Abstract
The need to examine the behavior of different user groups is a fundamental requirement when building information systems. In this paper, we present Ontology-based Decentralized Search (OBDS), a novel method to model the navigation behavior of users equipped with different types of background knowledge. Ontology-based Decentralized Search combines ontologies and decentralized search, an established method for navigation in social networks, to model navigation behavior in information networks. The method uses ontologies as an explicit representation of background knowledge to inform the navigation process and guide it towards navigation targets. By using different ontologies, users equipped with different types of background knowledge can be represented. We demonstrate our method using four biomedical ontologies and their associated Wikipedia articles. We compare our simulation results with base line approaches and with results obtained from a user study and find that our method produces click paths that have properties similar to those originating from human navigators. The results suggest that our method can be used to model human navigation behavior in systems that are based on information networks such as Wikipedia.
Introduction
One of the challenges in building information systems is the need to develop interfaces suited to a range of different types of users. Different types of users, such as novices, experts, generalists or specialists will, in general, display considerably different knowledge about a given domain. This specific knowledge in turn influences their interactions with an information system. Gaining insight into human navigation behavior supports the construction of easy-to-use software and information systems that are ready to accommodate a broad range of user types.
In this paper, we investigate ways of modeling navigational behavior of human users in information networks. Humans navigating an information network (such as Wikipedia) generally do not know the network topology in its entirety. They are therefore not always familiar with the global network structure but navigate based on assumptions and local information only. Experiments by Stanley Milgram and others [21,31] have shown that humans are very effective at finding short paths based on local information in offline as well as in online
In this paper we present a novel method for simulating human navigational click behavior in
This method is new in that it uses an explicit representation of the background knowledge in the form of an ontology. Research in psychology suggests that humans store concepts in their minds hierarchically [7].
Research questions
In this work, we will address the following three research questions:
RQ1
Can ontologies contribute useful information to modeling navigation in information networks? And how does OBDS using real-world ontologies perform in comparison to OBDS using randomly generated ontologies and random walks?
RQ2
Does Ontology-based Decentralized Search (OBDS) produce valid results, i.e., are the simulated navigation paths similar to those produced by human navigation?
RQ3
When using OBDS, what ontology is best suited to produce human-like navigation results?
To demonstrate our method, we use the information network formed by a set of biomedical Wikipedia articles and the connections (hyperlinks) between them. We show that several different biomedical ontologies can be used as background knowledge to inform navigation simulations, much as humans use their acquired knowledge for navigation.
Contributions
Our main contribution is the demonstration of the general suitability of existing real-world ontologies to inform models of human navigation, specifically decentralized search on information networks such as Wikipedia. To the best of our knowledge, our work presents a novel method and a novel application of ontologies. By comparing the navigational paths generated by our simulations with several baseline approaches and with data obtained from a user study, we show that our method yields results similar to those produced by actual human users. The results suggest that OBDS can be used to simulate human navigational behavior in information networks, which can be useful for addressing issues arising in the development of systems that are based on networked information. These findings are relevant for researchers interested in new applications for ontologies and for researchers interested in modeling navigation in information networks using ontologies as background knowledge.
The rest of this paper is structured as follows: In Section 2 we place our work in the context of previous research and related work. In Section 3 we discuss materials and methods, and we present the results in Section 4. We end with a discussion.
Related work
In the context of this paper, the related areas can be divided in three: navigation in social networks, navigation in information networks and ontologies.
Navigation in social networks
This paper particularly addresses navigation in social networks via decentralized search algorithms. Fundamentally, decentralized search describes a way of solving a pathfinding problem in a social network. Starting from an arbitrary start node (i.e., a person) within the network, the objective of decentralized search is to find a way to a given target node. The algorithm, however, does not possess global knowledge of the network and can therefore only take decisions based on local knowledge. The term
The idea of decentralized search, as used in our navigation simulations, was made popular by Stanley Milgram’s widely discussed small-world experiment [21,31] in the 1960s. In the experiment, participants in Boston and Nebraska received a letter containing information about a target person (a Boston stock broker). They were then asked to forward the letter to one of their acquaintances, to bring the letter closer to the target person. The resulting median chain length of six intermediates for successful chains of letters coined the term “six degrees of separation”. By taking only the limited knowledge of each participant into account at each step, the search effectively constituted a form of decentralized search. The result illustrated the so-called
In 1998, Watts and Strogatz [33] characterized networks exhibiting small-world characteristics as exhibiting a high clustering coefficient and a low characteristic path length and demonstrated the actual existence of this type of small-world networks in a film actor collaboration network, the power grid of the western United States and the neural network of C. elegans, a small roundworm.
In 2000, Jon Kleinberg proved that for the type of small-world networks proposed by Watts and Strogatz [14], no effective decentralized search algorithm could exist that always found a path connecting two nodes in subpolynomial time. However, Kleinberg presented a more generalized version of the model for which he then proved that a decentralized algorithm capable of finding short paths existed.
Kleinberg later extended his model of decentralized search to include hierarchies [15], where the term
Miao et al. [20] have studied decentralized search in collaboration networks. Collaboration networks differ from information or social networks in that the information flow in them is driven by tasks. This means that the edges in the network are formed by collaboration on tasks. In their study, the tasks were software bugs. Developers who were assigned a bug they could not eliminate themselves forwarded it to another developer who they believed could handle it. By establishing several forwards in a row, this workflow can be viewed as a type of decentralized search, as all decisions about the next hop were taken independently by multiple participants.
Adamic and Adar [4] studied decentralized search in the e-mail network of the HP labs and found that decentralized search according to hierarchies based on connectedness and office cubicle distance worked best.
Decentralized search is also used in peer-to-peer file sharing protocols such as Gnutella or KaZaA. With a low characteristic path length and a high cluster coefficient, the Gnutella network displayed small-world characteristics in 2003 [18].
Navigation in information networks
In this paper, decentralized search, a navigation model originally developed for social networks, is applied to information networks.
One of the most prominent models related to search in information networks is information foraging [26]. Information foraging is based on foraging theory in biology. In order to survive, animals have adopted methods which maximize the energy gained from food sources. In the theory of information foraging, search in information networks is not guided by background knowledge but by information scent, with each article and link emanating a distinct scent, which is dependent on the target of the search. For instance, when searching for information on penguins, a link leading to an article about Antarctica would provide more scent than a link leading to an article about the Sahara desert.
In this paper, information networks are studied on the example of Wikipedia. However, genuine navigation paths from Wikipedia are difficult to obtain, as the goals of users are often hidden and not explicitly visible, and logs of click trails are hard to obtain. With 60–70%, the fraction of teleports is furthermore significantly higher on Wikipedia than on general web sites [8]. This might be due to the fact that users visit Wikipedia to satisfy specific information demands rather than to browse articles. However, there exist valid reasons to navigate Wikipedia, which will be detailed in the description of the navigation scenarios in Section 3.5.
Due to the difficulty of obtaining Wikipedia navigation paths, wiki games have been a popular replacement for Wikipedia navigation paths in recent research. Wiki games, such as Wikispeedia,1
In our own group, we have used decentralized search (with non-ontological background knowledge) in different contexts:
In 2011, Helic and Strohmaier compared the navigability of different tag hierarchy generation algorithms on data from Bibsonomy, CiteULike, Delicious, Flickr and LastFm [11]. The paper evaluated the suitability of tag hierarchies for navigation on tagging networks and proposed a novel tag hierarchy generation algorithm.
In 2012, Strohmaier, Helic et al. compared different folksonomy induction algorithms through decentralized search [29]. They showed that, based on evaluation through navigation, clustering algorithms developed for social tagging systems performed better than standard hierarchical clustering algorithms.
Helic et al. applied decentralized search to broad and narrow folksonomies on data from Mendeley [10] and found broad folksonomies better suited to supporting navigation.
Trattner et al. [30] compared decentralized search and human navigation behavior in information networks and showed that the simulation of decentralized search yielded very similar results to actual human navigation data on Wikipedia. In their work, Trattner et al. investigated different types of hierarchies as background knowledge and found that decentralized search based on a hierarchy generated from network features such as in- and outdegree simulated human navigation better than comparable hierarchies generated from external knowledge.
In ongoing research, Helic, Strohmaier et al. are studying the influence of stochasticity and different methods of selecting the next hop in decentralized search [12].
The previous work did not tap into existing ontologies as background knowledge, but used other approaches (such as automated methods) for this purpose. This paper goes beyond previous research by extending the simulation framework with ontologies and by applying Ontology-based Decentralized Search to the case of Wikipedia and to concrete ontologies for the biomedical domain.
Ontologies have been used in previous research to facilitate navigation in digital libraries. Papazoglou and Hoppenbrouwers [24] have used ontologies to retrieve related work when searching digital libraries. The research of Rajapakse et al. [28] shows efforts to navigate the digitally available literature related to dengue fever. Villela Dantes et al. [32] have studied the ontology-guided insertion of links into web pages. In their work, they classified web pages according to an ontology and subsequently inserted links to related topics into web pages to facilitate navigation.
These research papers share the effort to use ontologies to aid navigation. The objective of this paper lies in explaining and modeling user behavior by using ontologies as background knowledge. The ontologies are hence not used to guide human users but to simulate and possibly explain behavior.
This paper uses three ontologies from the biomedical domain. Biomedical ontologies play an important role in biomedical research [6] and are used for a range of purposes. In the biomedical domain, ontologies have been adapted more frequently than in other disciplines [23].
Materials and methods
Introductory example

Alice’s Wikipedia Navigation Scenario. Looking for a disease, Alice goes to Wikipedia and starts from a hypothetical portal containing links to a number of common diseases. Alice then navigates her way through the Wikipedia network.
To illustrate our work, let us introduce the following example, depicted in Fig. 1:
Characteristics of the data sets used for our work. The tables display statistics about the examined ontologies as well as the set of Wikipedia articles mapping to those articles
At each step, Alice is only aware of the links leading away from the current article. She is familiar with some of the article titles, and is able to relate them to one another through what we refer to as her
To simulate Alice’s usage of Wikipedia, we first mapped a subset of biomedical Wikipedia articles to their corresponding ontology concepts in three biomedical ontologies. Given these mappings, the simulation could then calculate distance information on the ontology. For each potential outgoing link that Alice could click, the simulation computed the shortest path between the article behind that link and the target article. This distance information was used as a proxy measure to estimate the distance to the target article in the Wikipedia network. The distance information gained this way was not necessarily optimal or even correct, but generally provided a good guess to guide the navigation.
In this manner, the simulator was able to make an educated guess about what link to follow, just as Alice could roughly place the outgoing articles into categories.

Structure of the four top levels of the ontologies used in our research. The figure shows the structure for ICD-10, MeSH and SNOMED CT. The root node is displayed in black and bold in the center of each plot. The figures show all ontology concepts up until a distance of four from the root node. Color indicates distance, with red being close to the root and blue being farther away. SNOMED CT contains roughly 40,000 nodes in the subgraph displayed, which is significantly more than ICD-10 (∼12,000) and MeSH (∼10,000).
In the rest of the Materials and Methods section, we describe the ontologies used to inform the simulator (Section 3.2), the Wikipedia articles and how we obtained them (Section 3.3), our method of Ontology-based Decentralized Search (Section 3.4), the navigation scenarios (Section 3.5), the user study (Section 3.6) and the simulator implementation (Section 3.7).
We used the following three ontologies and terminologies (all from the biomedical domain) as background knowledge. With these ontologies, we were able to i) extract articles from Wikipedia and ii) guide the next-step selection in the simulator.
The
The
Table 1 displays statistics about the data sets used for this paper. The row denoted
We used a dump of the English Wikipedia from December 2011 to extract articles from the biomedical domain corresponding to ontology concepts. We then mapped the articles to the ontologies by parsing the articles’ info boxes.
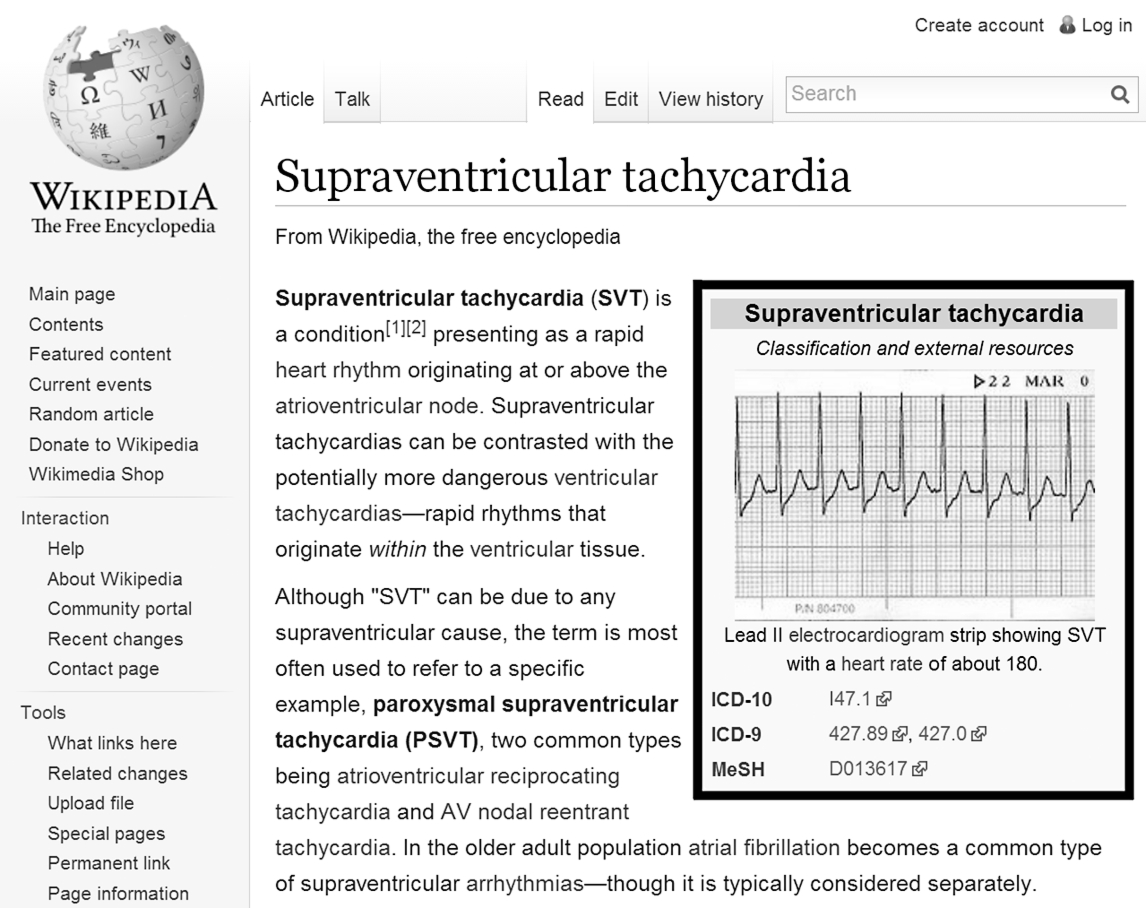
Example for an infobox template used in disease articles on Wikipedia. Disease articles commonly make use of an Infobox disease template, which offers fields for ontology codes. We used template fields in the Infoboxes to map Wikipedia articles to their ontology counterparts.
In disease articles, the
SNOMED CT is proprietary and as such not present in Wikipedia info boxes. As a consequence, we could not directly relate Wikipedia articles to the ontology concepts. We therefore used semantic mappings from BioPortal [36] to map Wikipedia articles to SNOMED CT. We mapped a total of 1,593 Wikipedia articles from both ICD-10 and MeSH to SNOMED CT with this method.
Introduction
Decentralized search is a method of solving a pathfinding problem in a network without a central control unit. Starting from an arbitrary start node within the network, the objective of decentralized search is to find a way to a given target node. The term
In a social network, the decision of where to forward the problem is generally based on the expected knowledge and capability of that particular next node (person). For our simulations, we assumed that all nodes shared a common background knowledge expressed as an ontology. This assumption made our algorithm less “decentralized” in a certain sense because all the decisions were now made by the same entity (our simulator). Just like in the original decentralized search however, at each node the simulator could only access information about that particular node’s local network neighborhood. The background knowledge represented additional knowledge about the network necessary to efficiently find a short path to the target. When looking for an employee in a company for example, this knowledge could represent the organizational hierarchy – with the restriction that the search can only be forwarded to acquainted employees, which would for instance be the case with personal recommendations.
In the theory of network navigability, Jon Kleinberg showed that networks that are formed according to a background hierarchy (i.e., a tree) are efficiently navigable [15], provided the search agent has access to that background hierarchy during the search. This method, called Hierarchical Decentralized Search, has been successfully applied in previous research ([12], [29]). This paper extends this application by using ontologies as the background knowledge.
Simulations
Based on Alice’s navigation example, we implemented our navigation simulations based on decentralized search. Recall that Alice did not remember the exact name of her navigation target, but she could roughly place it in a category. She then found her way to this target using her own background knowledge.
To be able to represent this scenario in the computer simulations, the title of the target article (but not its position in the article network) was directly known to the simulations. This modeled the same scenario – the somewhat familiar article Alice was trying to reach. The next link to click was determined by calculating the distance from the current node to the target node on the background knowledge. For instance, in the example shown in Fig. 1, if we assume the current position to be at the
To avoid loops, the simulation explored each node in the network only once. However, the simulation could backtrack to the last visited nodes (up until the starting portal, if necessary), just as Alice would use her browser’s back button. This was used in case of dead ends (articles with no unvisited outgoing links) or at articles providing only links leading further away from the target (according to the ontology information). At any given point, the simulation could also jump back to the starting portal directly, modeling a home button in an information system.
Usage of ontologies
This use of existing ontologies represents a substantial change in the motivation of the background knowledge: As opposed to previous work in this area, the background knowledge is now exogenous to the network. What this implies is that the hierarchy is based on knowledge independent of the network that the agent navigates on. All ontologies used in the application of Ontology-based Decentralized Search in this paper play a key role for their corresponding domain in their research fields. They are hence representative for a good part of the knowledge in these domains. This provides OBDS with a foundation to more accurately represent the intuitions of human navigation behavior.
The use of ontologies and the associated semantic information open up a range of new possibilities for the application of the background knowledge:
In the experiments conducted for this paper, Ontology-based Decentralized Search was used with three different ontologies (ICD-10, MeSH and SNOMED CT) that were not filtered. This meant that all concepts and relation types present in the ontologies (and mapping to the data sets) were used as the background knowledge. In terms of relation types, ICD-10 and SNOMED CT consist of is-a relations. For MeSH, which is more a controlled vocabulary than a proper ontology, the relations are of the types is-a and part-of and are both used without clear distinction. For this paper, Ontology-based Decentralized Search ignored any properties associated with the ontology concepts.
The results of the simulations were then compared on the same information network, which meant that the three ontologies effectively modeled different user groups on the same set of data. While all three ontologies represent expert knowledge, they still serve different purposes: ICD-10 is a disease classification widely used by insurance companies, physicians and hospitals. SNOMED CT is a terminology of clinical terms, and MeSH is a terminology for the indexing of journal articles in the medical domain. We hence assume these ontologies to be representative of expert knowledge in their respective biomedical fields.
Limitations
Previous research on decentralized search has studied the stochasticity of human navigation behavior [12]. As the aspiration for this work was not to improve the navigation model but to examine the suitability of ontologies to support navigation, this more complex navigation model was beyond the scope of this article.
For this work, we assumed the ontologies to be static representations of user knowledge. As the number of tasks in the users study was limited to 15 and all tasks were independent of each other, we assumed learning to be negligible. In future efforts, we would also like to include a learning component, such as users memorizing preferred paths through the network or making new associations while navigating.
The model used for this paper is also limited to a single domain, and hence to a single ontology at a time. Fulfilling multiple constraints at once is not possible as of now. Future work could look into more extensive domains that need to be covered by multiple ontologies.
Navigation scenarios
We studied two different search scenarios, both of which started from a hypothetical Wikipedia portal.
Starting portal
We started the navigation from a hypothetical Wikipedia portal featuring a selection of suitable articles: the 25 health conditions listed in the main navigation toolbar of

Starting portals used in navigation simulations. For ICD-10, MeSH and SNOMED CT we used a portal obtained by mapping navigation bar articles from
Single-target search
The first scenario was analogous to Alice’s introductory example. This models the scenario of having a vague concept in mind (but not the exact term), or having a concept on the tip of one’s tongue. The goal of single-target search is then to rediscover this concept via navigation on the Wikipedia article network.
Examples for clusters of Wikipedia articles used in exploratory search. The table shows three examples of clusters used in our simulations. We used TF-IDF features and k-means clustering to automatically group Wikipedia articles into semantically related groups of two to fifteen articles
In single-target search, the simulation started at the portal and proceeded to a single target article using Ontology-based Decentralized Search.
Multiple-target search
In the second scenario, we studied Ontology-based Decentralized Search in the context of exploratory search. In exploratory search, users explore a space of resources rather than trying to find one specific target [19]. The objective of multiple-target search is therefore to gain an overview of a certain field of concepts. One example could be to estimate a range of diseases associated with certain symptoms (such as “What diseases could show the symptoms of heart burn, fatigue and headaches?”).
In the simulations for multiple-target search, the difference to single-target search was in the targets, which consisted of target sets of two to fifteen articles (instead of only a single one). The rest of the simulation (starting portal, decentralized search, background knowledge) was conducted in the same way as the single-target search.
We used clusters of semantically similar Wikipedia articles as our target sets and applied
Maximum number of clicks
For all evaluations, a maximum number of 20 clicks for the single-target scenario and 40 clicks for the multiple-target scenario were assumed. While these limits are certainly variable in the setting of navigation, the chosen numbers represent a reasonable limit of potential clicks that a user might undergo to find the target article on an information network such as Wikipedia. Analysis of Wiki games has revealed average path lengths of fewer than seven clicks, with some games taking up to 30 clicks [9,35]. Longer paths might also skew the results, as they are more likely a result of guesswork and not of an exploitation of background knowledge.
To evaluate our simulations, we carried out a user study on Wikipedia navigation. Eight participants without any particular background in medicine were asked to navigate Wikipedia, modeling the scenario of navigating to find diseases. All of them were graduate students in different fields (but not in medicine) at Stanford University at the time of the user study.
With the OBDS approach, we assume that the ontology represents an expert with profound knowledge of the entire domain. As such, we expect experts to be able to be able to classify diseases to their approximate position in the ontology, which is precisely why the simulations have access to the entire ontologies as background knowledge. As we conducted our study with layperson users, however, we could only expect them to be only familiar with a smaller set of diseases.
The study used the data set of ICD-10, SNOMED CT and MeSH, containing 1,593 Wikipedia articles. Naturally, a large share of these articles turned out to be too specialized for participants not particularly familiar with the medical domain (with article names such as
The setup for the user study consisted of a server containing the subset of Wikipedia with an interface similar to Internet encyclopedia (see Fig. 4). The interface showed the articles used by the study, as well as information about the current task. As in our simulations, backtracking (using the back button in the browser) and jumping back to the portal by clicking a home link were enabled at all times. Each step of the users was logged.
We asked each of our participants to complete a total of 15 navigation tasks. A task consisted of finding a given target node (or a set of target nodes) in the subset of the Wikipedia network. As in the simulator, the starting point for a task was always the portal, and participants could only click on links to articles within the data set. To deal with potential frustration, participants were given the possibility to abort the current task if they had not found the target(s) after half of the maximum number of steps (20 for single targets and 40 for multiple targets).
Implementation
The experiments presented in this paper were conducted on a decentralized search simulator. This simulator was an extension of previous work by Helic, Strohmaier et al. ([11,29]) and implemented in
Results
Evaluation metrics
Based on work by Krioukov and Papadopoulos [17] we used
In accordance with Strohmaier and Helic [12], we define the
Let
We further extend these metrics with the
For all our evaluations, we assumed a maximum number of 20 clicks for the single-target scenario and 40 clicks for the multiple-target scenario.
We established comparisons with random and optimal solutions by including a random walk, randomly generated ontologies and a shortest-path solution.
Random walk
The random walk consisted of following a random link (or tracking back) at each step, not taking already visited nodes or potential targets among the neighboring nodes into account. The comparison with the random walk showed us how much more information the OBDS approach provided to the navigation in comparison to a completely random behavior.
Randomly generated ontologies
For this comparison, we constructed a randomly generated ontology counterpart to each ontology used in our simulations. To this end, we used the number of nodes and edges as input for the configuration model approach of generating a random graph with the same number of nodes and edges [22]. As the resulting graph was not necessarily connected, we subsequently randomly connected all graph components and then removed the number of additional edges created in this process from other parts of the resulting graph without deconnecting it.
This comparison showed us how much information the OBDS approach gained by taking the structure of the ontologies into account (but not yet the correct mappings). Furthermore, evaluating with randomly generated ontologies took the structured search behavior of decentralized search into account: Decentralized search, in our implementation, did not re-explore already visited nodes, could backtrack and always recognized links leading to a target node among the current node’s neighbors. This gave this method a distinct advantage over the pure random walk.
Shortest-path solution
Finally, for the comparison with the optimal approach, we computed a shortest-path solution. In the single-target scenario, this meant that we always used the shortest possible path in the graph for connecting the portal to the target node. For the multiple-target scenario, an exact solution would have required solving an instance of the traveling-salesman problem, which is computationally expensive. To circumvent this issue, we approximated the perfect solution with a nearest-neighbor approach that always took the shortest possible path to the nearest neighbor. This allowed us to compare to the (approximately) optimal solution. It is important to note that this was only possible with global knowledge of the graph topology, which users do not posses in a decentralized search scenario.
Evaluation
To compare the performance of OBDS with different ontologies as background knowledge, we evaluated multiple ontologies on the same set of Wikipedia articles. This allowed us to inspect the ontologies side by side, facilitating comparison. Figure 5 (left column) displays the results for OBDS using ICD-10, MeSH and SNOMED CT.
Figure 5 shows that the success ratio for OBDS with ICD-10, MeSH and SNOMED CT was well above both the random walk and OBDS with the randomly generated ontologies. When comparing the biomedical ontologies, the results show that OBDS with ICD-10 performed best, followed by MeSH and SNOMED CT for the success ratio. Cochran’s Q test showed statistically significant differences for the success ratios (
For the stretch, ICD-10 lead to the best results followed by SNOMED CT, which in turn fared slightly better than MeSH (all differences statistically significant by Friedman tests with Post-hoc Wilcoxon signed-rank tests with Bonferroni correction, all for
For the accumulated success ratio, ICD-10 and MeSH informed the navigation significantly better than SNOMED CT. In addition, ICD-10 was better than MeSH up until close to the maximum number of steps. OBDS with the biomedical ontologies performed better than with randomly generated ontologies, which in turn were better than the random walk (all differences statistically significant by Friedman tests with Post-hoc Wilcoxon signed-rank tests with Bonferroni correction, all for
One explanation for the variations in the performance for ICD-10, MeSH and SNOMED CT could be the different popularity of the ontologies. As the most widely used statistical classification system in the world, ICD-10 evidently has the highest influence on Wikipedia editing behavior. Not only do most disease articles reference their corresponding ICD-10 code, but also disease categories are arranged based on the ICD-10 model. This could explain why ICD-10 is slightly better suited to navigate the encyclopedia’s article network in these evaluations. Many disease articles on Wikipedia also reference MeSH, making the same arguments apply for it as well. While both MeSH and ICD-10 are available freely online, SNOMED CT is not available in all countries and perhaps less-known in the Wikipedia community. SNOMED CT is also not referenced in the infobox templates of disease articles, making it perhaps a poorer candidate to inform the navigation.
User study

Success ratio, stretch and accumulated success ratio for ICD-10/MeSH/SNOMED CT and the user study. The first column shows the results for ICD-10, MeSH and SNOMED CT, the second column the results for the user study. The rows show
For the user study, we evaluated the results of human navigators on a set of 100 manually selected targets and 20 manually selected clusters. We then compared the performance of OBDS, the computer-generated random baselines and the optimal solution on the same targets. Note that the subset of targets was a maximum distance of three hops away from the portal. The evaluations hence do not include any data points for longer shortest paths.
Figure 5 (right column) shows that the success ratio for the user study was fairly close to the simulator. For the single-target scenario, the overall success ratio was 92% for the user study and ranged from 79% to 91% for OBDS supported by the three biomedical ontologies. Cochran’s Q test showed statistically significant differences (

Path lengths produced by the user study and the simulator. The figures show the resulting path lengths for the single-target (a) and multiple-target (b) search scenarios. Navigation was limited to 20 and 40 steps, respectively, hence the high number of paths for these lengths (i.e., not all targets were found). The path distributions for the random walk and the randomly generated ontologies were left out for reasons of clarity.
For the single-target stretch, with an overall stretch of 1.74 the user study performed slightly better than the simulator, which displayed stretches between 1.78 and 1.84. A Friedman test showed that groups were significantly different (
For the multiple-target scenario, the accumulated success ratio showed that the user study fell within or just below the range of OBDS with one of the three biomedical ontologies. A Friedman test showed that statistically significant differences existed between the groups (
For post-hoc Wilcoxon signed-rank tests with Bonferroni correction, results varied over the tested values of
Kullback-Leibler divergence for the path length distributions produced by the simulator and the user study. The table shows the KL divergence from the user study to the ontologies and the optimal and random solutions. This metric measures the number of additional bits required to encode the original distribution, if another distribution is used in its place. The Randomly Generated Ontology column was computed using an average over the three randomly generated ontologies considered. The table shows that the user study was more similar to the ontologies than to the base lines for the single-target scenario. The results closest to the user study are displayed in bold
To obtain qualitative insight into the navigation process, we compared the resulting distribution of path lengths produced by both the user study and the simulations. This distribution can be seen in Fig. 6. We then computed the Kullback-Leibler (KL) divergence from the user study distribution to the other distributions. The Kullback-Leibler divergence measures the number of additional bits needed to encode the path length distribution, if the other distribution is used in place of the original (user study) path length distribution. The resulting values can be seen in Table 3. For the single-target search scenario, it is clearly visible that only OBDS with biomedical ontologies produced path length distributions close to the user study: All three ontology path length distributions had a very small KL divergence (0.08–0.18 bits) to the user study. This means that, as far as produced path lengths are considered, it appears justifiable to replace human navigation data with data produced by OBDS and a fitting ontology. The same cannot be asserted about randomly generated ontologies (nor the random walk or the optimal solution), which cannot be easily taken in lieu of the biomedical ontologies and yield similar results.
For the multiple-target search scenario, these claims cannot be made this clearly. However, this might be due to the path length distribution for the multiple-target scenario being rather sparse, which meant that a single path accounted for five percent of the path lengths (which is also reflected in Fig. 6b).
Details of the user study and the compared data sets. The table displays statistics about the user study and the ontologies. The most similar values to the user study are displayed in bold face. For the first three measures, we viewed the information about found targets and first hops as a vector of values, for which we calculated the angle to the vector containing the information for the user study (i.e., the cosine similarity). For the random walk, we averaged over 1000 random walks for each portal-target pair. The last two measures display the average per step usage of the back and home buttons for the different scenarios. In summary, the results confirm that what has appeared somewhat apparent from the success ratios and the stretch, i.e., that ICD-10 and MeSH displayed the most similar behavior to the user study. The randomly generated ontology column was computed using an average over the three randomly generated ontologies considered. The number of clicks ranged from 195 to 2000 for the single-target scenario (user study: 485) and from 111 to 800 fro the multiple-target scenario (user study: 419)
Details of the user study and the compared data sets. The table displays statistics about the user study and the ontologies. The most similar values to the user study are displayed in bold face. For the first three measures, we viewed the information about found targets and first hops as a vector of values, for which we calculated the angle to the vector containing the information for the user study (i.e., the cosine similarity). For the random walk, we averaged over 1000 random walks for each portal-target pair. The last two measures display the average per step usage of the back and home buttons for the different scenarios. In summary, the results confirm that what has appeared somewhat apparent from the success ratios and the stretch, i.e., that ICD-10 and MeSH displayed the most similar behavior to the user study. The randomly generated ontology column was computed using an average over the three randomly generated ontologies considered. The number of clicks ranged from 195 to 2000 for the single-target scenario (user study: 485) and from 111 to 800 fro the multiple-target scenario (user study: 419)
In addition to the path length distributions, we analyzed further aspects of the user study in comparison with OBDS (see Table 4).
First, we looked into the found targets and the first click after the portal. To compare these, we arranged the nodes into vectors and computed cosine similarities.
For the found targets, all three ontologies displayed high cosine similarity values. This reflects the results from Fig. 5, and is caused by high success ratios for the limited target set used in our user study which leads to the majority of the vectors containing ones at the same positions. As discussed in the examination of the success ratios, the user study differed statistically significantly from the random walk in both single and multiple-target search and in addition from the randomly generated ontologies in the single-target case (all
For the first hops (i.e., the very first click in the search), the clicks were distributed rather evenly. A uniformly random distribution would see each link clicked 3.7% of times. Our results showed distributions ranging from 1 to 17% and were thus fairly evenly distributed, explaining the values of the cosine similarity being in the same range for all groups. For the single-target scenario, ICD-10 displayed the most similar values to the user study in the single-target scenario, where it was also close to the optimal solution. This suggests that the participants were able to select the best first hop in most cases. In the multiple-target scenario, the user study was less similar to the other groups.
In addition to calculating similarities, we also inspected the average per-step probability of backtracking or clicking the home button. Statistical tests were conducted with Friedman tests with post-hoc Wilcoxon signed-rank tests with Bonferroni correction.
Both the simulation and the users had access to a back button (leading to the previously visited page) and a home button (leading back to the portal) at all times. As it turned out, the simulations used the home button only immediately after having found a target in the multiple-target search scenario. In all other cases, the best strategy given by our simulation constraints was backtracking. The user study showed different behavior from the simulator in several aspects: For single-target search, users backtracked less frequently (9% of clicks were back button clicks, versus 11–13% for the simulations, significant at the
For the multiple-target search, users backtracked more frequently (27% versus 17–18% for the simulator and used the home button less frequently (1% versus 2–3%). However, this was not statistically significant, possibly due to the small sample size.
In conclusion, backtracking appeared to be the most widely applied strategy for navigating out of dead ends and backtrack from less promising areas of the network.
Discussion
In this work, we studied simulated user navigation behavior via decentralized search. We introduced Ontology-based Decentralized Search (OBDS), a novel navigation simulation method based on decentralized search which uses ontologies as background knowledge. We showed that our method can be successfully applied to navigation in information networks, and demonstrated that it can be applied on Wikipedia when supported by biomedical ontologies.
In the following, we want to focus our discussion on the research questions raised in this work:
RQ1
Can ontologies contribute useful information to modeling navigation in information networks? And how does OBDS using real-world ontologies perform in comparison to OBDS with randomly generated ontologies and random walks?
We found that ontologies can indeed inform navigation in information networks. OBDS with medical ontologies as background knowledge was able to outperform the random baseline approaches significantly.
RQ2
Does Ontology-based Decentralized Search (OBDS) produce valid results, i.e., are the simulated navigation paths similar to those produced by human navigation?
We addressed this question by comparing properties of the simulated navigation paths with properties produced by humans in a study. We found that the click paths produced by OBDS matched success ratios, stretches and accumulated success ratios of human paths better than pure random walks and OBDS with randomly generated ontologies. Furthermore, the resulting path length distribution of the user study was best matched by OBDS with biomedical ontologies.
RQ3
When using OBDS, what ontology is bested suited to produce human-like navigation results?
From our results, it seems that ICD-10 and MeSH seem to perform best. However, the overall differences between the ontologies were not very strong, and it is subject of ongoing research to further identify differences in the performance of OBDS with different ontologies.
Further comments
We’ve limited our work to ontologies and Wikipedia articles from the biomedical domain in this paper. In this domain, ontologies have been adapted more frequently than in other disciplines [23], play an important role in biomedical research [6] and are used for a range of purposes. Without this frequent use, the results might be different in other domains. Another important aspect was the ready availability of infobox templates on Wikipedia articles, which facilitated the mappings to the ontologies. However, the principles of our method apply for other domains as well.
Influence of ICD-10
The International Classification of Diseases (ICD-10) has found widespread use and probably influences and inspires Wikipedia editors. On Wikipedia, disease articles are almost always indexed by ICD-10 as the first entry in the article infoboxes. Furthermore, the category system for the disease articles of the English Wikipedia is organized according to ICD-10. These two facts and the wide use of ICD-10 have quite possibly also influenced the link creation behavior on the encyclopedia as well as the general knowledge of the test participants. This might be an explanation of why ICD-10 seems to be best suited to model human navigation behavior in our case study.
User study
In comparison to the simulator’s performance, participants in the user study performed better for single-target search and worse for multiple-target search. This is also influenced by the fact that users aborted 30% of their multiple-target navigation tasks before having found all the targets, while the simulations ran for whole number of possible steps.
Building user models
By using different ontologies as background knowledge, our results could help researchers and engineers build and evaluate user interfaces with different user types. The ontologies compared in the results were rather similar and mostly shared the same domain. In future work, it will be interesting to compare ontologies that do not cover the entire domain, modeling specialist users, or combining ontologies to form a more complete coverage of the domain. Another idea might be to prune the ontologies at a certain depth, modeling broad generalist knowledge that does not extend beyond a certain depth.
Action selection
The simulations in the form we presented followed a deterministic greedy action selection model, in that it always selected the most profitable link according to the background knowledge. Related research has shown that users might be better modeled using epsilon-greedy action selection mechanisms with dynamically changing epsilons [12]. In follow-up research, our work could be extended with stochastic action selection mechanism such as epsilon-greedy. This would also lead to another potentially crucial aspect of the present simulations, namely the need to evaluate games multiple times with potentially varying results. One could expect that these adaptations would help to finetune and validate any future attempts at modeling human navigation. However, we leave this task to future research.
Future work
The user study we presented was limited in that it was restricted to a subset of target nodes because of the requirement that the target be familiar to participants without a medical background. Since the simulation results for these targets were very close to the results of the participants, it can be hypothesized that the user behavior for the whole set of targets is likewise similar. It is up to future research to show more details of the comparison of human users and decentralized search.
Another aspect was the limitation of the user study to a subset of the target articles. The user study tried to approximate non-medical students with expert biomedical ontologies. While this worked to a certain extent, it will be interesting to see further user studies with medical experts and compare their results on the entire data. Since the user study was limited to eight participants, conducting a similar study with a larger number of users could reveal more interesting results, especially for the multiple-target scenario.
The chosen portal, based on WebMD.com, undoubtedly influenced the navigation results. It is up to future work to compare different portals and shed a light on possible differences.
The idea to navigate to one single predefined target might seem somewhat artificial in the case of user behavior concerning explorative tasks. However, one idea to improve on this might be to calculate the TF-IDF features of the target node beforehand and subsequently navigate until a page (or a number of pages) similar enough to the TF-IDF features has been found (which does not need to be the predefined target page). This could model the case of users exploring areas of the network.
Other potential research questions might include the limitation of visible links to links in the upper part of Wikipedia articles and comparing the results on non-English editions of the encyclopedia. Past research [13] has already compared different methods of extracting background knowledge from the actual network used for navigation. These background knowledges were based on network features such as centrality or degree. It would be interesting to directly compare these extracted background knowledges with ontologies and analyze the differences.
Conclusions
In this work, we have presented a novel, ontology-based method (Ontology-Based Decentralized Search) for simulating human navigation in information networks such as Wikipedia. Our results provide technical answers to several questions regarding the use of ontologies in decentralized search: We have not only presented a method to integrate ontological background knowledge into decentralized search, but also found that ontologies can serve as
Summarizing, our findings are relevant for researchers interested in new applications for ontologies or interested in modeling navigation in information networks using ontologies as background knowledge.
Acknowledgements
This research was supported by the Austrian Science Fund (FWF) grant P24866 and by NIH grant U54-HG004028. Part of the research of the first author was carried out during a visit at the Stanford Center for Biomedical Informatics Research and was supported by a Marshall Plan Scholarship granted by the Austrian Marshall Plan Foundation.
