Abstract
Background
Multiple system atrophy (MSA) and Parkinson's disease (PD) are caused by misfolded α-synuclein spreading throughout the central nervous system. While familial PD is linked to several α-synuclein mutations, no mutations are associated with MSA. We previously showed that the familial PD mutation E46K inhibits replication of MSA prions both
Objective
Here we sought to further interrogate α-synuclein misfolding to identify the structural determinants that contribute to MSA strain biology.
Methods
We engineered a panel of cell lines harbouring both PD-linked and novel mutations designed to identify key residues that facilitate α-synuclein misfolding in MSA. We also used Maestro
Results
In many cases, our modelling accurately identified mutations that facilitated or inhibited MSA replication. However, Maestro was occasionally unable to predict the effect of a mutation, demonstrating the challenge of using computational tools to investigate intrinsically disordered proteins. Finally, we used our cellular models to determine the mechanism underlying the E46K-driven inhibition of MSA replication, finding that the E46/K80 salt bridge is necessary to support α-synuclein misfolding.
Conclusions
Our studies used a structure-based approach to investigate α-synuclein misfolding, resulting in the creation of a powerful panel of cell lines that can be used to interrogate MSA strain biology.
Plain language summary
In patients with Parkinson's disease (PD) and multiple system atrophy (MSA), the protein α-synuclein misfolds into distinct shapes, or strains, causing accumulation of protein aggregates in the brain. Increasing evidence indicates that the shape α-synuclein adopts determines which disease a patient will develop. As a result, it is pivotal that we understand the factors that contribute to protein misfolding in disease. In this study, we used computational modelling to predict the effect of PD-causing and novel mutations on α-synuclein misfolding into the MSA disease conformation. We then tested these predictions in cell lines expressing the same mutations to determine if α-synuclein isolated from MSA patient samples can induce protein aggregation in the presence of each mutation. Using this approach, we not only identified key mutations in the α-synuclein gene that influence the ability of the protein to misfold into the MSA strain, but we also determined the mechanism by which one of these mutations, the PD-causing E46K mutation, exerts its inhibitory effect on α-synuclein misfolding in MSA.
Introduction
Synucleinopathies are a group of neurodegenerative movement disorders, including Parkinson's disease (PD) and multiple system atrophy (MSA), that are defined by distinct clinical and pathological features arising from the accumulation of misfolded α-synuclein proteins in the brain. While PD patients develop abnormal gait, resting tremor, muscle rigidity, and REM sleep behaviour disorder,
1
MSA patients are distinguished by the presence of autonomic failure, which can present as orthostatic hypotension, difficulty controlling urinary movement, and impaired body temperature regulation.
2
Additionally, while PD is more prevalent and patients typically develop disease in their 60 s or later, MSA is a rare disease with an earlier age of onset.1,2 Along with these clinical differences, PD is defined by α-synuclein accumulating into Lewy neurites or Lewy bodies (LB) within neurones,
3
whereas the pathological hallmarks of MSA include glial cytoplasmic inclusions (GCI) in oligodendrocytes and, to a lesser extent, neuronal cytoplasmic inclusions in neurones.
4
Interestingly, while both diseases are defined by the presence of α-synuclein inclusions,5,6 mutations in the α-synuclein gene,
Recent advances in cryo-electron microscopy (cryo-EM) enabled resolving the structure of misfolded α-synuclein fibrils from human patient samples, resulting in confirmation that α-synuclein adopts distinct structures in MSA 7 (Figure 1) and PD 8 patients (Supplemental Figure 1A). The α-synuclein fibrils isolated from MSA patient samples are defined by the presence of two asymmetrically arranged protofilaments, both of which adopt a Greek key motif that is stabilized by a salt bridge between residues E46 and K80 (Figure 1). Additionally, several positively charged residues project into a pocket at the protofilament interface that contains a negatively charged non-protein density. 7 It should also be noted that three fibril structures were resolved from MSA patient samples (PDB IDs 6XYO, 6XYP, and 6XYQ), which contain variability in subunit A but a mostly consistent structure in subunit B. In contrast, the α-synuclein Lewy fold contains a single filament with a modified Greek key motif stabilized by a salt bridge between E35 and K80 (Supplemental Figure 1A). The Lewy fold also contains a negatively charged non-protein density; however, in this structure it acts to stabilize the positive charges from four internal facing lysines, none of which include residue E46. 8
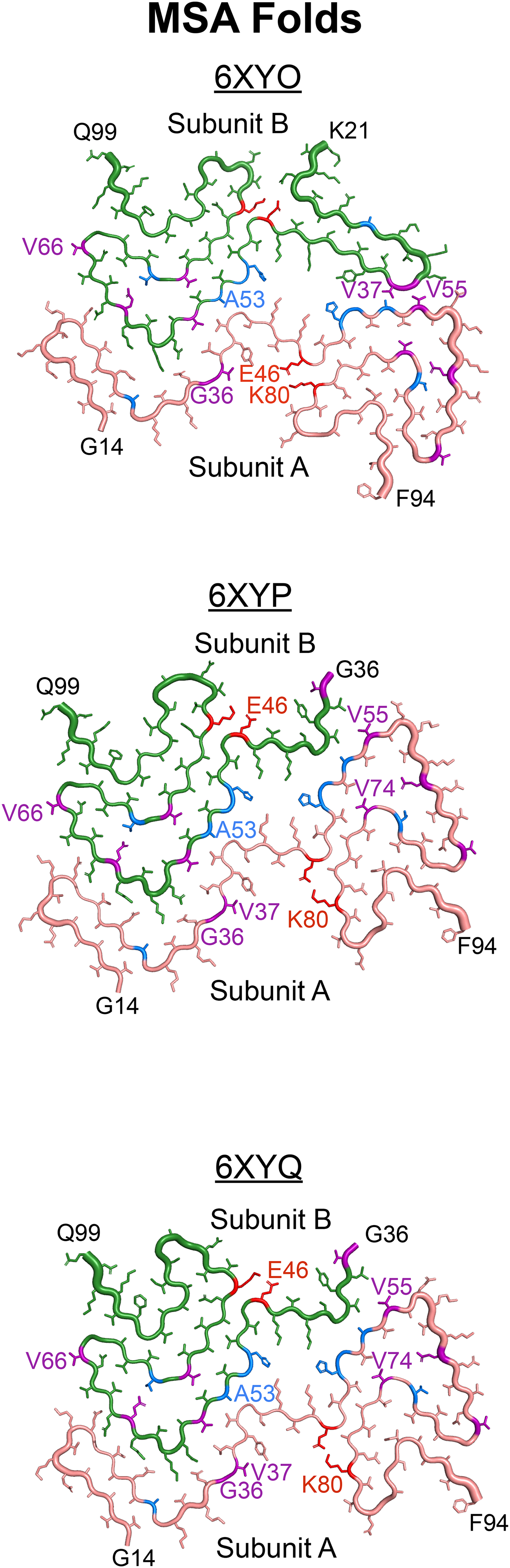
Using the MSA cryo-EM structures as an opportunity to develop hypotheses about how α-synuclein misfolds in disease, here we sought to test the effect of both PD-causing and novel mutations designed to interfere with protein misfolding on MSA propagation
Materials and methods
Human tissue samples
The Massachusetts Alzheimer's Disease Research Center (MADRC) supplied frozen midbrain tissues from two patients with neuropathologically confirmed MSA and two control samples. The NIH NeuroBioBank supplied tissue from two control patients, and the Sydney Brain Bank supplied frozen samples from three neuropathologically confirmed MSA cases. Demographic information about the patient samples used is included in Supplemental Table 1.
Cell line development
Wild-type (WT) and mutant (E46K, H50Q, G51D, A53E, G36K, V37F, V55Y, E61Q, V66F, V74I, and V74P single mutations and the A30P,A53T double mutation) human α-synuclein cDNA sequences fused to enhanced yellow fluorescent protein by an 18 amino acid flexible linker (EFCSRRYRGPGIHRSPTA) were synthesized and cloned into the pcDNA3.1(+) expression vector by GenScript. The α-syn140-YFP sequences were then subcloned into the pIRESpuro3 vector (Takara) using restriction cloning with EcoRV (5’) and NotI (3’). Gene sequence and insertion were confirmed by Sanger sequencing before subsequent use. Other mutant constructs (A30G, A53V, T72M, K80E, K80N, K80Q, K80W, and the double mutant E46K,K80E) were generated by PCR amplification of the pIRESpuro3 vector containing the WT α-syn140-YFP sequence using primers designed to introduce each mutation, or the E46K mutation for the double mutant. Gene sequence and insertion were confirmed by Sanger sequencing before subsequent use.
HEK293T cells (ATCC) were cultured in Dulbecco's modified Eagle's medium (DMEM; Corning) supplemented with 10% fetal bovine serum (FBS), 100 μg/mL penicillin, and 100 μg/mL streptomycin (ThermoFisher), referred to hereafter as complete media. Cultures were maintained in a humidified atmosphere of 5% CO2 at 37 °C. Cells were plated at a density of 5.7 × 105 cells per well in a 6-well plate overnight in complete media before adding 1 μg of plasmid DNA incubated with 3.5 μL Lipofectamine 2000 (ThermoFisher) for 20 min. Stable cells were selected in complete media containing 1 μg/mL puromycin (ThermoFisher) for 48 h before generating monoclonal lines by limiting dilution of polyclonal cells in 384-well plates. The resulting monoclonal lines were frozen in liquid nitrogen. Lysates from the lines were collected in 1× radioimmunoprecipitation assay (RIPA) buffer containing 50 mM Tris-HCl, pH 7.5 (ThermoFisher), 150 mM NaCl (Sigma), 5 mM EDTA (ThermoFisher), 1% nonidet P-40 (ThermoFisher), 0.5% deoxycholate (ThermoFisher), and 0.1% sodium dodecyl sulfate (SDS; ThermoFisher). Cell lysates in RIPA buffer were frozen, thawed, and clarified using two low-speed spins (500 ×
α-Synuclein prion bioassay
Frozen brain samples were used to make a 10% (wt/vol) homogenate in calcium- and magnesium-free 1× Dulbecco's phosphate buffered saline (DPBS) using an Omni Tissue Homogenizer with disposable plastic soft tissue tips (Omni International). Aggregated protein was isolated from the samples using phosphotungstic acid (PTA; Sigma-Aldrich) as described.9,10 Briefly, the brain homogenates were incubated with 2% (vol/vol) sarkosyl (Sigma) and 0.5% (vol/vol) benzonase (Sigma) at 37 °C under constant agitation (850 RPM). Sodium PTA, dissolved in 10% (wt/vol) in double-distilled H2O and brought to pH 7.0, was added to a final concentration of 2% (vol/vol) and incubated overnight as described. Samples were centrifuged at 13,200×
The α-syn140-YFP cell assay was performed as previously described.10,11 Briefly, monoclonal subclones stably expressing WT or mutant α-synuclein were cultured in complete media in a humid 5% CO2 environment at 37 °C. Cells were grown in 384-well plates with black polystyrene walls (Greiner) with growth medium containing 0.012 µg/well of Hoechst 33342 diluted in 1× DPBS (Thermo Fisher) to visualize nuclei. Plates with cells were incubated at 37 °C for 2–4 h. PTA-precipitated patient samples were diluted in 1× DPBS prior to incubation with Lipofectamine 2000 (Thermo Fisher) at room temperature for 1.5 h. Warmed OptiMEM (Thermo Fisher) was added to each sample prior to plating the sample in 6 replicate wells. Optimized assay conditions are reported in Supplemental Table 2. Plates were covered with a membrane to prevent drying during incubation for 4 d at 37 °C in a humidified atmosphere with 5% CO2. The plate was imaged using the MicroXLS (Molecular Devices) collecting a DAPI and FITC image (for Hoechst and YFP, respectively) from 5 areas per well. The images were analyzed with MetaXpress software using parameters developed to detect and quantify intracellular aggregates in live cells using pixel intensity and size thresholds. The measure of aggregation was calculated as the total fluorescence measured across all aggregates per cell in each well (× 106 arbitrary units [A.U.]).
Modeling α-synuclein mutations on the MSA cryo-Em structure
The MSA α-synuclein filament PDB file (6XYQ) was increased to a hetero 12-mer using MOE (Chemical Computing Group) using the build_fibril.svl. From this file, Maestro (Schrödinger) was used to optimize each amino acid side using the Predict Side Chain Panel, then minimized using the OPLS4 force field. Minimization was performed to an RMSD of 10−9 kcal/mol/Å for convergence. From the minimized structure, each mutation was simulated manually and mutations in each heterodimeric sequence were performed independently. To maintain the same ground state energy, the first step in the analysis pipeline was to create a compensating mutation at an amino acid with few neighbouring side chains to serve as a control (shown in Supplemental Table 3). Using the V55Y mutation as an example for the analysis procedure, the minimized structure underwent a computational E83Y mutation on all 6 layers of subunit A in the total fibril structure. Using this control, we computationally made a V55Y mutation and a concomitant Y83V mutation on all 6 layers. After re-predicting the position of all side chains in proximity to both the Y55 and V83 residues using Predict Side Chain panel, both the mutated and control (E83Y) structures were minimized to convergence using the OPLS4 force field. To calculate the relative energy change for each mutation (in kcal/mol), the total calculated ensemble energies for the mutated subunits were subtracted from the structures with only the compensating mutations reported in Supplemental Table 3. For V55Y, the subunit B mutations followed an identical process except that the control remained E83Y on subunit A. In some cases, the E83Q mutation was used. We note that while the E83Q mutation is known to accelerate α-synuclein misfolding kinetics in a cell-free assay, 12 this effect should not impact our structural modelling data.
Statistical analysis
Data are presented as mean ± standard deviation. Cell assay data were analyzed using GraphPad Prism software. Mean fluorescence intensity values and standard deviations were calculated for each cell line by averaging the fluorescence intensity values from the 6 technical replicate wells. Reported
Results
Parkinson's mutations exert differential effects on MSA prion replication in vitro
Building on our previous findings that the familial PD mutation E46K
13
prevents MSA prion replication both
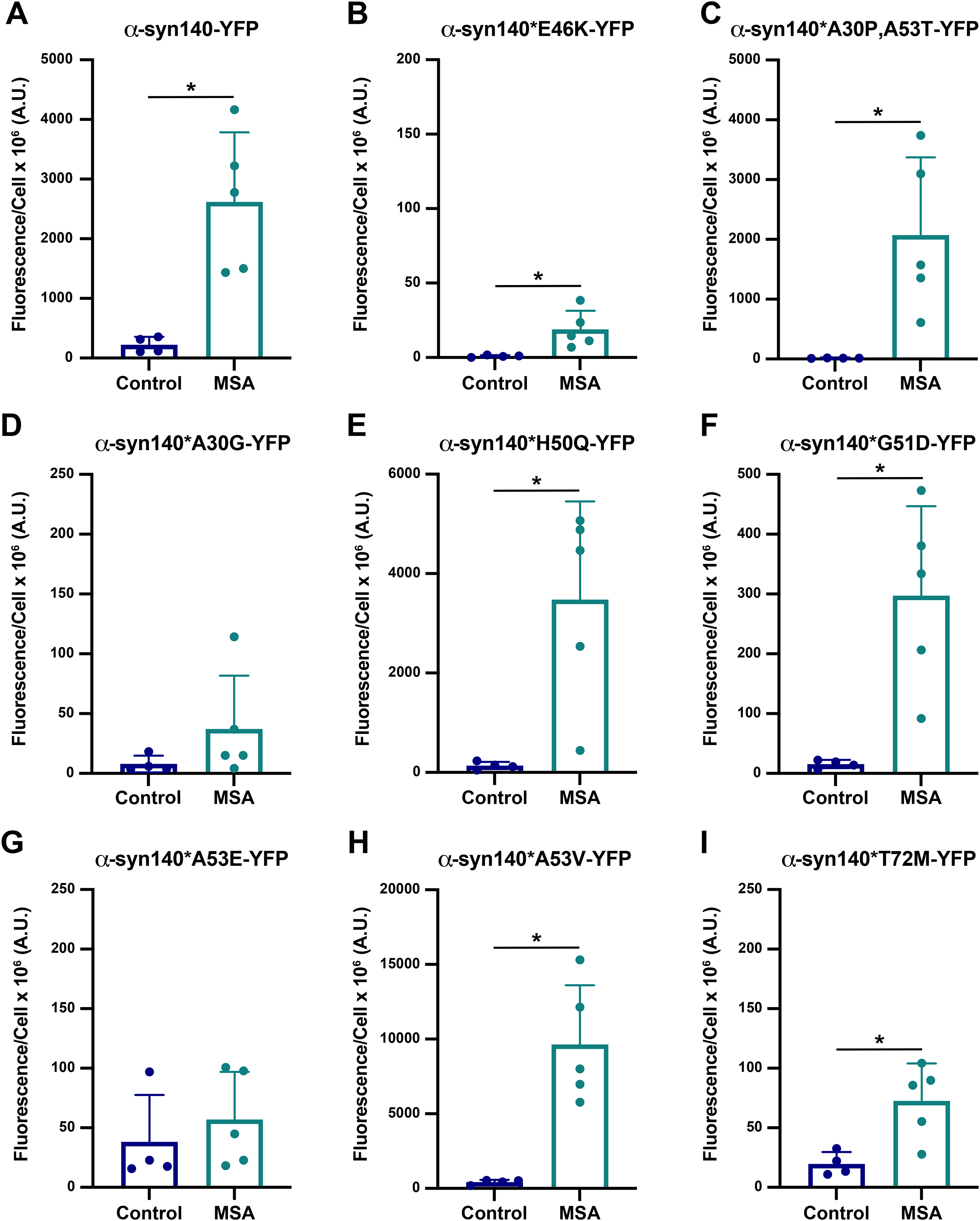
Our initial studies focused on the first three
Total energy change after mutagenesis using PD-causing mutations in MSA structure 6XYQ.
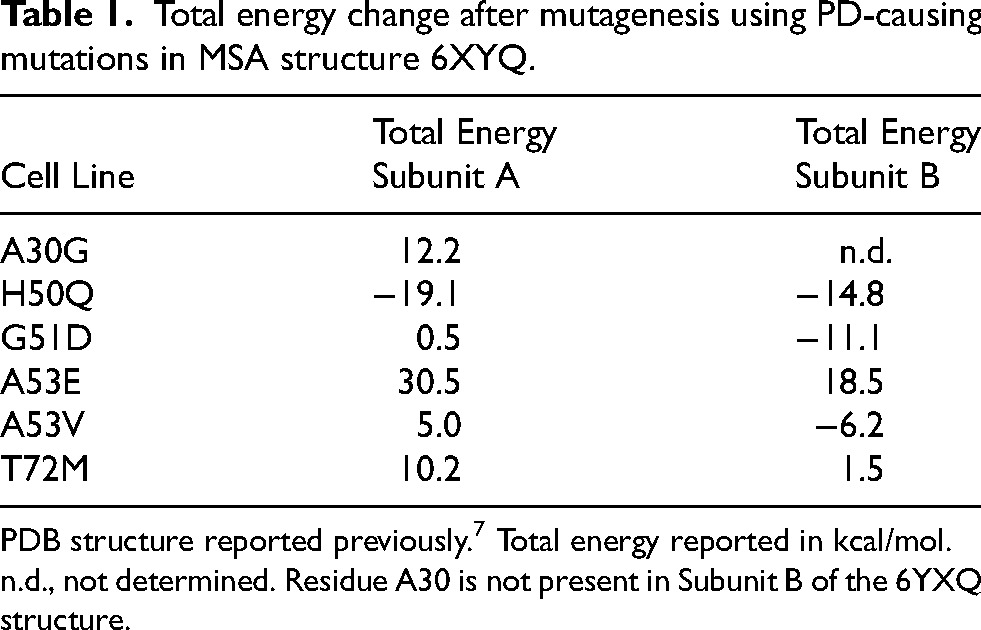
PDB structure reported previously. 7 Total energy reported in kcal/mol.
n.d., not determined. Residue A30 is not present in Subunit B of the 6YXQ structure.
To determine how well Maestro predicted the effect of α-synuclein mutations on MSA replication
Overall, the ability of Maestro to predict the effect of PD-causing point mutations on α-synuclein misfolding in MSA was reasonably accurate. Moreover, the cellular results contribute important data to the growing literature showing that the structural differences in α-synuclein fibril conformations in PD 8 and MSA 7 result in unique biological properties between the two strains.11,21–23 Unexpectedly, we found that the effect of mutations at two residues—A30 and A53—are mutation-specific. For example, while the A30P, A53T, and A53V mutations enable MSA replication, the A30G and A53E mutations block this activity. While an explanation for these residue-specific effects is not clear based on the mutagenesis modelling, future studies will focus on investigating this phenomenon.
Structurally informed mutations reveal varied biological activities between MSA patient samples
Preformed fibrils (PFFs) made from recombinant α-synuclein are widely used throughout the literature as a tool to investigate synucleinopathies, including PD and MSA. Previously, we showed that PFFs made from both WT and mutant protein exhibit unique biological and biochemical properties in cell and animal models of disease.11,14,24 These findings are supported by cryo-EM structures of PFFs, which misfold into conformations that are distinct from those found in PD and MSA patient samples (reviewed in
25
). To further investigate the differences in biological consequences that arise due to differences in MSA versus PFF structure, we developed a library of α-syn140-YFP cell lines expressing novel mutations predicted to interfere with protein misfolding in the ‘rod’ (PDB ID: 6CU7), ‘twister’ (PDB ID: 6CU8),26–28 or MSA fibril structures.
7
Comparing the rod and twister polymorphs (shown in Supplemental Figure 1B, C),
26
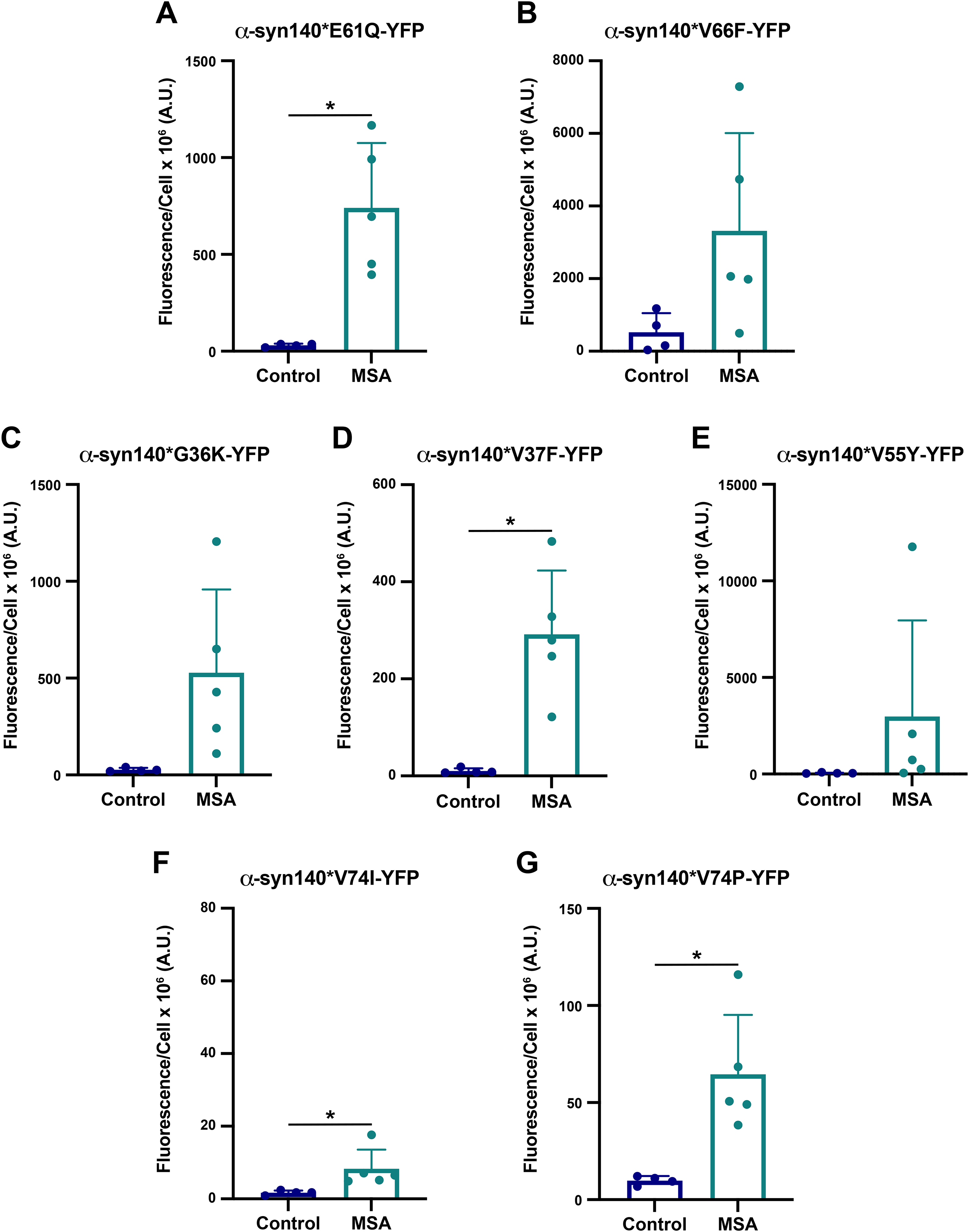
we chose to test the effect of mutations at residues E61 and V66 on MSA replication. Our initial assessment of these two structures suggested that the E61Q mutation would alter the stability of the pocket formed by the Greek key motif (which is present in most α-synuclein structures) with a greater disruption to the twister conformation than the rod, where the residues are more tightly packed together. Similarly, mutating the valine at position 66 to a phenylalanine (V66F) would disrupt the intermolecular zipper at the protofilament interface in the twister polymorph without affecting the rod, where V66 projects outward toward solvent. We subsequently used Maestro to predict the effect of these two mutations on the MSA fibril structure and found that the E61Q mutation enhanced the energetic favorability of the MSA fibril while the V66F mutation was neutral (Table 2). Inserting these mutations into the α-syn140-YFP cells, we found that all 5 MSA patient samples tested infected cells expressing the E61Q mutation (Figure 3A; Supplemental Table 5;
Total energy change after mutagenesis of structurally-informed mutations in MSA structure 6XYQ.
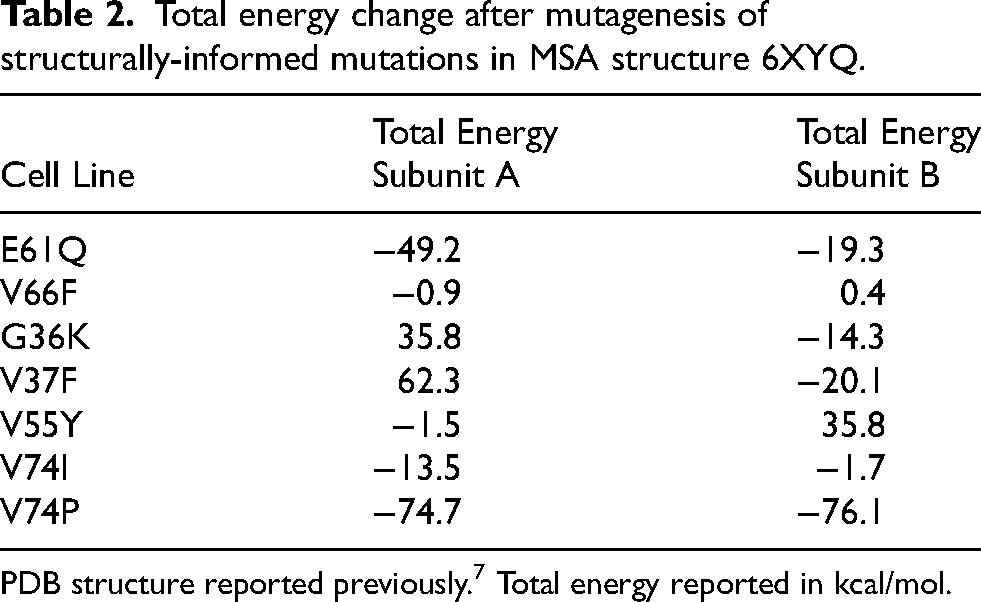

PDB structure reported previously. 7 Total energy reported in kcal/mol.
Analyzing the three MSA fibril structures resolved by cryo-EM,
7
we predicted that mutating residues G36, V37, and V55 would allow us to disrupt the interface between the two protofilaments in the MSA fibrils, with a particularly strong effect on 6XYO. To achieve this goal, we tested the G36K, V37F, and V55Y mutations both
Finally, to disrupt the twister conformation with minimal impact on MSA replication, we mutated the valine at position 74 to either an isoleucine or a proline. Consistent with this objective, Maestro modelling indicated that both the V74I and V74P mutations would reduce the total energy of the α-synuclein conformation, though the effect of the V74P mutation was much stronger, suggesting MSA should be able to propagate (Table 2). However,
It is worth noting that when we used Maestro to model the effect of mutations that were designed based on protein structures, it variably predicted our
The E46K mutation blocks MSA replication by disrupting the E46/K80 salt bridge
Several of the α-synuclein cryo-EM structures, including the MSA filaments, contain Greek key motifs that are stabilized by an intermolecular salt bridge that typically forms between residues E46 and K80 (reviewed in 25 ). In light of our previous findings that the E46K mutation blocks MSA replication,11,14 we predicted that this effect occurs because the two lysines repel one another when the salt bridge is lost, destabilizing the misfolded conformation. 29 To test this hypothesis, we generated four novel mutations designed to disrupt the E46/K80 salt bridge at position K80. In these mutations, we swapped the positively charged lysine for a negatively charged glutamic acid (K80E), predicting this would create a similar repellant interaction as the E46K mutation. We anticipated that the polar but uncharged asparagine (K80N) and glutamine (K80Q) would allow us to test the role of the positive charge on the lysine in salt bridge formation, given that the asparagine and glutamine have similar R groups to the lysine. And, finally, the bulky hydrophobic tryptophan (K80W) was selected to crowd out the packing of the R groups, preventing α-synuclein from adopting the Greek key motif. Using Maestro, we first confirmed that the E46K mutation is predicted to have a destabilizing effect on both subunits in the MSA fibril structure, with an increase of 70.2 and 51.6 kcal/mol, respectively (Table 3). We then tested the effect of the four K80 mutations using the same approach. While K80E, K80Q, and K80W all showed an increase in total energy, the K80N mutation showed a decrease in total energy for subunit A but an increase for subunit B.
Total energy change after mutagenesis targeting the E46/K80 salt bridge in MSA structure 6XYQ.
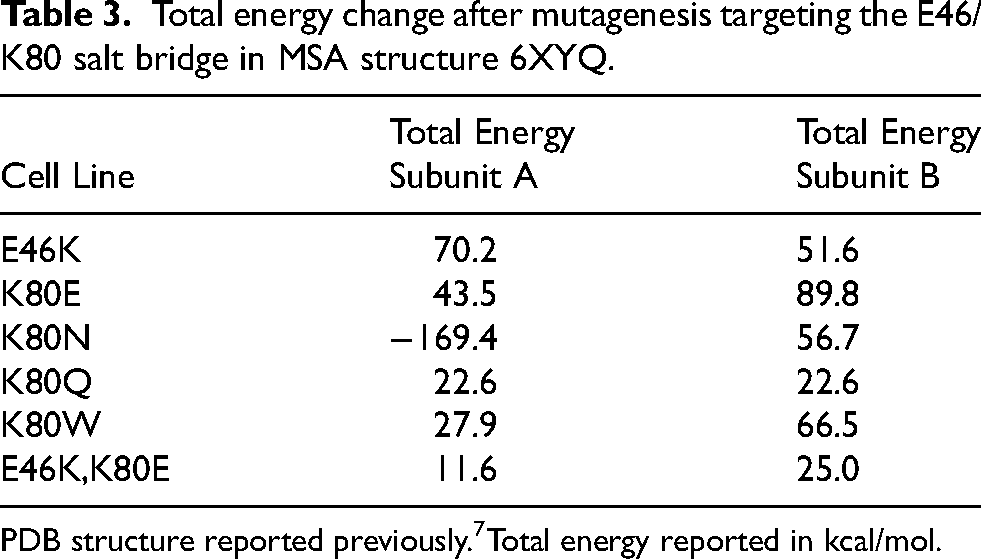

PDB structure reported previously.7Total energy reported in kcal/mol.
Testing these mutations
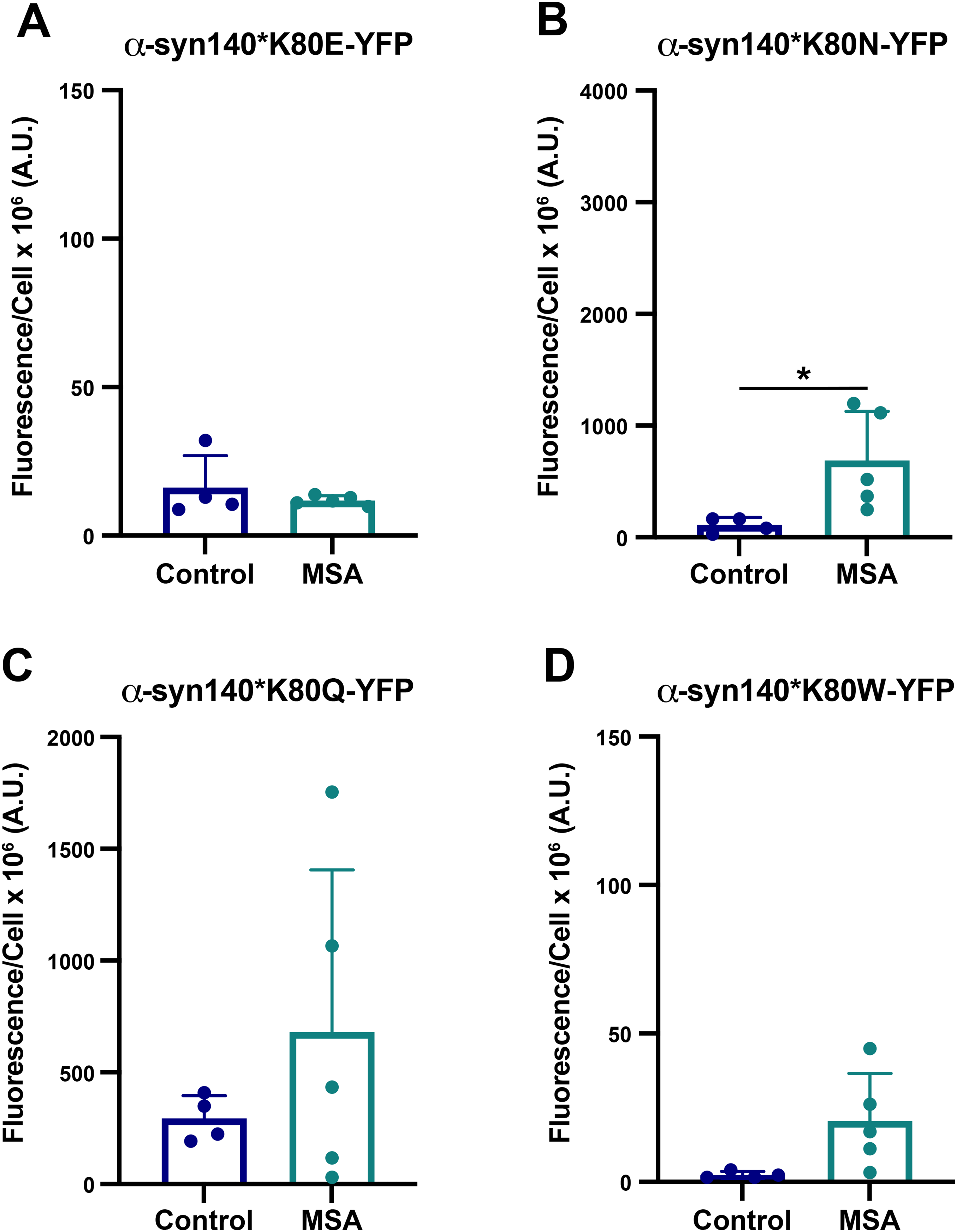
Having established the importance of the E46/K80 salt bridge in MSA prion replication, we hypothesized that swapping the residues would enable salt bridge formation, thereby re-establishing MSA replication
Discussion
Beginning with the solid-state NMR structure of α-synuclein fibrils reported by Tuttle et al., in 2016,
30
the number of reported structures deposited in Protein Data Bank has grown substantially due to advances in cryo-EM enabling the resolution of fibrillar structures made from recombinant protein26–28,31–38 or isolated from human patient samples.7,8,39 However, as our understanding of the conformations that α-synuclein can adopt has increased, the resulting biological consequences of these structural differences have not been elucidated. Here, to start addressing this pivotal question for the α-synuclein conformations in MSA, we paired an
Prior to the Goedert and Scheres labs determining the α-synuclein structures in MSA
7
(Figure 1) and LB diseases (Supplemental Figure 1A),
8
we used our cell-based assay to investigate the ability of MSA prions to replicate
We were particularly interested in evaluating the effect of the G51D and A53E mutations on MSA replication due to reports showing that patients with these mutations develop both LB and glial pathology, the latter of which resembles the GCIs in MSA patients.17,18 Anticipating these mutations would exert a similar effect on MSA replication, we were surprised to find that the A53E mutation completely inhibited α-synuclein misfolding
In addition to investigating the effect of PD mutations on MSA propagation, we used the reported MSA and PFF structures to design non-pathogenic mutations with the goal of disrupting α-synuclein misfolding into specific conformations (Table 2; Figure 3). As expected, the E61Q and V66F mutations, which were designed to interfere with the rod and twister PFF conformations (residues noted in purple in Supplemental Figure 1B and C), largely facilitated MSA replication, as did the V74P mutation, though with a blunted response. Unexpectedly, the V74I mutation blocked MSA propagation. This is particularly interesting given that our Maestro analyses suggested that both the V74I and V74P mutations would reduce the total energy of the structure. By comparison, the G36K, V37F, and V55Y mutations were all designed to interfere with the protofilament interface in the MSA structures; however, V55Y exerted the strongest inhibitory effect on MSA patient sample propagation. It should also be noted that residue V55 is located next to a small unfilled pocket in the cryo-EM structure, and it is unclear if that pocket is filled with something exogenous. As a result, we were unable to account for these potential interactions in our calculations. While analyzing our cell assay data, we noticed that in some cell lines, such as V55Y, one of the MSA patient samples appeared as an outlier. We also noticed that there was often a large range in replication ability across the patient samples, as is seen in the V66F and G36K data. To determine if this variability is due to differences in titre, or the amount of replication competent α-synuclein present in each sample, we created a heat map showing cell infection relative to the average of the control samples for each patient sample on every cell line tested (Table 4). We observed that a single patient sample is neither responsible for the lowest nor the highest infection across the cell lines, suggesting that our findings are not due to differences in titre. Instead, these data suggest the potential presence of multiple α-synuclein strains in the MSA patient samples. Given that some mutations tested, for example V66F, should have no impact on MSA replication based on the location of the residue in the reported structures, and others, such as V55Y, should inhibit α-synuclein misfolding, it is highly likely that these unreported strains have conformations that are distinct from reported structures (both residues are highlighted in purple in Figure 1A). One caveat to this interpretation is that it is currently unknown what effect
Heat map of MSA patient sample cell infection relative to control.
Our previous finding that the E46K mutation inhibits MSA replication is consistent with research on the prion protein (PrP), which has shown that a single residue change in the protein sequence can inhibit the conversion of cellular PrPC into pathogenic PrPSc. Notably, a single residue change is responsible for the species barrier creating resistance of the
An important finding from this work is that protein modelling software is not always capable of accurately predicting the effect of each mutation on MSA strain biology. For example, as was the case for residues G36, V37, H50, and G51, the location of the residue within the templating region contributes to the reliability of the modelling data. There are three important caveats to our approach that may contribute to these discrepancies. First, our methods used implicit waters in the force field calculations instead of explicit waters. Second, there are pockets in the MSA fibril structures, some of which are known to be occupied by non-protein densities, however, because it is unknown what molecules are present in these pockets, we are unable to account for their interactions with mutated residues. And third, we are using thermodynamic calculations to try to predict a kinetic process. While Maestro can be used to assess the capacity of an established structure to house a mutated residue, it is unable to predict the ability of the mutant substrate to misfold into a specific conformation.
Attempting to address this limitation, our initial approach to modelling the effect of each mutation using Maestro focused on mimicking the experimental design of our
Total energy change after mutagenesis of 3 versus 6 protein layers in MSA structure 6XYQ.
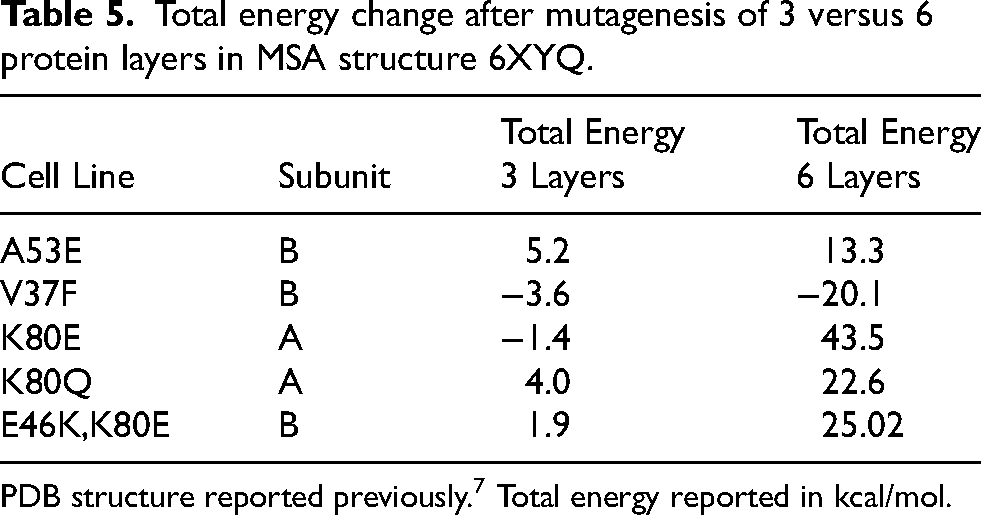

PDB structure reported previously. 7 Total energy reported in kcal/mol.
A final critical limitation to our
The remarkable advances made in structural biology methodology, particularly for prionogenic proteins, have had a tremendous impact on our understanding of protein misfolding and strain biology. The availability of these structures now serves as an opportunity for hypothesis development focused on elucidating the structure-function relationship underlying the ability of each prion strain to exert unique biological consequences in disease. However, the nature of structural techniques limits our understanding of fibril formation to a single snapshot from the brain of a deceased patient, failing to capture the dynamic processes that lead to the final fibrillar product. In this study, we paired structural analysis with
Supplemental Material
sj-docx-1-pkn-10.1177_JPD-240296 - Supplemental material for Structurally targeted mutagenesis identifies key residues supporting α-synuclein misfolding in multiple system atrophy
Supplemental material, sj-docx-1-pkn-10.1177_JPD-240296 for Structurally targeted mutagenesis identifies key residues supporting α-synuclein misfolding in multiple system atrophy by Patricia M Reis, Sara AM Holec, Chimere Ezeiruaku and Matthew P Frost, Christine K Brown, Samantha L Liu, Steven H Olson, Amanda L. Woerman in Journal of Parkinson's Disease
Supplemental Material
sj-docx-2-pkn-10.1177_JPD-240296 - Supplemental material for Structurally targeted mutagenesis identifies key residues supporting α-synuclein misfolding in multiple system atrophy
Supplemental material, sj-docx-2-pkn-10.1177_JPD-240296 for Structurally targeted mutagenesis identifies key residues supporting α-synuclein misfolding in multiple system atrophy by Patricia M Reis, Sara AM Holec, Chimere Ezeiruaku and Matthew P Frost, Christine K Brown, Samantha L Liu, Steven H Olson, Amanda L. Woerman in Journal of Parkinson's Disease
Footnotes
Acknowledgments
ORCID iDs
Funding
The authors disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by grants from the NIH (R01NS121294 and accompanying supplement and R21NS127002) to A.L.W. Control tissue samples were provided by the NIH NeuroBioBank and by the Massachusetts Alzheimer's Disease Research Center (MADRC), which is supported by 1P30AG062421. Multiple system atrophy patient samples were provided by the MADRC and the Sydney Brain Bank at Neuroscience Research Australia, which is supported by The University of New South Wales and Neuroscience Research Australia.
Declaration of conflicting interests
The authors declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: A.L.W. is an Editorial Board Member of this journal but was not involved in the peer-review process of this article nor had access to any information regarding its peer-review. S.H.O. and A.L.W. are the co-founders of Allagus Therapeutics, which did not contribute financial or any other support to these studies. S.H.O. and A.L.W. are inventors on U.S. Patent Application PCT/US2023/072173, which includes data reported here.
Data availability
The data supporting the findings of this study are available within the article and/or its supplemental material.
Supplemental material
Supplemental material for this article is available online.
