schema D.1 solves this inconsistency. Overall, the number of instances referenced by the
The number of referenced and not referenced chemical compound instances in Wikidata subsets from 2015 to 2022 and the Wikidata query service (queried on 14 February 2023)
In this paper, the problem of subsetting in Wikidata was reviewed. As a continuously edited KG, Wikidata has a massive amount of data which cannot be queried from the SPARQL endpoint in all cases. Its weekly RDF and JSON dumps are maintained for a short period of time and hosting a Wikidata dump is costly. On the other hand, research and applications may need a specific scope of its data. Subsetting provides a platform to extract a dedicated part of the data from Wikidata, reducing the overall cost and facilitating the reproducibility of experiments.
The paper surveyed all available subsetting approaches over Wikidata and other KGs and explained their advantages and limitations. In the context of Wikidata, four subsetting approaches are distinguishable as practical subsetting tools that can be deployed to extract a given defined subset: WDSub, WDumper, WDF, and KGTK. The performance, accuracy, and flexibility evaluations were then established over these four practical tools by defining several subsetting use cases. The results show that in terms of performance (i.e., the speed of extraction), WDF is the fastest tool and it can extract a subset in less than 4 hours. In terms of accuracy (i.e., extracting what is defined exactly, not more or less) the results show that WDF extracts all items and statements exactly as they are present in the input dump. The ratio of missed items is less than 0.05% all tools missed less than 4% of items and that can be justified by the inconsistencies and syntax errors in the input dumps. In terms of flexibility (i.e., how much the tool allows the designer to define complex subsets on different parts of the Wikidata data model), three use cases have been defined and several subsets have been extracted from Wikidata dumps from 2015 to 2022. At first, a subsetting on different classes of the Gene Wiki WikiProject was performed and all tools supported such a subsetting. Then the subsets of genes and taxons were extracted based on having English, Spanish, Farsi, and Dutch labels and comments, which WDSub and KGTK supported such filtering. In the end, subsets of referenced chemical compounds were extracted and only WDSub was able to perform filters on references. The flexibility tests show that the most flexible tool for subsetting is WDSub, mainly because of its defining language which is ShEx and has the flexibility of SPARQL queries. During the subsetting, valuable information was gained about the amount of data in Wikidata from 2015 to 2022.
In KG subsetting, many open questions and future work remains. The first open question is subsetting other KGs, such as DBPedia, where the vocabulary is different and the dumps are not in JSON. There are also many massive collections of data supporting RDF, such as Uniprot and PubChem that can be the subject of subsetting. Future work also includes building more flexibility and performance with one tool. WDSub is the most flexible tool but when you have flexibility, your filters take a longer time to be applied on the input dump items. SparkWDSub [23] is an under-development subsetting tool for Wikidata based on WDSub, which implements graph traversal for subset creation. To improve the speed, SparkWDSub uses the Apache Spark platform to distribute the computation. This tool is in the initial stages of development. Live subsets are the other future path. In this study (as well as in other related projects) several topical subsets have been extracted for which reusability is one of the main features. Over time with the new edits coming, the gap between these subsets and the corresponding data in Wikidata will increase. This gap can be reduced by repeating the subsetting process regularly, and by reducing the interval to an acceptable level (e.g., one day), end users can reach practically live subsets. A better solution is not to spend the extraction time for each repetition, instead, to generate the subset and apply the edits in real-time by establishing an active link between the Wikidata database and the subset. The main challenge in this task is hosting issues and the fact that Wikidata does not have a public API for establishing active links to the best of our knowledge. Subsetting suffers from not having proper documentation for tools, definitions, and use cases. It is essential to aggregate and document all subsetting definition efforts as a training wiki, in which users can effectively learn and define desired subsets in a reasonable time. Having such a wiki, further performance, accuracy, and flexibility tests can be established in different fields.
Footnotes
Acknowledgements
This paper has progressed in several hackathons and tutorials of the ELIXIR BioHackathon-Europe series and SWAT4HCLS, and we would like to thank the organizers and participants. Suggestions and intellectual contributions of Dan Brickley, Lydia Pintscher, Eric Prud’hommeaux, Thad Guidry, and Filip Ilievski are greatly appreciated. This project has benefited from part of the following research grants: project PID2020-117912RB, ANGLIRU: Applying kNowledge Graphs to research data interoperability and ReUsability. The Alfred P. Sloan Foundation under grant number G-2021-17106 for the development of Scholia. The project R01GM089820 from the National Institutes of General Medical Sciences.
Gene wiki ShEx
The ShExC shape expressions that is used to extract Gene Wiki subsets via WDSub is as follow:
Wikidata dumps dates
The exact dates, size, and download URL of Wikidata dumps used in the flexibility experiments
| Dump | Exact date | Size | Download URL |
| 2015 | 2015-06-01 | 4.5 Gb |
|
| 2016 | 2016-06-13 | 7.19 Gb |
|
| 2017 | 2017-08-21 | 15.7 Gb |
|
| 2018 | 2018-01-15 | 26.48 Gb |
|
| 2019 | 2019-01-21 | 48.14 Gb | https://archive.org/download/wikibase-wikidatawiki-20190121/wikidata-20190121-all.json.gz |
| 2020 | 2020-11-02 | 83.94 Gb |
|
| 2021 | 2021-05-31 | 93.93 Gb |
|
| 2022 | 2022-06-30 | 107.66 Gb |
|
Genes + taxons labeling and commenting ShEx
The ShExC shape expression that is used to extract Genes and Taxons subsets via WDSub based on labels, descriptions, and aliases in four languages is as follow:
