Abstract
Annotations enrich text corpora and provide necessary labels for natural language processing studies. To reason and infer underlying implicit knowledge captured by labels, an ontology is needed to provide a semantically annotated corpus with structured domain knowledge. Utilizing a corpus of adverse event documents annotated for sepsis-related signs and symptoms as a use case, this paper details how a terminology and corresponding ontology were developed. The Annotated Adverse Event NOte TErminology (AAENOTE) represents annotated documents and assists annotators in annotating text. In contrast, the complementary Catheter Infection Indications Ontology (CIIO) is intended for clinician use and captures domain knowledge needed to reason and infer implicit information from data. The approach taken makes ontology development understandable and accessible to domain experts without formal ontology training.
Keywords
Introduction
Many natural language processing (NLP) studies rely on annotated corpora to create models for text classification, information extraction, named entity recognition, question answering, summarization, and text generation. Often, semantic annotation is done to capture domain knowledge within the text. Annotated corpora are frequently generated by annotators based on an annotation guideline, which provides the standard and rules for how to label text using specified terms. This annotation guideline is usually similar to a terminology, unless the corpus was annotated using an ontology. To further enrich an annotated corpus by capturing, reasoning, and inferring the underlying associated domain knowledge, an ontology is needed.
To demonstrate semantic annotation terminology and ontology development, the use case is based on clinician-presented needs for identifying sepsis from adverse events (AEs). Improperly cared for peripheral intravenous catheter (PIVC) medical devices can lead to unwanted and unintentional events that harm patients, such as AEs like phlebitis, bloodstream infections (BSIs), and sepsis. However, PIVCs are poorly documented in clinical records because of routine use among inpatients, and sepsis is also poorly documented outside the intensive care units (ICUs). The lack of explicitly documented concepts makes it challenging to directly detect and annotate PIVC-related phlebitis, BSI, and sepsis for quality surveillance to improve care. Thus, indications for the presence of PIVCs, phlebitis, infections, and sepsis are annotated instead. Those annotations are structurally preserved as a terminology, and the clinical knowledge required to reason about the indications is represented in an ontology. Additional details for the use case are provided in Section 2.
This paper provides a detailed and concrete description of the methodology utilized for ontology development of an annotated corpus based on a use case from the clinical domain. The main contributions presented are:
Describing the development process for constructing a terminology that can represent an annotated corpus. Specifically, a terminology for indexing annotated AE documents. Presenting the development process for the terminology’s corresponding ontology, which represents domain knowledge and allows inference of implicit knowledge in a specific domain. The corresponding ontology in the use case represents clinical domain knowledge specifically for annotated catheter-related and infection-related signs in AE documents. Releasing a terminology and ontology that can be applicable to identifying and reasoning about sepsis in an AE corpus.
This paper significantly extends the papers [66] and [67], by adding an ontology with instances and including evaluation of the correctness and ability to answer competency questions. In addition, instances from the annotated corpus in [67] were added into the terminology, the terminology was evaluated using competency questions, and an ontology was developed to answer competency questions with clinical knowledge.
Based on the presented use case in Section 2 and objective in Section 2.2, an Annotated Adverse Event NOte TErminology (AAENOTE) and corresponding Catheter Infection Indications Ontology (CIIO) were developed. Section 5 details the terminology construction and development process to represent annotated documents, and Section 6 presents the results and evaluations for the AAENOTE. To address shortcomings in the terminology for annotated documents, Section 7 describes the ontology development process for domain knowledge representation of documented content. Additionally, Section 8 presents the results and evaluations for the CIIO. Finally, Section 9 discusses the findings, limitations, representations, accessibility, and utility.
Use case background and motivation
Sepsis from peripheral intravenous catheter-related phlebitis and infection adverse events
As the most commonly used medical device in hospitals, PIVCs are inserted into the peripheral vein to administer intravenous (IV) fluids, IV medications, and blood transfusions [1]. Improper management of PIVCs or the infusions connected to the PIVC can lead to phlebitis, which is either infectious, mechanical, or chemical inflammation of the vein [12,23,47]. Independent of cause, all PIVC phlebitis share many symptoms like redness and swelling near a patient’s infusion insertion site for infectious, mechanical, or chemical phlebitis, making it difficult to distinguish. Furthermore, all PIVC-related phlebitis causes AEs like significant pain, PIVC failure that delays treatment, and compromises future venous access. Infectious phlebitis may lead to BSI due to: 1. migration of bacteria at the insertion site, 2. bacteria migrating through the catheter tract or catheter hub, 3. contaminated infusate, or 4. bacteria from an existing infection in the bloodstream attaching to the catheter [69]. BSIs can potentially cause sepsis and occur when bacteria enter the bloodstream [26].
Even though PIVCs are frequently used, they are routinely not documented in clinical records [1]. Moreover, sepsis is also poorly documented outside the ICUs [49]. This lack of documentation makes it challenging to perform retrospective and real-time systematic quality surveillance of PIVC-related phlebitis or BSIs to identify learning opportunities for improving PIVC care to lower phlebitis-related, BSI-related, and PIVC-related AE incidents. Hence, AE reports or documents, which are customarily used to report PIVC failures, were selected as the clinical text for this project. To capture documented observable patient states and infer underlying knowledge of PIVC-related phlebitis or BSI from clinical text, an ontology that models clinical knowledge representation and reasoning is necessary.
Use case objective
The use case objective was to develop a model for representing and reasoning about PIVC-related BSIs in the unstructured free-text of AE reports, describe the development process, and discuss the discoveries and limitations. From the research question “is there a connection between BSIs and PIVCs at the hospital?”, competency question requirements for an ontology representing and reasoning about PIVC-related BSIs were identified by clinicians as follows:
Does patientA have phlebitis, and was it infectious, chemical, or mechanical phlebitis? Does patientA have an infection? Does patientA have a BSI? How many patients have an infection or BSI? Which patients have sepsis? Does patientB have a catheter? Does patientB have a PIVC? How many catheters does patientB have, where are they, and why does patientB need them? Does patientC have an infection and catheter? If so, was patientC’s infection associated with a catheter?
Related work
Annotation, tagging, and ontologies for natural language processing
Many studies focus on the relationship between annotation, tagging, and ontologies for NLP development. These studies include annotating corpora, NLP extraction, and classification tasks. Below are some studies that have shared their findings, issues, and possible solutions.
Annotated using the Uberon multi-species anatomical ontology [34], the Colorado Richly Annotated Full-Text (CRAFT) Corpus is a resource for NLP development which is also semantically annotated with concepts from eight Open Biomedical Ontologies (OBO) and terminologies [4,6]. However, while annotating with the OBOs, they discovered that the OBOs are not developed for annotation because there are overlapping terms within the different OBOs, context-specific definitions, and semantic ambiguities. Additionally, some OBOs do not follow the OBO Foundry principle of using relations from the OBO Relation Ontology (RO) [57] to link concepts [5]. Therefore, to improve OBOs for semantic annotation of biomedical documents, the researchers proposed desirable ontology implementations such as, but not limited to, integrating overlapping OBOs terms, resolving ontology-specific ambiguities, and expanding relations [5].
A study comparing how anatomy ontologies are used for annotations discovered annotation and ontology issues [61]. Annotations from three public datasets were compared to anatomical terms in the Foundational Model of Anatomy (FMA) [50,51] and Uberon [34] ontologies using the Zooma and Ontology Mapper software tools. Manual and semi-automated preprocessing were done to normalize terms, but there were few matches between the ontologies and annotations, mainly because of strict matching. Additionally, the user-provided annotation labels resulted in mismatches, such as annotating a phrase with multiple ontology terms or using an abbreviation or adjective for an anatomical part instead of the anatomical ontology term. Ontology issues include missing anatomical synonyms used by the annotators and differing anatomical terms in the ontologies because the ontologies are designed for different purposes and made by different design decisions. The study concluded that mapping terms to an ontology requires a large amount of time, effort, and manual curation. Furthermore, an ontology’s design decisions and scope will affect users trying to match annotations to an ontology, and ontologies must be used to understand their potential.
The Unified Medical Language System (UMLS) is a collection of standard biomedical terminology [8], and it has been used to process text by extracting concepts, relations, and knowledge (i.e., link or annotate text with standard terminology) [2]. A software capable of finding and linking biomedical text to terminology concepts in the UMLS Metathesaurus is MetaMap [3]. However, the developers of MetaMap mention that improvement is required for detecting similar names, acronyms, and abbreviations and resolving ambiguities by possibly distinguishing concepts using word sense disambiguation.
In [37], an overview of studies using knowledge bases for entity coreference resolution were discussed. Among those studies was the OntoNotes project, which annotated a multilingual corpus for different levels of semantic structure in the text [25,44,45]. One of the annotation levels includes linking OntoNotes word senses to the Omega ontology [25,46,68]. Near-synonymous word sense pools were created by specialists who grouped sense distinctions from WordNet and dictionaries based on similar definitions. This enables machines to automatically tag senses more accurately and improves inter-annotator agreement due to difficulties determining WordNet distinctions directly in the text [68]. Before each sense pool was linked to a concept in the Omega ontology [42], each sense pool was verified by machine and humans [68].
Ontology development methods and evaluation
There are many ontology development methods, such as: the Enterprise ontology’s Uschold and King [62], the TOronto Virtual Enterprise (TOVE) ontology’s Grüninger and Fox [20], METHONTOLOGY [14], the On-To-Knowledge Methodology (OTKM) [60], and NeOn [59]. Of those methods, Uschold and King [62] and Grüninger and Fox [20] follow a sequential sequence of phases, whereas METHONTOLOGY [14], OTKM [60], and NeOn [59] are iterative. Whether sequential or iterative, the previously mentioned methods and 2 reviews [10,43] have shown that ontology development typically includes the phases: specification, conceptualization, formalization, implementation, and maintenance. During those phases, the knowledge acquisition, evaluation, and documentation phases also commonly occur either as a separate phase or concurrently with other phases. Appendix A provides a summary of the methods for each phase.
During and between phases, ontology evaluation judges an ontology’s content to a reference, such as requirement specifications, competency questions [20], or the real-world [18,19]. Evaluation includes: (1) verification that the ontology has the correct informal natural language definition and formal ontology language definition, and (2) validation that the ontology represents the world it was created for [18,19]. In theory, there are many criteria for evaluation, but in practice, most studies only use the expressiveness and practical usefulness criteria [11]. Expressiveness is the number of competency questions answerable by the ontology [11,20,38], and practical usefulness is the number of problems an ontology can be applied to [11,38].
Relevant ontology resources
Overview of relevant resources for this study
Overview of relevant resources for this study
There lacks an ontology specifically for sepsis-related BSI, infection signs, anatomical locations, medical devices, and procedures. However, pre-existing ontologies can contain relevant concepts. For example, the Infectious Disease Ontology (IDO) [27] has sepsis and hospital-acquired infection entities. Sign and symptom entities are present in the Ontology for General Medical Science (OGMS) [39], and vital sign entities exist in the Vital Sign Ontology (VSO) [17,64]. Anatomical locations can be described using anatomical entities of the Foundational Model of Anatomy Ontology (FMA) [15,51] and anatomical spatial location descriptor entities from the Biological Spatial Ontology (BSPO) [7]. Because AE reports are used, the adverse event entities in the Ontology of Adverse Events (OAE) [21,40] might also be relevant. Furthermore, relationship object properties in the Open Biological and Biomedical Ontology (OBO) Relation Ontology [48,57] could be used to link different entities together to capture more information.
In addition to ontologies, there are also potential relevant terminologies and taxonomies. For example, there are different procedure, medical device, and catheter terms in the National Cancer Institute Thesaurus (NCIT) [35,36]. Potential relevant standardized nursing practice language is found in the International Classification for Nursing Practice (ICNP) terminology [13], Nursing Interventions Classification (NIC) taxonomy [9], and NANDA International Nursing Diagnoses Classification taxonomy [22]. Furthermore, infusion phlebitis-related information can be obtained from the 1998 Visual Infusion Phlebitis Scale [30] and the 2021 Infusion Therapy Standards of Practice Updates [28]. Concepts or terms from these resources can be used to expand the ontology if deemed necessary by ontology users. The relevant ontologies, terminologies, taxonomies, and clinical guidelines can be found in Table 1.
Synthetic dataset
Documents for annotation are from an AE synthetic dataset. The documents are based on unstructured free-text AE notes within the extracted AE reports from the electronic incident reporting system at St. Olavs hospital, Trondheim University Hospital in Trondheim, Norway, between September 2015 to December 2019 [67]. The synthetic dataset contains 100 AE notes or documents manually created and verified by a nurse to ensure clinical data is anonymized. The Norwegian Regional Committees for Medical and Health Research Ethics (REK) has granted ethical approval to use AEs in this paper (approval no 2018/1201/REKmidt, 26814).
Annotated synthetic dataset
The synthetic documents were annotated by 8 annotators with clinical backgrounds over 4 annotation sessions [67]. Each annotator annotated 10 documents in session 1 and 20 documents in the remaining 3 sessions (i.e., 70 documents annotated over 4 annotation session). This resulted in 560 annotated synthetic AE documents, as shown in Fig. 1 (i.e., 8 annotators * 70 annotated documents over 4 sessions = 560 total annotated synthetic AE documents). In each annotation session, annotators followed the annotation guideline and used the Brat rapid annotation tool (BRAT) [58] to annotate the documents. Then, documents were evaluated and manually screened to identify ambiguities for revising the annotation guideline. This process was repeated 3 additional times with a new set of documents and a revised annotation guideline.

From unannotated documents to an annotated corpus for populating instances in a terminology. 100 synthetic adverse event (AE) unstructured free-text documents were manually generated. Those synthetic documents were annotated by 8 annotators over 4 annotation sessions using revised annotation guidelines. Each annotator annotated 70 documents (i.e, 10 documents in session 1 and 20 documents in the remaining 3 sessions) to produce a total of 560 annotated synthetic AE documents.
An annotation guideline was developed based on the clinical research question: Is there a connection between BSIs and PIVCs at the hospital? Discussions with nurses provided insight into how catheters can be distinguished explicitly by the name or implicitly based on the anatomical insertion site or procedure mentioned. This formed into four domain-specific What are the different signs of infections, specifically for BSIs, sepsis, or infected PIVCs? What are the signs for different types of catheters? Where are the anatomical insertion sites of catheters? What procedures, interventions, and activities can be related to catheter use?
Answers to domain-specific questions were then sorted into the following 7 main categories:
All categories except
A preliminary annotation guideline was created using the seven categories and four relationships. Annotation guidelines from each of the 4 annotation sessions are available online1
In annotations, categories are known as entities for labeling a span of words or phrases. Whereas in ontologies and terminologies, categories are known as classes. To separate the annotation guideline from the terminology and ontology, the annotation categories and entities are in
Design decision for annotations
The terminology was developed using the bottom-up approach based on the annotation guideline refinement process from 4 iterations. Competency questions were not used to create this terminology. This terminology is meant to assist annotators who want to label text and allow users interested in performing downstream analyses to adjust the granularity of labels. The objective is solely to represent the annotated corpus and provide structure to the terminology used by annotators. Thus, included individuals are based on concrete examples from the annotated corpus. A simplified example of how annotation labels in annotated documents are added to the terminology as individuals is provided in Fig. 2.
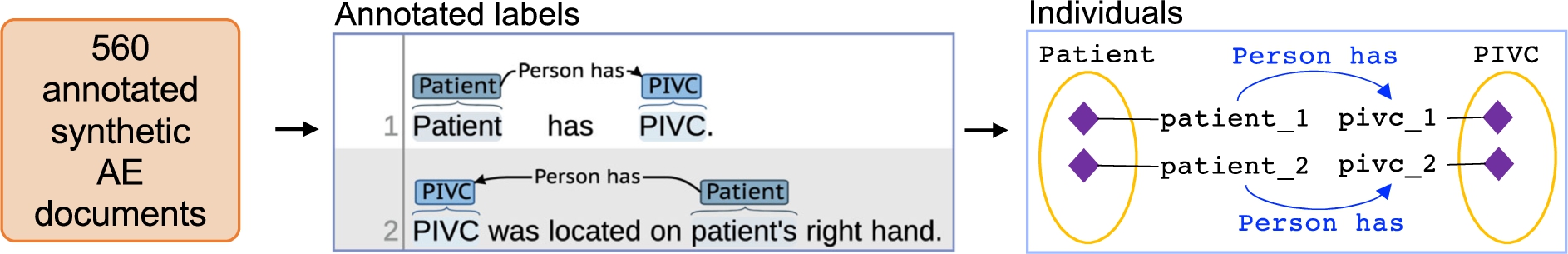

Using an annotated corpus to populate individuals in a terminology. Each of the 560 documents were translated into an individual, and each label within a document was also translated into an individual. In the simplified example, an annotated document has 2
Instead of reusing and re-defining existing ontologies, it was easier and simpler to develop a terminology based on what is documented in the data. For instance, although the FMA contains relevant anatomical parts, the ontology was too complex and detailed to be incorporated easily into the terminology to fit the use case’s purpose. Additionally, the purpose was to include only concrete items documented in the terminology and not provide terminology for all existing items. By opting to simplify the terminology, it was easier to build the terminology directly based on the annotation guideline and then modify the terminology to incorporate feedback from discussions with clinicians.
The categories, attributes, and relationships in annotation guidelines described in Section 4.3 correspond to classes, data properties, and object properties in terminologies (Table 2 and Fig. 3).
Convert annotation guideline to terminology
Convert annotation guideline to terminology

Terminology development. The annotation guideline from the fourth annotation session was converted into a terminology. Annotation categories were converted into ontology classes, relationships into object properties, and attributes into data properties. Then the individuals of documents and labels were added. Additional modifications were incorporated as needed, such as removing ambiguities, re-organizing hierarchies, and adding missing concepts. This resulted in the AAENOTE which models and provides an index of annotated documents.
The terminology was developed from the annotation guideline by translating each hierarchy of entities into a class hierarchy, using attribute information to add data properties, and converting relationships into object properties. During development, the terminology was modified to remove ambiguities by adding new class hierarchies and modifying class names, object properties, and data properties. Specifically, to remove ambiguity between symptoms, infection signs, and device malfunction signs, the
Annotation Guideline vs Annotated Adverse Event NOte TErminology (AAENOTE). Annotation categories and entities are in
The results from all 4 annotation sessions were included as individuals in the terminology, but the terminology only reflects results based on the last annotation guideline. To accommodate revisions in the annotation guideline, annotation categories that were revised in the guideline are updated in the terminology correspondingly. For instance, removed annotation categories are reflected by changing the granularity of the removed category to a higher level. Annotation categories can also be re-organized to become subclasses of a different class. Moreover, newly added annotation categories are directly added as new classes in the terminology. An example is depicted in Fig. 4.

Handling annotation guideline revisions in the terminology. If the
Although the terminology was not developed to answer competency questions, the competency questions were still used to determine what could be found in annotated documents. To answer competency questions, the annotated documents were imported into the terminology as individuals.
Annotated Adverse Event NOte TErminology (AAENOTE)
Annotated AE documents and their annotations are modeled by the Annotated Adverse Event NOte TErminology (AAENOTE). To increase accessibility, the terminology is in both English and Norwegian. There are 149 classes, 5 object properties, 27 data properties, and 4470 individuals. The 7 classes which form the main hierarchies are:
Relationships between the 7 class hierarchies can be formed using the following 5 object properties:
An example showing AAENOTE representing an annotated document using parts of the class hierarchies and class properties is shown in Fig. 5. The complete class hierarchies of AAENOTE are in Appendix B.1.
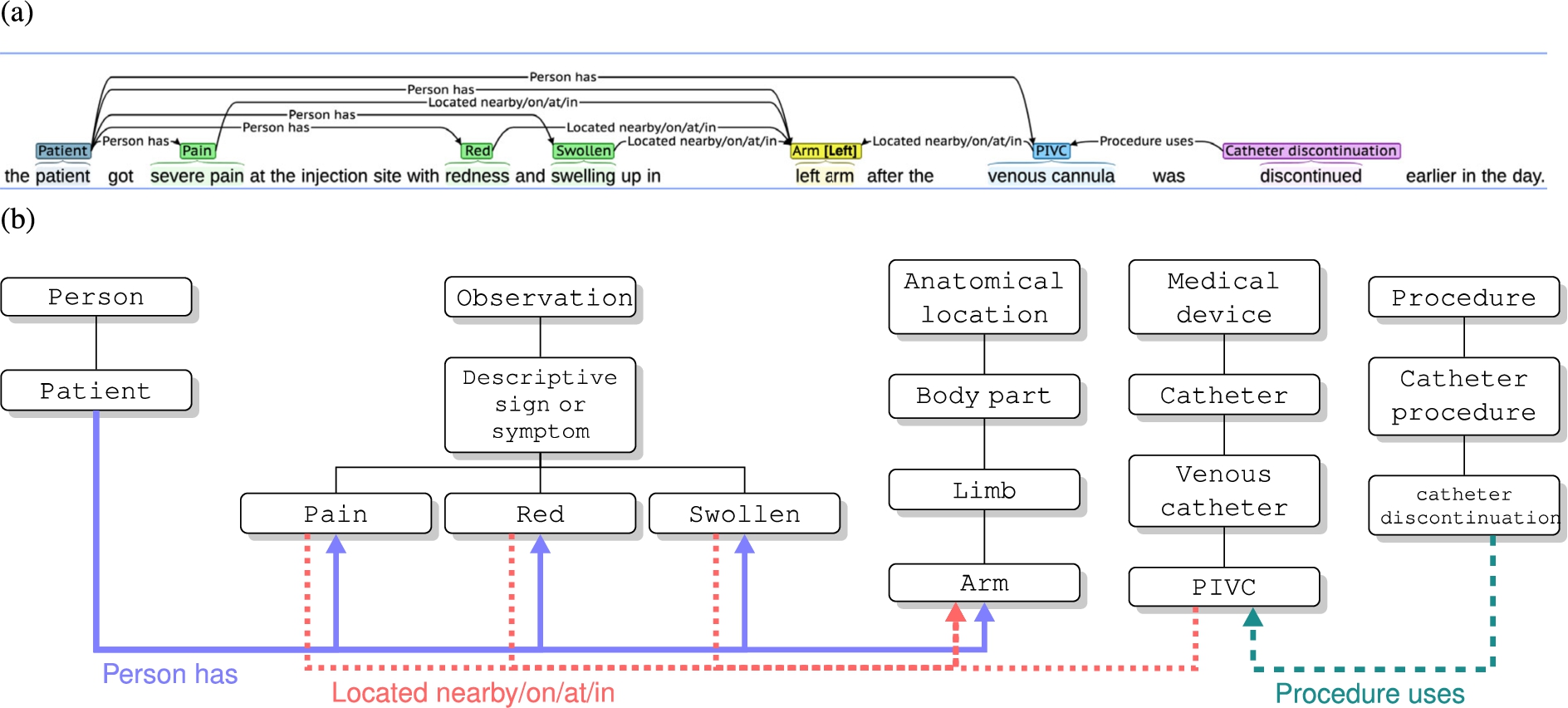
Annotated Adverse Event NOte TErminology (AAENOTE) representing an annotated document. (a) Example of an annotated document with annotation categories and relationships that link the categories together. (b) Part of the terminology class hierarchies used within the annotation example are shown in the white boxes. The 3 annotated relationships (i.e.,
Each annotated AE note or document is an individual of the
The purpose of AAENOTE is to model and provide semantic meaning to annotated AE notes or documents. The terminology was not developed based on competency questions, but it would be interesting to see what competency questions could be answered. Hence, AAENOTE was evaluated using the competency questions as requirements. Competency questions using AAENOTE can only be answered based on explicitly annotated classes or subclasses. Words or phrases that lack annotation are excluded from this terminology. Thus, only the annotation category labels provided by annotators are included as individuals of the corresponding classes.
Knowledge represented by the terminology can either be found explicitly, based on the direct classes and relationships, or be inferred implicitly, based on underlying concrete knowledge and indirect classes and relationships. For example, the competency question “Does patientA have an infection?” can be answered explicitly by finding an individual of the patient class who has an infection (i.e.,
SPARQL query results vary depending on the clinician’s interest in knowing how many instances a patient has for one class or a combination of that one class with other classes. For example, to find how many patients explicitly have an infection, the query can be written to find all instances where either: 1. individuals of the patient class have the object property “person has” to an individual of the infection class (i.e.,
AAENOTE SPARQL query result: patient has infection
Query result 1: Patient
Infection .
Query result 2–14: Patient
Infection and other Observation(s) .
AAENOTE SPARQL query result: patient has infection
Query result 1:
Query result 2–14:
AAENOTE SPARQL results for observations located at the skin
Query result 1:
Query result 2–10:
Overall, explicit queries and basic implicit queries in AAENOTE can answer 7 of the 9 competency questions. However, these queries still lack the clinical knowledge needed to include more implicit queries by combining additional observations, anatomical locations, and/or procedures to identify indications. In Appendix B, Table 7 provides the terminology classes and relationships used to form explicit and implicit queries to explicitly and implicitly answer each competency question. Additionally, concrete underlying knowledge used to make inferences is also provided. Results can be found in Appendix B.
Design decision for domain knowledge
In this use case, the clinicians’ need is to focus on identifying and inferring a patient’s state based on documented observations in AE documents related to PIVCs and BSIs. A patient’s underlying state can be measured by monitoring devices that measure vital signs (e.g., blood pressure, pulse, and respiratory rate) or exhibited by observable signs and symptoms (e.g., pain, fever, chills, and mobility impairment). Those measurable and observable signs and symptoms are then documented by clinicians in the electronic health record (EHR) to record patient conditions and communicate with other clinicians. When an AE incident could have or has happened, clinicians will go through the documentation to recall what occurred and report it in a separate AE document.
To limit the scope of modeling the clinical knowledge ontology, 15 documents were used as examples to form the classes and individuals. Each document was split into sentences to identify catheter and infection indications at the sentence-level and document-level. At the sentence-level, individual sentences were presented to clinicians who determined what observations, anatomical locations, or procedures within the text are needed to determine catheter and infection indication. Only clinician-identified sentences with indications were included as individuals in the ontology. At the document-level, individual sentences from a document were presented together, allowing clinicians to identify indications based on additional information from a more complete documented story. Presenting the document as separate sentences allowed clinicians to identify concepts within a limited example to determine what can and cannot be determined based on limited information. Whereas, allowing a clinician to see the whole document presented more possibilities and helped identify necessary data combinations for indications of catheters and infections.
The focus of the ontology includes catheter indications and the clinicians have identified that it is important to identify infusion phlebitis. Thus, infusion phlebitis was included in the ontology as rules based on the 1998 Visual Infusion Phlebitis Scale [30] mentioned in the 2021 Infusion Therapy Standards of Practice Updates [28]. Furthermore, causality is not within the scope because the exact reason for chemical and mechanical reactions resulting in infection-like signs are more likely found at the body’s cellular or genetic-level in pathophysiology studies [29,55] and unlikely to be documented in AE documents.
Anatomical locations in this ontology were kept simple and similar to the AAENOTE. Clinical guidelines for catheter insertion into anatomical locations are very specific (e.g., a central venous catheter is inserted in the jugular vein until it reaches the superior vena cava [31]) because clinical guidelines provide instructions on how to perform a task properly. However, clinical documentation is more general (e.g., central venous catheter in the chest) because this is common clinical knowledge, and the documentation is written for other clinicians to understand. To match the ontology with available documented data, this ontology relies on general anatomical location terminology. If clinicians deem it necessary, clinical guidelines can be included in a separate ontology focused on identifying catheter locations based on clinical guidelines and the FMA. Inclusion of clinical guidelines to identify specific catheter insertion sites and placement requires anatomical knowledge. For instance, to identify a central venous catheter’s general anatomical location using a clinical guideline and the FMA anatomy ontology, the ontology would need to:
Identify the jugular vein insertion site and the superior vena cava placement. Infer that the jugular vein is in the neck and the superior vena cava is present within the superior and middle mediastinum [65], annotated as anatomical location chest. Convert the terms into more general terms that match the available data (i.e., vena cava is in the chest).
Representing domain knowledge
Discussions with clinicians about example documents and indications formulated the classes, object properties, data properties, and rules within the ontology. Then, the provided indications were sorted and summarized to match the ontology closely. Afterward, indications were verified by clinicians and included in the ontology using SPARQL queries. A list of indications can be found in Appendix C.3.1 to Appendix C.3.7.
Ontology results for domain knowledge
Catheter Infection Indications Ontology (CIIO)
The Catheter Infection Indications Ontology (CIIO) represents clinical knowledge for signs of infections and catheters to identify PIVC-related BSIs and was developed to accompany the AAENOTE. Similar to AAENOTE, this ontology is also in both English and Norwegian. There are 57 classes, 10 object properties, 16 data properties, and 187 individuals. The 7 classes which form the main hierarchies are:
Relationships between the 7 class hierarchies can be formed using the following 10 object properties:
An example showing how a sentence is represented using the class hierarchies and class properties of CIIO to model documented clinical knowledge is shown in Fig. 6. The complete class hierarchies of CIIO are in Appendix C.1.

Catheter Infection Indications Ontology (CIIO) clinical knowledge representation. (a) A sentence from a document used to identify documented clinical knowledge. Annotations are based on terms from the Annotated Adverse Event NOte TErminology (AAENOTE). (b) CIIO has a
Each sentence documented in the report is an individual of the
Designed to capture and reason about clinical catheter-related and infection-related signs and symptoms documented in an AE report, the CIIO provides the missing clinical domain knowledge for the AAENOTE. CIIO can answer 8 of the 9 competency questions based on assumptions and indications. The assumptions are that 1 AE document represents 1 patient and all sentences within a document are likely to describe concepts within the same event (Appendix C.2). Indications for catheters and infections are provided in (Appendix C.3). Additionally, the ontology classes and relationships used to answer each competency question is detailed in Appendix C.4.
Discussion
Ontology development method comparison
The clinical problem drove this study, and the objective was not to apply an ontology development method. Hence, a specific ontology development method was not applied. However, certain steps taken are similar to the pre-existing methods and this study does include the typical phases of specification, conceptualization, formalization, implementation, maintenance, knowledge acquisition, evaluation, and documentation. An overview of the process is shown in Fig. 7, and similarities to other methods can be found in Appendix A.
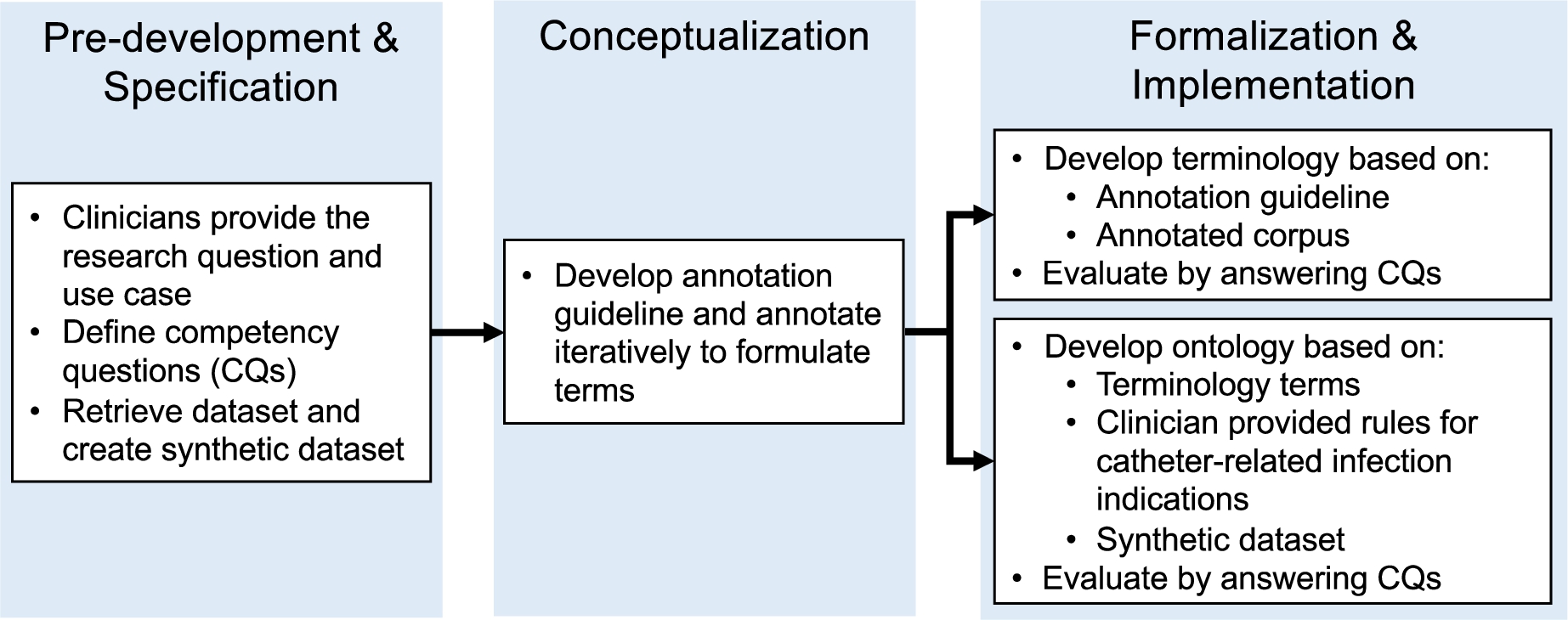
Development phases for Annotated Adverse Event NOte TErminology (AAENOTE) and Catheter Infection Indications Ontology (CIIO).
During the pre-development and specification phases of the terminology, clinicians provided the research question and use case. Those were utilized to define the competency questions. Additionally, the AE dataset was retrieved, and an AE synthetic dataset was created. The conceptualization phase was performed by iteratively developing the annotation guideline and annotation sessions. Afterward, the formalization and implementation phases of the terminology were developed iteratively based on the annotation guideline and using instances from the annotated corpus to answer competency questions for evaluation. Knowledge acquisition occurred during all phases with insight, guidance, and feedback from clinicians. Documentation is provided in the annotation guidelines, the annotated corpus, and the evaluation of competency questions. The annotation guidelines document changes in terms over time, and the annotated corpus documents knowledge acquisition from the text. Answers to each competency question are documented using natural language for clinicians and SPARQL queries for computer scientists.
Ontology development is similar to the terminology’s pre-development, specification, and conceptualization phases. However, the formalization and implementation phases differ. The ontology iteratively incorporated clinical knowledge that can be annotated in AE documents using terminology terms to answer competency questions for evaluation. Knowledge acquisition was provided through the annotated corpus, clinician-provided catheter indication rules, and clinician-provided publications containing phlebitis rules. Additionally, clinicians iteratively reviewed and verified documented sentences to match the rules and competency questions. Ontology documentation includes the assumptions (Appendix C.2), rules for catheter and infection indications (Appendix C.3), and how competency questions were answered (Appendix C.4).
Although UMLS includes clinical terminology (i.e., SNOMED CT and ICD-10), the terms are often a combination of different concepts. For example, phlebitis has many options and is combined with different locations, such as “phlebitis of the lower limb vein,” “phlebitis of the portal vein,” and “retainal phlebitis.” Additionally, swollen has many options, such as “foot swelling,” “swollen nose,” and “tongue swelling.” The June 10, 2022 version of SNOMED CT has 361,907 classes.2
The clinician asks questions about the condition of a physical patient and the patient’s PIVCs, but AAENOTE is about words in the AE document regarding a patient event. The correspondence between clinical condition and document content is represented by CIIO. So, the answer to a question about a patient’s condition will be answered using the document’s annotated text. Understanding a query means translating it from clinical concepts to concepts within the document’s content. Here, the terminology is used to fit and answer questions. SPARQL queries can answer most competency questions, and the results can be used as consistency checks. For example, SPARQL can be used to count and make quantitative queries about the number of catheters and devices. Likewise, qualitative results enabled clinicians to verify if results matched their expectations of clinical events (i.e., the anatomical location of specific catheters) or why the AE was reported (i.e., incorrect medical devices used in a particular procedure).
There are several limitations to the AAENOTE. Although this terminology does not cover sepsis, it does cover events that could lead to sepsis. This terminology lacks the clinical knowledge required to answer several competency questions more in-depth. Moreover, it is not always possible to determine what a patient has because of the document’s content or provided annotations. Most documents do not explicitly mention a patient because these are AE documents, and it is often implied that the adverse event has happened to a patient. Annotators will often not link the patient to all possible observations, anatomical locations, medical devices, or procedures because typically, one AE document refers to one event or patient. Furthermore, referent tracking and resolution are not handled by AAENOTE. Thus, multiple mentions of a label or individual do not indicate whether it is the same item or a different item. For example, given the annotated document in Fig. 2, the terminology cannot determine if the 2
CIIO scope and limitations
The CIIO is an abstract ontology with instances populated using the terminology. Only parts of the AAENOTE necessary for creating queries with clinician-provided indications were included or extended. This provides flexibility, allows for easier ontology maintenance, and separates the needs of clinicians who use CIIO and annotators who use AAENOTE.
Based on assumptions and indications, competency questions can be answered using SPARQL queries. The queries retrieve documents and translate the content for the user by identifying concepts necessary to answer the competency questions. Thus, the retrieved documents and concepts can provide sufficient information for clinicians to further decipher retrieved answers. For example, the exact reason why a patient needs a catheter cannot be determined by the query unless there is a direct relationship (i.e.,
The lack of detailed documentation inhibits the query’s ability to answer certain questions. This includes why a patient needs a catheter as previously stated, counting catheters within a patient, and where the catheters are located in a patient. Counting the exact number of catheters per document is not possible because multiple sentences within a document could be describing the same catheter or multiple procedures could use the same catheter. The exact anatomical location of catheters per document cannot be determined for several reasons. First, multiple sentences within a document could be describing the same catheter at the same location but with more general terms (i.e., arm instead of hand). Second, the location’s position not being documented makes it difficult to distinguish if a body part is on the same side. Finally, various procedures can be performed at the same location. For example, given an example document, “The patient received IV fluids in elbowA and IV antibiotics in right handB. Right armC showed signs of phlebitis.” Here, handB is part of armC because both are on the right side, but elbowA might or might not be part of armC. Additionally, armC is likely a more general term for handB. Furthermore, an additional anatomical ontology is needed to infer the possible locations based on catheter type.
Purpose of separate terminology and ontology
Even though the ontology uses terms from the terminology, the terminology and ontology are separate. They are separate because of their different purposes and functionalities. Additionally, separation provides downstream analysis flexibility for researchers. It also simplifies evaluation and allows for easier maintenance. Furthermore, separation enables a better understanding of the terminology’s and ontology’s limitations.
The terminology and ontology were developed for different purposes using different methods. AAENOTE is intended to be useful for annotators who are annotating documentation and a way to provide them a structured terminology with varying granularity. In comparison, CIIO is intended to be used by clinicians to clinically reason about a patient state. The design process of AAENOTE is heavily based on the explicit terms used in annotations and not on competency questions. It uses the bottom-up method and the annotation guideline development process to capture semantic annotations. In contrast, the design process of CIIO is based on the competency questions, which focus on patient states. Designed with a top-down method, it is based on concepts naturally used by clinicians to describe patients. Thus, the terminology and ontology have different purposes and functionalities.
Separating the terminology from the ontology enables annotators to annotate concepts with standard terms and clinicians to reason about the annotated concepts. Here, the ontology does not impact the terminology annotators can use. Instead, the ontology provides knowledge for the terminology. Thus, our methods avoid the significant amount of time, effort, and manual curation previously required to map terms to an ontology [61]. Instead, our ontology utilizes concepts in the terminology and is limited by the competency questions, clinical guidelines used, and clinician-provided rules. In downstream analyses, researchers can freely choose to use the terminology to quickly retrieve documents with specific annotations, the ontology to reason and infer clinical knowledge, or both.
The terminology indicates concepts annotated in documents. Using terms from the terminology ensures that the included clinical knowledge within the ontology represents the knowledge documented in the text that can be annotated. As the ontology develops further, it is possible to conceptualize additional terms required to answer the competency questions. Those terms can then be added to the terminology and annotation guideline for additional data curation.
In this paper, separating the terminology made it easier and quicker to evaluate syntax and semantics because the ontology only has 187 instances compared to the terminology’s 4470 instances. Mixing the indexed annotation terminology with a clinical knowledge ontology would be outside the ontology’s scope, decrease ontology reusability, and increase the complexity of ontology maintenance. Additionally, the terminology can cover a broader scope of documents not in the ontology. Finally, using competency questions to evaluate the terminology and ontology separately reveals the distinct limitations of both. The inability to answer competency questions can be due to either the lack of knowledge or lack of necessary content within the data.
Representing annotated data and revisions
Annotating data provides data meaning, and the corresponding annotation guideline and terminology provide additional structured semantic meaning. Additionally, the terminology can represent knowledge and disambiguate annotation entities and relationships. Each annotation session uses a slightly different annotation guideline that has been revised based on the previous annotation session. Hence, revisions in the annotation guidelines include added, re-organized, and removed categories. Since results from 4 different annotation sessions are included as individuals in the AAENOTE, this indicates the terminology can handle different versions of annotated data while preserving semantic meaning. It is also possible to easily customize the granularity (i.e., superclasses, classes, or subclasses) and extend or retract the terminology based on clinician needs without breaking the terminology.
To alleviate the problem with overlapping terms and ambiguities experienced by [5] and remove mismatches between the annotations and the terminology experienced by [61], all annotators in our study could only use provided annotation labels from the terminology. Using concrete concepts from the annotation guideline based on what can be found in the documentation instead of other pre-existing ontologies lowers the complexity and simplifies the terminology.
Representing annotated documents the way clinicians view patients
The AAENOTE is a terminology that provides an index of what is annotated in a clinical document. It is not used to design a language’s syntax, grammar, or terms because AAENOTE is a terminology for understanding the language and underlying meanings. Instead, it is the interpreted formalized language that has been translated into basic statements for reasoning. To capture relevant information, the underlying document was represented by annotated labels and relationships for the task of question answering and text understanding instead of solely retrieving information. Hence, the terminology focuses only on items of interest and is blind to items not within the terminology.
The corresponding CIIO is an ontology that models clinical knowledge missing from AAENOTE. It provides the missing clinical knowledge required to reason about the presence of catheters and infections documented in clinical text. Although the data modeled is documented text, it enables clinicians to think about the data as an individual patient because they already do this routinely when documenting patient states.
Understandability and accessibility for domain experts
The approach in this study made ontology development understandable and accessible for the domain experts without formal ontology training. Furthermore, the employed approach made it possible for clinicians to understand and be part of the design process. In practice, the approach was a necessity to progress in developing the ontology to incorporate clinical knowledge.
Clinical utility
The collected competency questions and requirements are largely met. Thus, the main objective of developing a terminology and ontology that clinicians and hospital systems can use to get a systematic overview of identifying and reasoning about PIVC-related phlebitis, infection, and sepsis in an AE corpus has been met. Furthermore, our ontology is a step toward automated and continuous quality control that move beyond today’s focus on repeated point prevalence quality controls, like the Peripheral Intravenous Catheter mini Questionnaire (
The developed ontology is of value for sepsis because of its purpose, clinician involvement during development, and intended use. The ontology focuses on identifying indicators of catheter-related phlebitis or infections that can lead to sepsis by utilizing the clinicians’ documentation and perspectives. Throughout the whole development, clinicians were involved as the users, domain experts, and data annotators. Furthermore, clinical knowledge within the ontology was captured similarly to how clinicians ask questions, document observations, and view documents as patients. The intent is to eventually implement the ontology into a quality surveillance system to automatically detect the presence of PIVC-related phlebitis and BSIs to improve PIVC care and lower sepsis incidents. Thus, only documented content can be included as data, and the ontology must directly correspond to and represent concepts documented within the AE documents from the clinician’s perspective.
Future work
For the sepsis-related use case, the synthetic AE dataset used for annotations is a placeholder for the real Norwegian AE dataset and clinical records from the EHR. Future work includes utilizing the current AAENOTE to annotate the real Norwegian AE dataset and clinical records. And to evaluate if clinical knowledge from the CIIO can still be applied and expanded on new data. Additionally, the ontologies could be applied to AE documents at other Norwegian hospitals to assess how similar documentation and knowledge are between different hospitals. The ontologies can be directly translated to other Scandinavian languages (e.g., Swedish and Danish) and applied similarly at other Scandinavian hospitals. The design and representations are largely language-independent and should be easy to transform for English clinical text about adverse events. After all, international literature suggests that the phenomena related to PIVC and devices are language-independent [1]. It would also be possible to provide multi-language querying over multi-language AE documents to enable cross-language repositories [63]. Furthermore, supervised machine learning methods can be employed to identify PIVC-related BSIs and classify patients requiring additional monitoring.
Conclusion
The development process resulted in a terminology and an ontology, specifically, the Annotated Adverse Event NOte TErminology (AAENOTE) which models annotated classes in annotated documents and the Catheter Infection Indications Ontology (CIIO) which models clinical knowledge for catheter and infection indications. Although there is a clinical focus here, the methodology for creating a terminology from an annotation guideline for semantically annotated data and a domain knowledge ontology to represent knowledge can be utilized in other domains to provide additional semantic meaning to annotated datasets in other domains.
Data availability
The AAENOTE, CIIO, and SPARQL queries for this paper are in the GitHub repository branch “swj” of
Footnotes
Acknowledgements
The authors would like to acknowledge the eight annotators for producing the annotated clinical adverse event (AE) corpus. Funding was provided by the Computational Sepsis Mining and Modelling project through the Norwegian University of Science and Technology (NTNU) Health Strategic Area. Additionally, ethical approval was granted by the Regional Committees for Medical and Health Research Ethics (REK), approval no. 26814.
Ontology development methods and evaluation similarities
There are various ontology development methods and the steps in this study have some similarities to other methods Table 6.
