Abstract
The research goal of this work is to investigate modeling patterns that recur in ontologies. Such patterns may originate from certain design solutions, and they may possibly indicate emerging ontology design patterns. We describe our tree-mining method for identifying the emerging design patterns. The method works in two steps: (1) we transform the ontology axioms in a tree shape in order to find axiom patterns; and then, (2) we use association analysis to mine co-occuring axiom patterns in order to extract emerging design patterns. We conduct an experimental study on a set of 331 ontologies from the BioPortal repository. We show that recurring axiom patterns appear across all individual ontologies, as well as across the whole set. In individual ontologies, we find frequent and non-trivial patterns with and without variables. Some of the former patterns have more than 300,000 occurrences. The longest pattern without a variable discovered from the whole ontology set has size 12, and it appears in 14 ontologies. To the best of our knowledge, this is the first method for automatic discovery of emerging design patterns in ontologies. Finally, we demonstrate that we are able to automatically detect patterns, for which we have manually confirmed that they are fragments of ontology design patterns described in the literature. Since our method is not specific to particular ontologies, we conclude that we should be able to discover new, emerging design patterns for arbitrary ontology sets.
Keywords
Design patterns in ontology engineering
Today’s methodological guidelines in ontology engineering (see, for instance, the NeON methodology [33]) suggest to
ODPs have been proposed as a method analogous to design patterns in software engineering [11,12,28] that aim to provide good quality solutions to recurring modeling problems. Blomqvist et al. [4] have proposed various types of ODPs, e.g., content, structural or lexico-syntactic ones. Ontology patterns may also be specific for a certain domain, for example, Aranguren et al. [3,10] developed ontology patterns specific for biology. Many of the proposed patterns can be found in two repositories, the ODP Portal,1

Examples for the three types of patterns covered in this paper: 1. Axiom Pattern with Variables (APV), 2. Axiom Pattern without Variables (APNV), and 3. Class Frame Pattern (CFP).
In the current practice, ontology authors create the ontology patterns manually, and sometimes, they upload them to one of the ontology patterns repositories. However, developing such patterns is very laborious. Moreover, ODP repositories are not yet comprehensive – not all recommended design solutions are recorded in these repositories of patterns. Even with the availability of such repositories, domain experts still have difficulties to find and apply a suitable modeling pattern, when having to choose among several, possibly abstract patterns.
In many cases, recurring patterns of axioms may exist in ontologies, even if they have not been officially published as a part of a recommended design pattern. We call such empirical patterns
For the purpose of this work, we will identify three different types of patterns: Axiom Patterns with Variables (APV), Axiom Patterns without Variables (APNV), and Class Frame Patterns (CFP). Figure 1 shows examples for the three types of patterns with the goal of introducing the terminology. We provide the formal definitions of the three pattern types in Section 3.2.
Our research objective in this work is to investigate patterns that occur frequently in individual ontologies, as well as in a group of ontologies, such as the ones stored in an ontology repository. Our work is guided by the following research questions:
Do certain patterns recur in ontologies? Can we generalize over such patterns to mine more generic templates?
Do such patterns appear within a group of ontologies?
Do such patterns exist on the axiom level? Do they exist on the level of sets of axioms?
Are we able to automatically detect fragments of documented ODPs?
The main contributions of our work are:
We propose a method based on tree mining for discovering frequent
We propose an association–analysis method to discover frequent
We conduct an experimental analysis on BioPortal ontologies with the goal to discover frequent ontology patterns.
Our analysis reveals that: (i) We are able to identify recurring patterns in ontologies, both at the axiom level, and at the level of sets of axioms; (ii) We are able to automatically extract non-trivial, and significantly-frequent patterns without variables; (iii) We are able to discover patterns with variables; and (iv) We are able to automatically mine fragments of already known ODPs. All results obtained during the experiments are available online at:
The rest of this paper is structured as follows. Section 2 summarizes the related work. Section 3 introduces the problem of tree mining, and formally defines the notions of ontology axiom and class frame patterns. Section 4 describes the BioPortal dataset used in this study, and the proposed methods. Section 5 describes the results of our experiments. We discuss our results in Section 6, and conclude in Section 7.
Prior research on the topic of extracting ontology patterns has been quite scarce. Some previous works dealt with studying the syntactic properties of OWL ontologies on the Web. In their work, Wang et al. [39] presented statistics on the occurrence frequency of OWL language constructs, and the structure of ontology class hierarchies, in a corpus of ontologies. Zamazal et al. [34] conducted a study on collections of OWL ontologies with the aim of determining the frequency of several combined name and graph patterns, which potentially indicate underlying structural clusters. These works mainly deal with lexical patterns, and do not tackle mining recurring fragments in ontologies.
Mortensen et al. [23] studied the use of ODPs in BioPortal ontologies. The authors encoded 68 ODPs from two online pattern libraries (Manchester ODP Catalog, and the ODP Portal) using the Ontology Pre-Processor Language (OPPL).3
In our previous work [19], we presented a method, Fr-ONT, for mining patterns in ontologies. However, our data-driven method worked only on ontologies with instances. The method iteratively constructed new ontology classes, and checked their frequency in terms of the number of instances rather than mining frequent fragments in existing axioms, as we do in our current work.
In their work, Khan and Blomqvist [17] detected content ODPs in existing ontologies. Their method works top-down, starting from existing ODPs and trying to find their instantiations in an ontology. In our method, we do not require an ODP as an input, and we mine (possibly new) patterns bottom-up. Similarly, Šváb-Zamazal et al. [35] considered the problem of detecting (logical) patterns top-down, starting from a particular pattern. Thörn et al. [37] studied potentials and limits of graph-algorithms for discovering ontology patterns based on a definition of what structures are considered patterns. Their conclusion was that graph-pattern algorithms appear inefficient for finding patterns in ontologies. Tempich and Volz [36] performed a statistical analysis of the DAML ontology library. They mostly studied the language primitives with a goal to establish a benchmark for Semantic Web reasoners.
The approach taken by Mikroyannidi et al. [21,22] with the Regularities Inspector for Ontologies (RIO) is the closest to our approach. The authors used clustering to identify regularities in the usage of entities in axioms within an ontology. The authors defined the distance based on the similarity of the structure of ontology axioms [21]. Thus, the process of clustering groups axioms (more precisely, axiom templates) based on their similar structure. Our work differs from this approach in two important aspects. First, the method that we use is different, and is based on frequency- and association analyses. Second, we are able to discover sets of axiom templates (class–frame fragments), rather than only single-axiom templates, due to the use of association analysis, which can group axiom templates of very different structures. We discuss our approach in comparison to RIO in more detail in Section 6.5.
Tree mining
Tree mining is an area of data mining that deals with the discovery of frequent subtrees in tree-shaped data structures. Tree mining has been applied to several areas, such as, bioinformatics, web usage mining, and mining XML files [42]. We use the
In the following paragraphs, we will introduce some basic definitions and terminology.
A
A
A
A
A
An
An
A
In this work, we assume that all trees are rooted and labeled, and that all forests are labeled.

By the
The
Our research is concerned with identifying patterns in ontologies represented in the
An
Entities are the basic building blocks of OWL ontologies, defining the vocabulary of an ontology. An OWL vocabulary
OWL provides several constructors to combine entities into more complex class expressions. The complex class expressions are defined inductively using the following grammar:4 We use the Manchester syntax [15] for OWL ontologies throughout the paper.
For the analyses described in this paper, we consider two classes of logical axioms, namely subclass axioms
Following the terminology of Horridge et al. [16], we define the

In other words, a class frame
Simple ontology on anatomy serving for the illustration purposes
As part of this work, we also identify patterns containing variables. We define the following sets of symbols (variable names) that are not in the vocabulary of
An
A sample axiom pattern with variables (APV) corresponding to our illustrative example from Table 1 is:

An
An example of an axiom pattern without variables is the following (we replaced the entity ids with their labels to improve readability. See details in Appendix Table 10):

A
A sample class frame pattern (CFP) with a variable left-hand side is:

An
An
The examples for the different types of patterns are also shown in Fig. 1.
BioPortal ontologies
For the experimental evaluation, we downloaded on July 25, 2015 a snapshot of all ontologies from the BioPortal ontology repository5
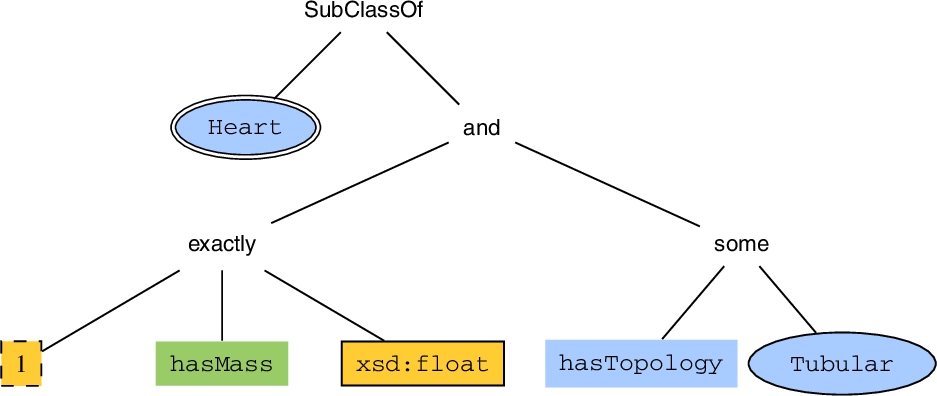
Tree representation of an axiom:
Our aim is to find frequent patterns based on OWL subclass and equivalent class axioms, such that their left-hand side is a named class. We convert every axiom to a single tree, which then is used as an input for the SLEUTH algorithm (Section 4.3). We build the tree by recursively processing the arguments of each constructor, starting with
OWL constructor concerning classes or datatypes (e.g., intersection), corresponding to
named class, named datatype, corresponding to
named object or datatype properties, corresponding to
named individual, literal value (e.g., in
cardinality values in cardinality restrictions, corresponding to
facet datatype restriction (e.g., in limiting range of integers);
left-hand side of subclass axioms (always a named class);
As an example, let us consider the following axiom:

The tree corresponding to the axiom is presented in Fig. 3.
Because SLEUTH operates only on numeric labels, every distinct pair (type and name) is assigned an unique integer value. These values are stored to decode frequent trees back.
By applying this method to the BioPortal ontologies, we obtained a set of 331 non-empty forests, the smallest of which containing 4 trees, while the largest containing 1,833,925 trees. The histogram of the forests’ sizes is presented in Fig. 4. The median size of a forest was 629, while the average size was 25,308.4, with the standard deviation of 147,725.3.

Forests’ sizes histogram. The forests were obtained from 331 BioPortal ontologies, each converted to a forest of trees, where each tree corresponds to a
In order to extend and apply SLEUTH [42] – an efficient algorithm for mining frequent, unordered, embedded subtrees – to our use case, we needed to encode each tree into an efficient string representation, as described in this work [43]. Precisely, a tree
The
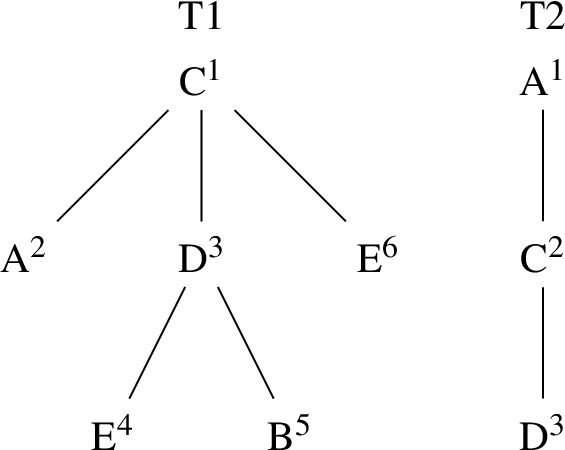
A forest of two trees, T1 rooted in C and T2 rooted in A. Numbers in the superscript represent an order of depth–first preorder traversal of the trees.
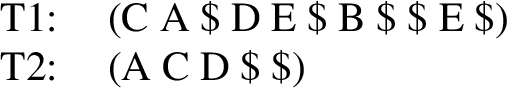
Depth–first preorder string encoding of the trees T1 and T2. $ is the special symbol to indicate backtracking.

Scope–list representation of the trees T1 and T2.

Scope–list representations for some more complex subtrees. Values in parentheses are
Despite its feasibility, the SLEUTH algorithm has two disadvantages, which renders it unsuitable for our use case. The first disadvantage is concerned with the embedded patterns which do not provide valuable information about axioms in an ontology. For example, consider these two axioms, corresponding to the axioms #1 and #4 in Table 1:

The trees corresponding to these axioms are presented in Fig. 6. One of embedded patterns occurring in the axioms is

Trees for axioms [
The second disadvantage of the SLEUTH algorithm is that it mines tree patterns in a single forest, which is not sufficient. Indeed, a single ontology forms already a forest, and we are also interested in mining patterns that occur in multiple ontologies. Thus, we need a method to mine tree patterns in a family of forests.
To address these two disadvantages, we modified the SLEUTH algorithm. To deal with the first issue (unintended patterns), we keep and extend only the induced subtrees. Other subtrees – those that are embedded, but are not induced – are discarded. This change does not affect the soundness of the algorithm, as all induced subtrees of a frequent, induced subtree must also be frequent, and are therefore constructed as well.
We addressed the second issue (mining in a family of forests) by introducing a new measure of support. Considering a family of forests

A family of two forests
Figure 7 shows an example explaining how this new support is computed. The support is also applied to the axiom pattern corresponding to the tree after decoding it. To better exemplify how the support is computed, consider the two ontologies shown in Table 1 and the axiom pattern with variables (APV):

Recall the example from Table 1. This axiom patterns has support 1 in Ontology 1 (matches axiom #2), and support 2 in Ontology 2 (matches the axioms #8 and #10). The support of the axiom in the family of trees (composed of Ontology 1 and 2) is 2, as it matches at least one axiom from each of the ontologies.
As the original SLEUTH implementation9 Available at:
Both SLEUTH and FF-SLEUTH compute the set of all frequent induced subtrees. Obviously, not all of the computed subtrees are useful for our purposes mainly due to two reasons.
First, we are interested only in maximal frequent subtrees – frequent subtrees for which none of their proper supertrees is frequent. Consider the forest in Fig. 8: there are many frequent subtrees of these two trees, such as, (

There is one maximal frequent subtree of these two trees, with nodes denoted by bold symbols. All subtrees of this maximal tree are also frequent, but are not useful for our analysis.

During mining we also discover frequent subtrees that do not contain
The second reason is that we aim to mine axiom patterns with or without variables. Therefore, we will consider only the frequent subtrees which contain
For the further processing, we will use only the frequent subtrees which fulfill both of the above-mentioned criteria – being frequent and having the proper form. The next step in our analysis is to transform the mined trees back into
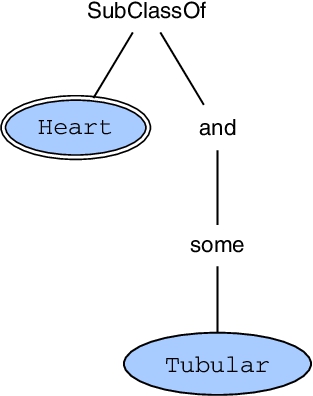
A frequent subtree with decoded labels that is an incomplete tree representation of some axiom and that (after completion) will form a frequent axiom pattern
After the completion, we obtain a frequent axiom pattern:
The complete algorithm for decoding and completion is published as a supplementary material.10
So far, we have described the methods for computing single axiom patterns. In this section, we describe how to compute class frame patterns (CFP) based on the discovered axiom patterns. In the simplest case, there might be axiom patterns that have a named class on their left-hand side (
In the classical formulation of the frequent itemset mining task, the inputs are: (1) a set of
The process for mining frequent class frame patterns has three phases and is illustrated in Fig. 11.

A three-phase process: mining frequent class frame patterns on top of the discovered frequent axiom patterns with use of propositionalisation.
In the first phase, we mine frequent axiom patterns over an ontology
In the second phase, we apply
A propositional (attribute-value) representation where attributes (features) are constituted by frequent axiom patterns and examples are constituted by ontology classes
In the third phase, we apply an off-the-shelf frequent itemset mining algorithm. The result of the algorithm will be a set of itemsets, which correspond to class frame patterns (
We define the
The
In the example from Table 2, the class frame pattern
Frequent axiom patterns in single ontologies
To discover frequent axiom patterns in single ontologies, we encoded each ontology into a forest using the method described in Section 4.2. We then used the original SLEUTH implementation on these forests to mine frequent induced subtrees with relative support treshold of
In order to present clearly the results, we divided our ontologies into six groups, depending on the number of

Various statistics for frequent axioms patterns computed for each single ontology from our BioPortal dataset (cf. Section 4.1). The ontologies are clustered into 6 groups depending on their sizes.
The number, support, size and depth of mined axiom patterns are presented in Figs 12(a)–12(d). Each figure contains a box plot for every ontology group showing: the median
In Fig. 12(a), we present the number of mined frequent axiom patterns for each ontology group from our dataset. In Fig. 12(b), we present the supports for the mined frequent axiom patterns. As we used relative support threshold of
Figure 12(c) shows the sizes of the frequent axiom patterns that we mined. Interestingly, in larger ontologies, the median size increases, and IQR decreases. Finally, Fig. 12(d) shows the depths of the frequent subtrees. Here, we can also observe that the median value is higher for larger ontologies.
We discovered that 96% of the ontologies (320 out of 331) in the dataset reuse patterns containing vocabulary from domain namespaces (namespaces other than
In the 321 ontologies, we found patterns of size at least 2; in 81 (24%) ontologies, we found patterns of size at least 5; in 17 (5%) ontologies, we found patterns of size at least 10; in 4 ontologies, patterns of size at least 20; and in one ontology (NEMO), we found 2 patterns of size 43. The biggest axiom patterns from some of the most used ontologies available in BioPortal are presented in the Appendix in Table 9.11 We display labels instead of IRIs, where possible.
The patterns with the highest support from a subset of most visited ontologies in BioPortal

One of the two the longest patterns discovered in a single ontology. Both longest patterns were discovered in the NEMO ontology, and have a size of 43. The presented pattern has support 27 and depth 13. The other longest pattern is nearly identical to the presented one.
Frequency statistics for namespaces corresponding to upper-level and cross-domain ontologies. The second column is the number of times a given namespace occurred in all patterns (multiple occurrences in a single pattern are possible). The third column shows the number of patterns containing the namespace. The fourth column shows the number of ontologies that contain at least one of these patterns
One of the patterns of size 43 in the NEMO ontology (Fig. 13) has resulted from repeating a substantial fragment of the class definition in the subclasses of the class
We discovered that 99.2% of all mined patterns contain at least one variable. Out of these, 26.1% contain a variable in the left-hand side and 89.6% contain a variable in the right-hand side.
We have also investigated the frequency statistics for namespaces occurring in axiom patterns. We looked mainly at top-level and cross-domain ontologies. We found that most patterns appear in OBO12 OBO stands for Open Biomedical Ontologies. The library of OBO ontologies can be found at:
To mine frequent patterns in the set of ontologies from the BioPortal repository, we used FF-SLEUTH with the support measure based on a set of forests (Section 4.4). Precisely, every axiom was translated to a tree (Section 4.2), and every ontology formed a single forest. In this way we were able to discover frequent patterns that occur in multiple ontologies independent of the relative sizes of the ontologies. If we were to combine all axioms to form a single, huge forest, then patterns from large ontologies (such as, NCIT) would dominate the results.
An experiment with a minimal support of 4 forests (i.e.,

Distributions of various statistics for patterns mined with minimal support 4. Top and right charts present histograms for one dimension each, while the charts in the middle present 2D-histograms for both statistics combined using varying color intensity.
Top ten axiom patterns found in the set of BioPortal ontologies, sorted by descending support and size
Table 5 shows the top-identified patterns with the biggest support. The top pattern with support 29 turned out to be an artifact of the way the OWL API converts OBO to OWL. We found one pattern without variables and three patterns with variables that appear in 27 ontologies.
The biggest subtree, which occurs in 14 ontologies, represents a pattern without variables (or fragment), and it is shown in Table 6. This pattern represents the logical definition of the class
An axiom pattern without variables (or fragment) corresponding to the biggest subtree discovered in a set of ontologies. It has size 12, depth 3 and support 14
We have also examined axiom patterns that have the largest size. The sorted list is available in Table 10 in the Appendix. The size of the patterns is presented in the first column of the table and their total number of occurrences (the number of ontologies where they occur) is shown in the second column. We can observe that the largest patterns come from OBO ontologies and that they include entities from upper-level ontologies (BFO, OBI, IAO, snap, span).
Similar to our investigation for single ontologies, we have also examined which namespaces occur more frequently in ontology patterns in the entire set with a minimal support of 4. As in the previous cases, the most frequently-occurring namespaces are the ones from OBO. The full list is shown in Table 7.
Namespaces occurring in frequent patterns mined with minimal support 4
We used the discovered axiom patterns to analyse class frame patterns (CFPs) as described in Section 3.2. To conduct the analysis, we used Orange3-Associate13
We have computed transactions for all ontologies and all their classes matching at least one frequent axiom pattern. Altogether, we have discovered 5,397 frequent CFPs. 2,335 (43%) of these patterns are composed of more than one axiom pattern. On average, there are 16.3 CFPs per ontology (with median value 11.0), containing an average number of axiom patterns equal to 2.7 (with median value 1.0), and with average support 233.8 (with median value 6.0). The biggest CFP (in terms of the number of axiom patterns) that we have discovered is composed of 17 axiom patterns, and the most frequent CFP that we have discovered has the support of 178,320.
Selected class frame patterns (CFP). First column displays the name of the ontology where the CFP was found. Second column contains the relative support
In the following paragraphs, we discuss some sample CFPs from specific ontologies. The CFPs are documented in Table 8.
The

(a) ‘Cell Line Cells’ design pattern for the CLO ontology [30] (top). (b) The selected corresponding class–frame fragments, which we have automatically mined (middle and bottom part of the figure).
The
The
The
The
The
The
The
One of the research questions that we are trying to answer is whether our methods are able to mine axiom patterns of ODPs described in literature. Figure 15 shows the CFPs that we have automatically mined for the Cell Line Ontology (CLO). After a manual investigation, we have subsequently established that the mined patterns reflect the ‘Cell Line Cells’ design pattern proposed by Sarntivijai et al. [30]. CLO is one of the largest BioPortal ontologies from our dataset (it contains 114,843
One surprising finding is the fact that we discovered several CFPs, which contain a part that is not included in the original ODP, namely: (‘has quality at some time’
The Manchester ODP Catalog and Ontorat [41] are two pattern repositories for biomedical ontologies that document or refer to patterns from ontologies contained in our dataset: CLO, OBI, OOEVV, BCGO, and CCO. We investigated whether we were able to mine the documented patterns. Our manual inspection confirmed that we were able to mine the patterns in CLO (described above), as well as the ’Assay’ pattern from OBI and an instantiation of a fragment of the ’Device’ pattern from the same ontology, and a pattern concerning the main classes of OOEVV [7].
Discussion
Research questions
We found patterns in every ontology in the experiment, with the exception of those that did not have any
We also noted that most patterns contain vocabulary from OBO ontologies (Table 4). This finding hints at the fact that modeling patterns and reuse are more prevalent in OBO ontologies than in the other ontologies in the dataset. This fact is not surprising as OBO ontologies follow the principles set forth by the OBO Foundry [31], which prescribe a strict set of rules for reuse and orthogonality of ontologies.
We have also observed that the median fragment size for smaller and larger ontologies (with less or more than 1,000 axioms, respectively) is fairly similar, between 2 and 3 (see Fig. 12(c)), although there are variations in IQR. This finding may indicate that most patterns are still fairly simple, rather than complex expressions, and are usually of the size of two or three. We discovered that the majority (99.2%) of all mined axiom patterns contain at least one variable, out of which 89.6% contain a variable in the right-hand side of the axiom.
We found that ontology patterns exist, not only in single ontologies, but across the set of investigated ontologies. In the latter case, the longest patterns discovered from the set of all ontologies (Table 10), are patterns without variables. They represent fragments from OBO ontologies, which have likely been copied from other ontologies. For example, the ‘curation status specification’ class (Table 6) is originally defined in the file
We also noted that several of the rows in Table 10 are fragments (i.e., APNV) of upper ontologies – such as BFO – or cross-domain ontologies – such as, OBI or IAO. One question that arises is whether these fragments may represent reusable ontology modules [32], which would be valuable also outside of the OBO community. To facilitate their reuse, such modules could be made available separately from the ontologies from which they originate.
We found patterns on both levels. We were able to mine frequent patterns from every ontology that contained
This result is intriguing taking into account that Mortensen et al. [23] found modest reuse of ODPs in BioPortal ontologies. These results are, however, not contradictory. The approach taken by Mortensen et al. is top-down – they test the occurrence of a set of several predefined patterns in the ontology dataset. This study found that the ontologies in BioPortal contain some of the structural patterns from Manchester ODP Catalog, and a few high-level content patterns from the ODP Portal.
In contrast, our approach mines patterns bottom–up, and can also detect parts of specific content ODPs. We call our mined patterns “emerging”, as they may not comply to predefined ODPs in existing repositories. Yet, these patterns appear in the studied ontologies, likely because they are valuable to the ontology authors and users.
We were able to establish manually that some of the patterns that we have automatically mined are fragments of ODPs proposed in literature. We have detected fragments of ODPs for CLO, OBI (the ‘Assay’ pattern), and OOEVV, which were documented in the Manchester ODP Catalog and in the Ontorat repository. For the ‘Device’ pattern for OBI from Ontorat, we have been able to detect an instantiation of a fragment of the pattern (

A sample logical ODP found in

A sample alignment ODP found in
Besides the exact parts of the proposed ODPs, we also found two other types of constructs:
We note that BioPortal hosts a relatively well-described set of ontologies. The ontologies are documented either in scientific publications, and/or on the webpages of the projects that developed them. Thus, it allowed us to identify the patterns used in the ontologies’ construction, and then to check whether our algorithm can mine the documented patterns. We also note that our approach is generic and it can be applied in other domains and with other datasets.
The authors of [4,12] distinguish six types of ontology design patterns: content, structural, correspondence, reasoning, presentation, and lexico-syntactic. Our method is suited to mine three of the six types of patterns: content, some structural (logical) and some correspondence (alignment) ODPs. However, our approach is not suited for discovering reasoning, presentation and lexico-syntactic ODPs.
The content ODPs are the main target of our method. We have shown in Section 5.4 that we can automatically mine a part of the ’Cell Line Cells’ content ODP. We have also mined frequent CFPs which contain specific domain vocabulary (Table 8). We have also shown that we can mine patterns that contain mostly variables – corresponding to a subtype of structural ODPs, namely logical ODPs. For example see Fig. 16.
There are two types of correspondence ODPs: re-engineering and alignment ODPs. Our method cannot mine re-engineering patterns, which represent transformation rules to create a new ontology from elements of a source model. However, our method can mine some of the alignment ODPs, in particular, those that express class equivalence and class subsumption. Our method can detect if an ontology reuses parts of another ontology, which comes from a different namespace. For example see Fig. 17.
Possible reasons for pattern occurrence
Throughout the paper, we mentioned possible reasons for which patterns occur in ontologies, summarized as follows:
Possible uses
We envision several uses of the methods and findings in this paper. First, our approach can be used to extract frequent fragments (APNV) from sets of ontologies – like the one shown in Table 6. These fragments may form generic reusable modules that might benefit the development of other ontologies. Second, ontology authors may run the mining algorithm to discover implicit patterns in ontologies that are developed collaboratively, and potentially adopt some of these patterns as recommended practices. Third, the mined patterns may be inspected manually, and then submitted to one of the online pattern repositories to enable their reuse. And fourth, the mined patterns can be used to create custom user interfaces – for example, in the form of templates – to enable their easier authoring and error checking. For instance, a custom user interface may allow only the entry of constructs that are conforming to the pattern definition, and thus, possibly, reducing authoring errors.
Our approach versus RIO
The RIO method developed by Mikroyannidi et al. [21,22] computes clusters of ontology entities. Then, for each cluster, it computes a set of axiom generalizations. Each generalization has an associated set of one or more matching axioms, which contain an entity from the cluster. A cluster aggregates axiom generalizations, which describe similar usages of subsets of clustered entities in the axioms. In contrast to our approach, generalizations within one cluster may involve largely disjoint sets of clustered entities. An entity may also appear in an axiom in various positions, both on the left-hand side and the right-hand side of the axiom.
We use the VIVO ontology to exemplify the differences between our approach and RIO’s. One of the clusters generated by RIO for the VIVO ontology gathers 9 entities. Five of these entities also match our class frame pattern (CFP) shown in Table 8. The RIO cluster is described by 43 axiom generalizations, which also include 2 axiom generalizations that correspond to the axiom patterns that we mined as a CFP for VIVO. However, the axiom generalizations in RIO only characterize subsets of the entities. Without further analysis, it is impossible to know what is the overlap between the generalizations. In this particular cluster, most axiom generalizations (more than a half of them – 22) cover only single axioms. One of the axiom generalizations from this cluster is ”
Limitations
Our approach has several limitations. One limitation is that we can only discover patterns occurring in the ontology itself. That is, we can only discover what is frequently expressed through ontology axioms. Please note that not everything, which is expressed visually in the ODPs from literature (e.g., using UML), can be represented with the types of OWL axioms that we consider in our approach. The reason is that these OWL axioms have a tree-shaped, and variable-free form. As a consequence, our mined class–frame patterns also have a tree-shaped form. In addition, it is important to notice that the ODPs proposed in the literature are just a recommendation, and the actual ontology modeling may not entirely conform to the recommended patterns.
Another limitation is also related to the tree-shaped form of the OWL axioms, and the effect of our two–step mining process of the class frame patterns. We mine class frames on top of the already discovered frequent axiom patterns. It might be the case that the variables appearing in a class frame pattern (as part of different axiom patterns) refer to the same entity. However, we cannot say currently whether this is the case. We can also not mine cyclic patterns. The motivation for our two-step method is to make the mining of class frames computationally feasible. Without this constraint, the search space for data mining algorithms becomes prohibitively large.
Although we are able to detect (emerging) design patterns automatically, our method cannot confirm whether a mined pattern is indeed a fragment of an ODP, and this needs to be confirmed manually.
Conclusions
In this paper, we described a two-step approach for automatically detecting axiom patterns in ontologies. Our approach is able to detect three different types of patterns: axiom patterns with variables, axiom patterns without variables (a.k.a., ontology fragments), and class frame patterns. We described the two methods used in our approach: (1) a tree mining method for discovering frequently recurring ontology axiom patterns; and (2) an association analysis method to discover frequent class frame patterns. We conducted an experimental analysis on a corpus of 331 BioPortal ontologies, and found that all ontologies in the corpus contain at least one of the three types of patterns. We also extracted ‘emerging’ design patterns (frequent class frame patterns) from the ontology corpus. We could confirm manually that some of these patterns are fragments of ODPs documented in the literature. Our approach is generic, and can be applied to ontologies from any domain.
As future work, we would like to explore application scenarios that would benefit from some form of inference, for which we would extend our approach to take such inference into account. We would also like to further apply and test our methods on other ontology repositories. We envisage that our data-driven approach for identifying ontology patterns will help expose emerging design patterns and potential ontology modules, and it will ultimately lead to a better reuse across ontologies in all domains.
Footnotes
Acknowledgements
This work was partially supported by the PARENT-BRIDGE program of Foundation for Polish Science, co-financed from European Union, Regional Development Fund (Grant No POMOST/2013-7/8). Agnieszka Ławrynowicz acknowledges the support from the National Science Center (Grant No 2014/13/D/ST6/02076). This work is also supported in part by grants GM086587 and GM103316 from the US National Institutes of Health.
Appendix
(Continued)
| Size |
|
Fragment (URIs and labels) | Ontologies |
| 8 | 9 | BCO, ERO, IAO, OBCS, OBIB, OBI, OPL, PCO, SDO | |
| ’obsolescence reason specification’ |
|||
| 8 | 9 | ADAR, ADO, BFO, CAO, ERO, HUPSON, OPL, PCO, SDO | |
| ’processual_entity’ |
|||
| 8 | 5 | BCO, OBCS, OBI_BCGO, OBIB, STATO | |
| ’action specification’ |
|||
| 8 | 4 | OBI_BCGO, OBIB, OBI, STATO | |
| ’nucleic acid extraction’ |
|||
| 7 | 9 | ADAR, ADO, BFO, CAO, ERO, HUPSON, OPL, PCO, SDO | |
| ’spatial_region’ |
|||
| 7 | 6 | BCO, OBCS, OBI_BCGO, OBIB, OBI, STATO | |
| ’specimen collection process’ |
|||
| 7 | 6 | OBCS, OBI_BCGO, OBIB, OBI, OBIWS, STATO | |
| ’organism’ |
|||
| 7 | 5 | OBCS, OBI_BCGO, OBIB, OBI, STATO | |
| ’measure function’ |
|||
| 7 | 5 | OBCS, OBI_BCGO, OBI, OBIWS, STATO | |
| ’sequence data’ |
|||
| 7 | 5 | OBCS, OBI_BCGO, OBIB, OBI, STATO | |
| ’processed material’ |
|||
| 7 | 4 | CL, OBI_BCGO, TAO, VSAO | |
| ’secretory cell’ |
|||
| 7 | 4 | OBCS, OBI_BCGO, OBIB, STATO | |
| ’information carrier’ |
