Abstract
Background:
Tobacco smoking and alcohol intake have been identified in observational studies as potentially protective factors against developing Parkinson’s disease (PD); the impact of body mass index (BMI) on PD risk is debated. Whether such epidemiological associations are causal remains unclear. Mendelian randomsation (MR) uses genetic variants to explore the effects of exposures on outcomes; potentially reducing bias from residual confounding and reverse causation.
Objective:
Using MR, we examined relationships between PD risk and three unhealthy behaviours: tobacco smoking, alcohol intake, and higher BMI.
Methods:
19,924 PD cases and 2,413,087 controls were included in the analysis. We performed genome-wide association studies to identify single nucleotide polymorphisms associated with tobacco smoking, alcohol intake, and BMI. MR analysis of the relationship between each exposure and PD was undertaken using a split-sample design.
Results:
Ever-smoking reduced the risk of PD (OR 0.955; 95%confidence interval [CI] 0.921–0.991;
Conclusion:
Using split-sample MR in over 2.4 million participants, we observed a protective effect of smoking on risk of PD. In contrast to observational data, alcohol consumption appeared to increase the risk of PD. Higher BMI had a protective effect on PD, but the effect was small.
INTRODUCTION
Observational studies have identified putative risk factors for Parkinson’s disease (PD) but their effect on aetio-pathogenesis, hence their value as targets for neuroprotection, remains unclear. The impact of three phenotypes in particular—tobacco smoking, alcohol intake, and high body mass index (BMI)—on risk of PD warrants clarification given their prevalence in the general population and their potential for modification should they modulate disease risk.
Observational data suggest that ever-smokers are 40%less likely to develop PD than never-smokers, with a dose-response relationship being observed with pack-years smoked [1]. Similarly, high alcohol intake was associated with a 22%lower risk of PD in a large-scale meta-analysis [2], although this may be driven in part by small-study bias [3]. A recent case-control study supported the negative observational associations with smoking and alcohol [4].
The role of BMI in risk of PD is unclear. Low BMI is apparent in patients with PD in case-control studies, however a meta-analysis of 10 prospective studies found no association between premorbid BMI and PD risk [5].
The extent to which such epidemiological associations are causal or driven by confounding factors and/or reverse causation remains unclear. Risk-averse individuals may be less likely to engage in unhealthy behaviours and more likely to develop PD [6]. Reverse causation is plausible given the long prodromal phase of PD [7]. Indeed, ease of smoking cessation and weight loss may be prodromal features of PD [8, 9].
Using Mendelian randomisation (MR), we sought to explore the nature of associations between PD risk and smoking, alcohol intake, and higher BMI. In MR, genetic variants associated with the exposure are used as instrumental variables to estimate the effect of the exposure on the outcome. The random allocation of genetic variants to offspring means that alleles are generally unrelated to confounding factors. Moreover, the germline genotype cannot be modified by the disease process, minimising bias from reverse causality. MR helps address limitations of randomized controlled trials: ethical (when the intervention is harmful, e.g., smoking) and pragmatic (when adherence cannot be guaranteed, e.g., alcohol intake, BMI modulation) [10].
METHODS
Participants
Participants were customers of 23andMe, Inc., a personal genomics company. Study protocols were approved by an external AAHRPP-accredited institutional review board and conducted in accordance with the Declaration of Helsinki principles. Participants gave informed consent to participate. Data was collected between 10/11/2007 and 1/1/2018.
Defining phenotypic traits
Parkinson’s disease status
PD cases were drawn from the 23andMe participants who self-reported a diagnosis of PD. For a fuller description of recruitment see Do et al. [11]. If multiple PD questions were completed, participants were scored as cases if they reported being a case in any survey. Despite questionnaire data providing less diagnostic accuracy than clinically ascertained data, satisfactory level of agreement between patient-reported PD diagnosis and neurologist assessment has been demonstrated in a small validation study [12]. Participants reporting a change in their diagnosis or uncertainty relating to their diagnosis were excluded. Controls were drawn from the 23andMe participants who self-reported never having been diagnosed with PD. We also removed cases and controls who self-reported having ever been diagnosed with: 1) atypical parkinsonism (e.g., dementia with Lewy bodies, progressive supranuclear palsy, multiple system atrophy, corticobasal degeneration) or a non-parkinsonian tremor disorder; or 2) stroke, deep vein thrombosis, or pulmonary embolism (to reduce the probability of including individuals with vascular parkinsonism). 19,924 PD cases and 2,413,087 controls were included in the analysis. For a description of how 23andMe research participants with PD compare phenotypically to those without PD, see Heilbron et al. (2019) [4].
Unhealthy exposures of interest
Participants self-reported tobacco smoking habits with response categories “ever-smoker (> 100 cigarettes smoked in lifetime)”, or “never-smoker (< 100 cigarettes smoked in lifetime)”. Our tobacco use phenotype followed the established standard definition used by the US Centers for Disease Control and Prevention (CDC) in their National Health Interview Survey [13]. Participants self-reported the number of alcohol measures consumed per day over the past two weeks, where 1 measure corresponds to 12 oz. of beer, 5 oz. of wine, or 1.5 oz of spirits. Response options were grouped as: 0, 0–1, 1, 2, 3, 4, 5 or more measures. Participants self-reported their mass (in kilograms) and height (in metres) squared, from which BMI was calculated. The majority of the exposure data was collected when a 23andMe research participant completed their first survey. If multiple tobacco use questions were completed, participants were scored as ever-smokers if they reported being an ever-smoker in any survey. If multiple alcohol use, mass, height, or PD questions were completed, the first non-missing value was taken from an ordered list of surveys. Responses were highly concordant across surveys (e.g.,
Genome-wide association studies
We performed new GWAS for each exposure using unrelated individuals. We selected unrelated individuals using a segmental identity-by-descent estimation algorithm [14]. Individuals were defined as related if they shared 700 cM identity-by-descent, including regions where the two individuals share one or both genomic segments identical-by-descent. This level of relatedness (∼20%of the genome) corresponds approximately to the minimal expected sharing between first cousins in an outbred population. Ancestry composition was performed as previously reported [15]. Inclusion was restricted to individuals of predominantly European ancestry to minimise confounding by ancestry.
DNA extraction and genotyping were performed on saliva samples by National Genetics Institute. Samples were genotyped on one of five Illumina-based genotyping platforms. The v1 and v2 platforms were variants of the Illumina HumanHap550 + BeadChip, including about 25,000 custom SNPs selected by 23andMe, with a total of about 560,000 SNPs. The v3 platform was based on the Illumina OmniExpress + BeadChip, with custom content to improve the overlap with our v2 array, with a total of about 950,000 SNPs. The v4 platform was a fully customized array, including a lower redundancy subset of v2 and v3 SNPs with additional coverage of lower-frequency coding variation, and about 570,000 SNPs. The v5 platform is an Illumina Infinium Global Screening Array (∼640,000 SNPs) supplemented with ∼50,000 SNPs of custom content. Samples had minimum call rates of 98.5%.
We phased participant data using either an internally-developed tool, Finch (V1-V4 genotyping arrays) or Eagle2 (V5 genotyping array) [16]. Finch implements the Beagle haplotype graph-based phasing algorithm, modified to separate the haplotype graph construction and phasing steps [17]. It extends the Beagle model to accommodate genotyping error and recombination, to handle cases where there are no consistent paths through the haplotype graph for the individual being phased. We constructed haplotype graphs for European and non-European samples on each 23andMe genotyping platform from a representative sample of genotyped individuals, and then performed out-of-sample phasing of all genotyped individuals against the appropriate graph. For the X-chromosome, we built separate haplotype graphs for the non-pseudoautosomal region and each pseudoautosomal region, and these regions were phased separately.
Imputation
Imputation panels created by combining multiple smaller panels have been shown to give better imputation performance than the individual constituent panels alone [18]. To that end, we combined the May 2015 release of the 1000 Genomes Phase 3 haplotypes with the UK10K imputation reference panel to create a single unified imputation reference panel [19, 20]. Multiallelic sites with N alternate alleles were split into N separate biallelic sites. We then removed any site whose minor allele appeared in only one sample. For each chromosome, we used Minimac3 to impute the reference panels against each other, reporting the best-guess genotype at each site [21]. This gave us calls for all samples over a single unified set of variants. We then joined these together to get, for each chromosome, a single VCF with phased calls at every site for 6,285 samples.
In preparation for imputation, we split each chromosome of the reference panel into chunks of no more than 300,000 variants, with overlaps of 10,000 variants on each side. We used a single batch of 10,000 individuals to estimate Minimac3 imputation model parameters for each chunk [21]. We imputed phased participant data against the chunked merged reference panel using Minimac3, treating males as homozygous pseudo-diploids for the non-pseudoautosomal region. Throughout, we treated structural variants and small indels the same as SNPs.
Association tests
We computed association test results by regression assuming additive allelic effects (logistic regression for case-control traits [PD, tobacco use], linear regression for quantitative traits [alcohol consumption, BMI]). We included covariates for age, sex, the top five genetic principal components to account for residual population structure, and indicators for genotype platforms to account for genotype batch effects. The association test
Principal component analysis
We performed the genetic principal components analysis using ∼65,000 high-quality genotyped variants present in all five genotyping platforms and a random sample of one million research participants with predominantly European ancestry. Principal component scores for participants not included in the analysis were obtained by projection, combining the eigenvectors of the analysis and the SNP weights.
Quality control of genotyped GWAS results
We excluded SNPs that: 1) had a call rate < 90%, 2) had a Hardy-Weinberg
Quality control of imputed GWAS results
We excluded SNPs with imputed r2 < 0.3, as well as SNPs that had strong evidence of a platform batch effect. For each SNP we identified the largest subset of the data passing other quality control criteria based on their original genotyping platform –either v2 + v3 + v4 + v5, v4 + v5, v4, or v5 only –and computed association test results for the largest passing set. The batch effect test is an F test from an ANOVA of the SNP dosages against a factor representing the V4 or V5 platform; we excluded results with
Additional quality control of GWAS results
Across both genotyped and imputed GWAS results, we excluded SNPs that had sample size of less than 20%of the total GWAS sample size. We also removed SNPs that did not converge during logistic regression, as identified by abs (effect) > 10 or stderr > 10 on the log-odds scale. We removed SNPs with MAF < 0.1%from linear regressions because these SNPs are sensitive to violations of the regression assumption of normally distributed residuals. If SNPs were both genotyped and imputed, and they passed QC for both, we used results from the imputed analysis.
After quality control, we had analysed 904,040 genotyped SNPs and 25,208,208 imputed SNPs.
Instrument construction
SNPs associated with each of the exposures at the genome-wide significance level (
Split-sample MR analysis
We performed MR analysis of the relationship between the exposures of interest and PD using a split-sample design, in accordance with published methods [23, 24]. Individuals from the 23andMe cohort were randomly allocated into two evenly sized groups (Table 1). Demographic differences between cases and controls across cohorts are shown in Table 2. The instrument-exposure association was measured in the first group (cohort 1), and the instrument-outcome association was measured in the second group (cohort 2), and MR analyses undertaken. We then repeated the MR analyses but using cohort 1 for the outcome and cohort 2 for the exposure. For each MR method, this resulted in two independent MR estimates, which were combined using an inverse variance weighted (IVW) fixed-effects meta-analysis.
Participant numbers in cohort 1 and cohort 2
Demographics of PD cases and controls in cohort 1 and cohort 2
PD, Parkinson’s disease; SD, standard deviation; N, number of individuals in a cohort.
The effect of an exposure on PD was calculated for each SNP using the Wald ratio method [25]. In the IVW analysis we performed a linear regression constrained through the origin of the variant-exposure and the variant-outcome associations for each instrument, weighted by their inverse variance.
We used four methods to assess the impact of bias on IVW estimates [22]. Heterogeneity in Wald ratio estimates, which can indicate bias due to horizontal pleiotropy, was assessed using the Cochran’s Q test and I2 index [26]. dividual variant contributions to Cochran’s Q was calculated, and variants were excluded in the heterogeneity filtering analysis if their contribution surpassed the Bonferroni-corrected 99.8th percentile of the χ2 (1df) distribution. We performed MR-Egger analysis to assess the magnitude of bias occurring due to horizontal pleiotropy, the weighted median method to assess for invalid instruments, and generalised summary data-based MR (GSMR) to filter out pleiotropic SNPs (Supplementary Table 1).
To minimize misallocation of variants to the exposure rather than outcome group, we applied Steiger filtering to all analyses to remove genetic variants that had a stronger correlation with the outcome than with the exposure [27].
Replication in the International Parkinson’s Disease Genomics Consortium (IPDGC) dataset
Summary statistics from the largest published PD GWAS meta-analysis were used as the outcome data for replication after excluding data from 23andMe (the “IPDGC dataset”) [28]. he outcome summary statistics used for this analysis included 15,056 cases, 18,618 proxy cases, and 449,056 controls, and there were 17,410,431 genotyped and imputed SNPs tested for association with PD. Recruitment and genotyping quality control are described in the original report [28]. Only exposure SNPs from cohort 1 were used in the replication analysis, and we excluded Only exposure SNPs from cohort 1 were used in the replication analysis, and we excluded SNPs on the X-chromosome and palindromic SNPs.
Sensitivity analyses
If tobacco use has a protective effect on PD, we would expect genetic variants that increase the tobacco use to decrease PD risk in tobacco users, but to have no effect on PD risk in non-users. If these variants are protective for PD in separate analyses of both ever-smokers and never-smokers, however, this suggests that the protective effect is mediated by a pleiotropic pathway unrelated to tobacco use. Given that ever/never-smoker status has a positive genetic correlation with “number of cigarettes smoked per day” (
Statistical analysis
Analyses were performed using R statistical software 3.3.2 (2016-10-31).
Data availability statement
Summary data is available via application at 23andMe https://research.23andme.com/dataset-access/.
RESULTS
Tobacco use
385 SNPs were associated with self-reported smoking status (ever- versus never-smoker) in cohort 1 (422 SNPs in cohort 2, Supplementary Tables 2 and 3). These instruments explained an average of 0.80%of the variance in tobacco smoking liability in the out-of-sample cohort (cohort 1 SNPs = 0.80%, cohort 2 SNPs = 0.79%). The F statistic (
In the IVW analysis, ever-smoking had the effect of reducing PD risk with an OR 0.955 (95%CI 0.921–0.991,
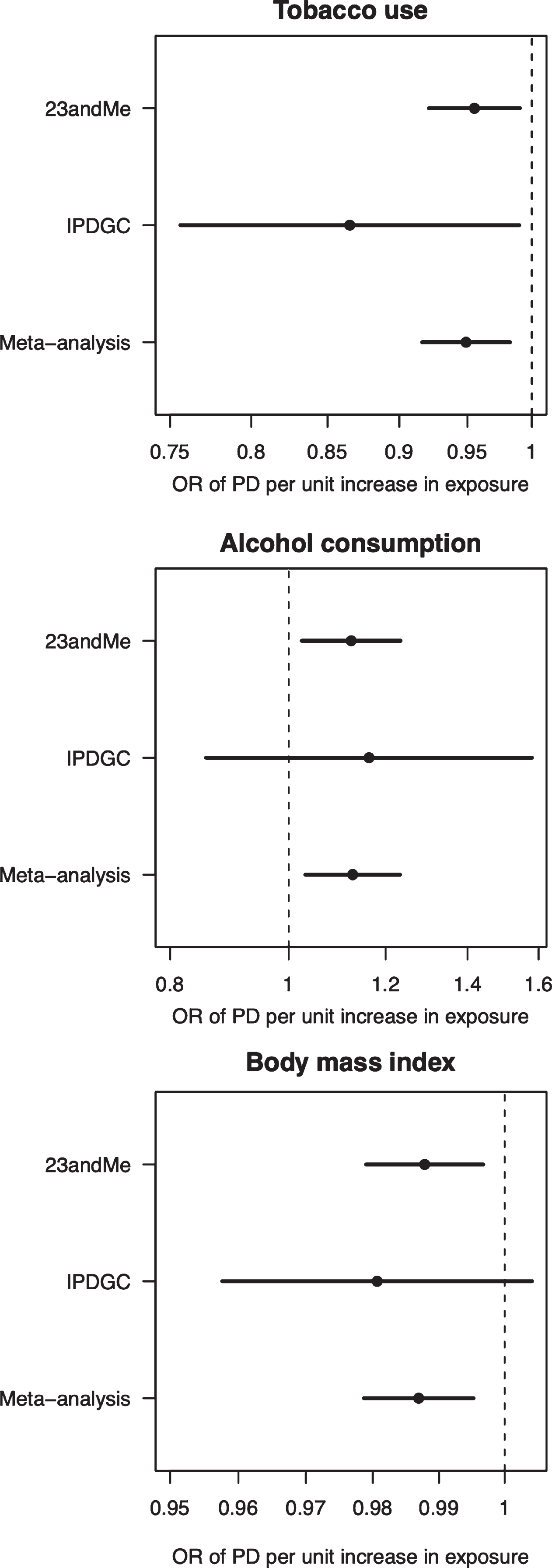
Forest plot of Mendelian randomisation causal association estimates between risk of Parkinson’s disease and unhealthy behaviours derived from meta-analysis of the 23andMe and IPDGC datasets. The pooled odds ratio (OR), derived from meta-analysis of the inverse variance weighted estimates, and 95%confidence intervals are shown. For smoking, the unit of exposure is never versus ever smoking. For alcohol, the unit of exposure is 1-group difference in daily alcohol intake. For BMI, the unit of exposure is 1 kg/m2. OR, odds ratio; PD, Parkinson’s disease.
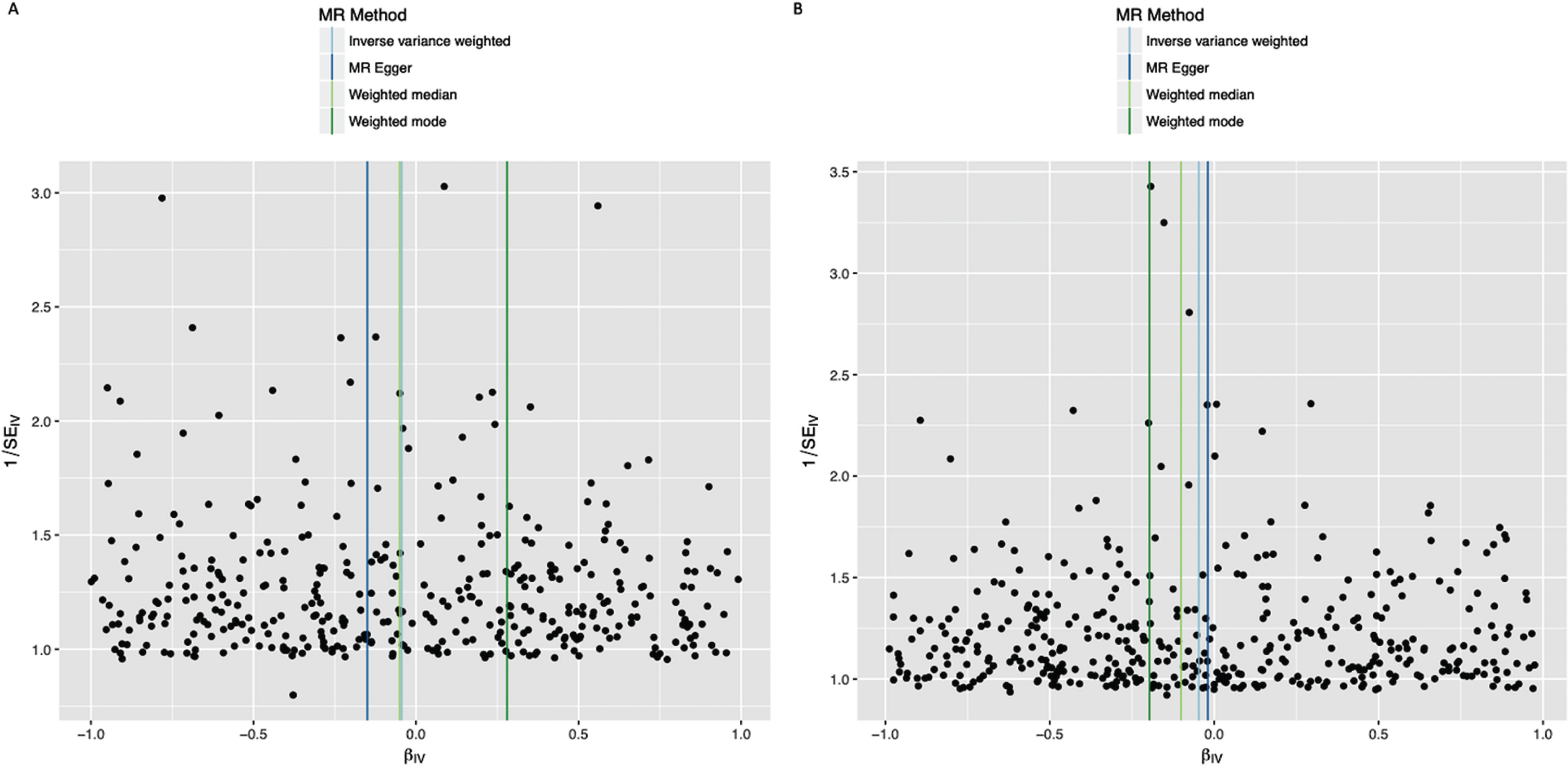
Funnel plots of individual variant effects for the smoking instrument (from cohort 1 (A) and cohort 2 (B)) plotted against the inverse of their standard error.
As a sensitivity analysis, we re-ran MR in the 23andMe dataset separately for ever-smokers and never-smokers. We found a similar protective effect, although less precisely estimated, in ever-smokers (OR = 0.966, 95%CI 0.885–1.055) and little evidence of a protective effect in never-smokers (OR = 1.030, 95%CI 0.966–1.097). We found similarly imprecise estimates when performing MR using a SNP in
Alcohol intake
129 SNPs were associated with self-reported alcohol in cohort 1 (124 SNPs in cohort 2, Supplementary Tables 7 and 8). These instruments explained an average of 0.32%of the variance in alcohol intake liability in the out-of-sample cohort (cohort 1 SNPs = 0.33%, cohort 2 SNPs = 0.31%). The F statistic (
In the IVW analysis, alcohol intake had the effect of increasing PD risk with an OR 1.125 for a 1-group increase in daily alcohol intake (95%CI 1.025–1.235,
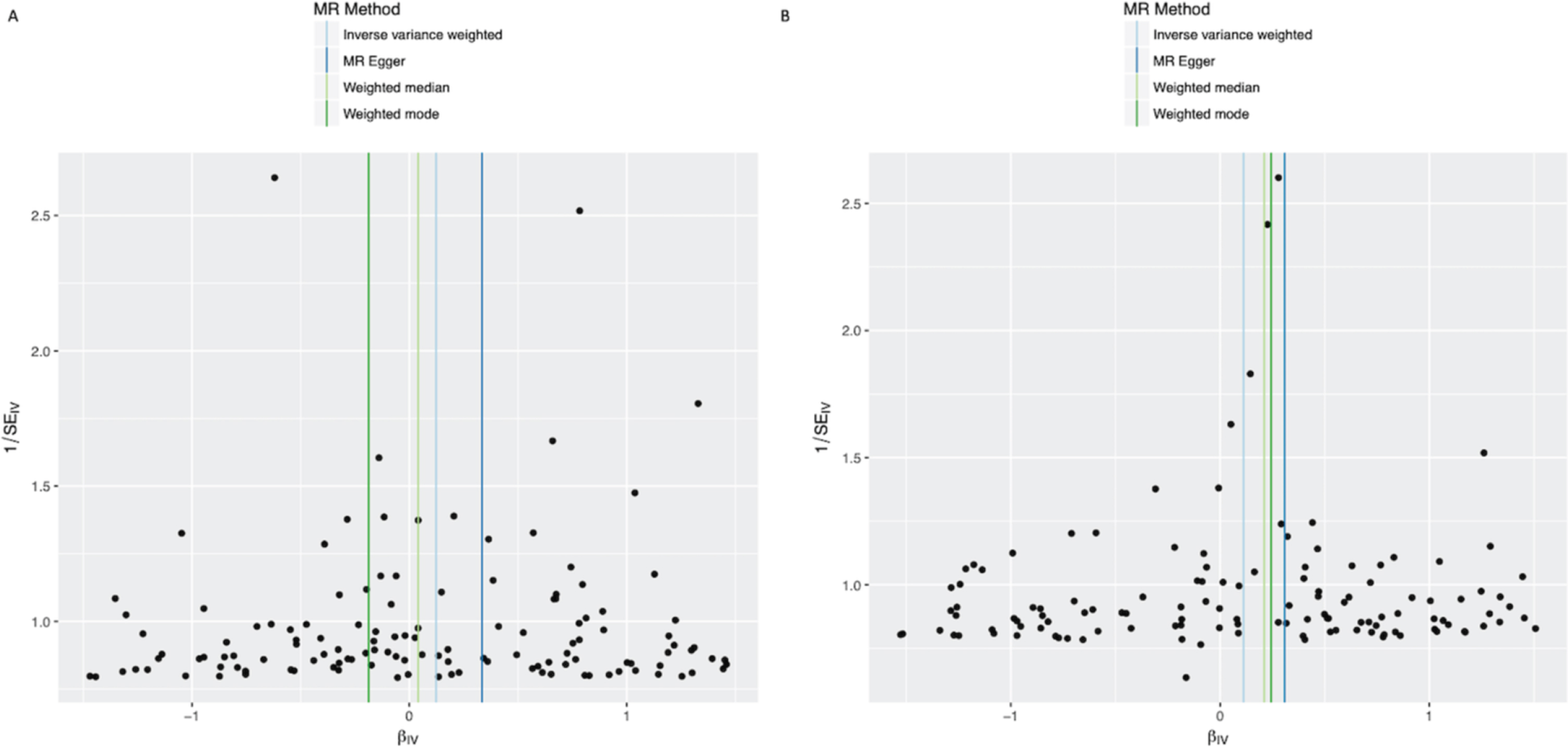
Funnel plots of individual variant effects for the alcohol instrument (from cohort 1 (A) and cohort 2 (B)) plotted against the inverse of their standard error.
BMI
729 SNPs were associated with self-reported BMI in cohort 1 (693 SNPs in cohort 2, Supplementary Tables 9 and 10). These instruments explained an average of 4.56%of the variance in BMI liability in the out-of-sample cohort (cohort 1 SNPs = 4.63%, cohort 2 SNPs = 4.50%). The F statistic (
In the IVW analysis, a genetically-estimated 1 kg/m2 higher BMI had the effect of reducing PD risk with an OR 0.988 (95%CI 0.979–0.997,
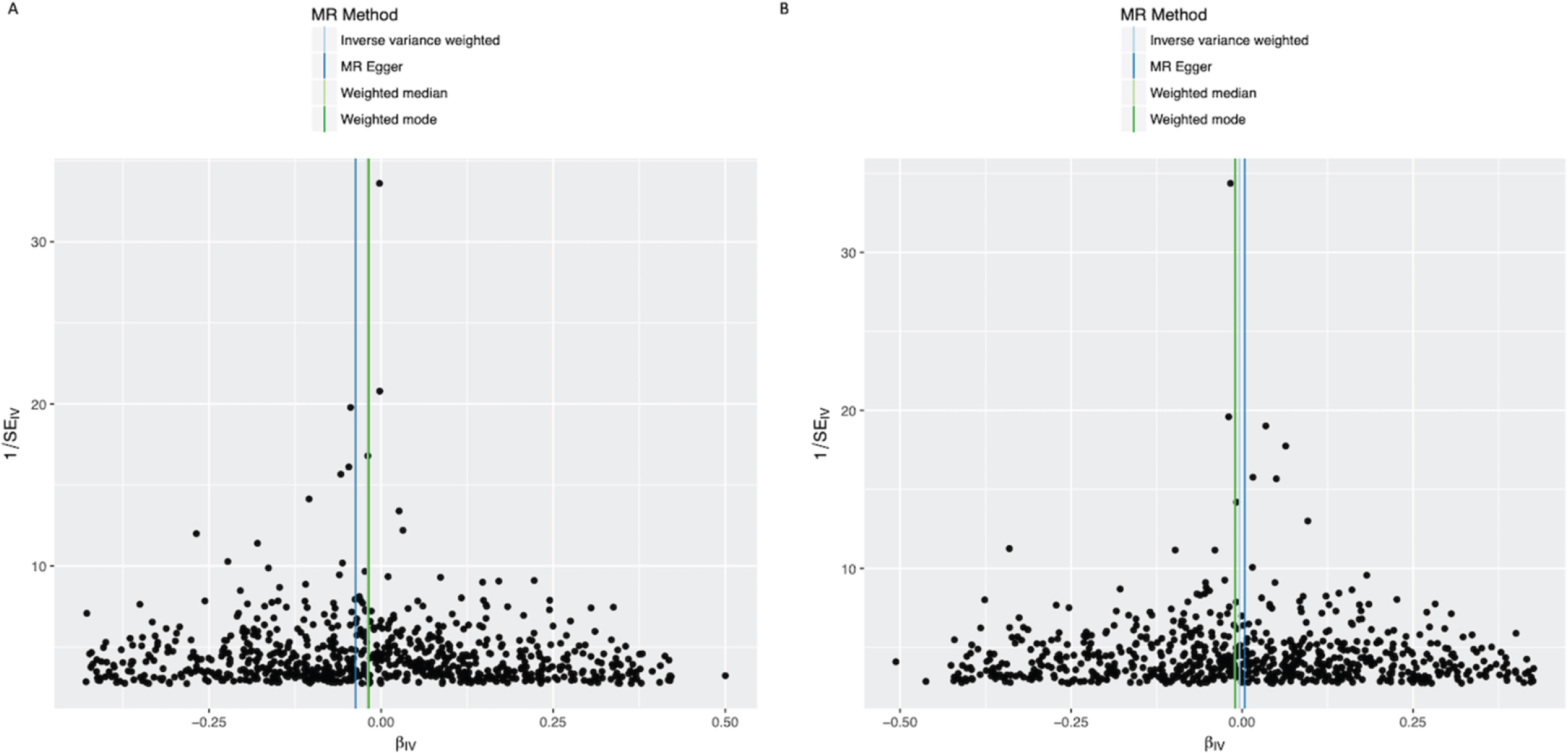
Funnel plots of individual variant effects for the BMI instrument (from cohort 1 (A) and cohort 2 (B)) plotted against the inverse of their standard error.
DISCUSSION
Using split-sample MR in a cohort of > 2.4 million participants, ever-smoking reduced the risk of PD by 5%compared to never-smoking. Sensitivity analyses suggested that the effect was unlikely to be driven by horizontal pleiotropy, outliers, or invalid instruments. The findings provide cautious support for a protective effect of smoking in the aetio-pathogenesis of PD.
These findings are concordant with observational studies demonstrating inverse associations between smoking and PD risk [1]. However, observational studies cannot definitively rule out reverse causality: in a study of 220,000 individuals, the protective effect of smoking on PD risk no longer held true among smokers who had quit > 20 years before recruitment, meaning that preclinical dopaminergic changes facilitating smoking cessation could not be excluded [31]. Similarly, a cohort study reported that parental smoking during childhood reduced future PD risk, however a transgenerational exposure, such as a toxin influencing parental smoking behaviour and PD risk in children, could not be excluded [32]. By circumventing the use of proxy measures to overcome unmeasured/residual confounding and reverse causality, the findings from MR analyses more robustly support a protective effect.
Our results are similar to other MR studies (Table 3) exploring the relationship between PD and tobacco use. Grover et al. found a significant protective effect (OR = 0.71 per log-odds of ever-smoking; 95%CI 0.57–0.90), while Nalls et al. found some evidence for a protective effect (OR = 0.94 per log-odds of ever-smoking; 95%CI 0.88–1.00,
Exposure and outcome sample sizes and reported inverse variance weighted Mendelian randomisation estimates (odds ratio and 95%confidence interval) for the effect of smoking on risk of PD in the present study (23andMe discovery cohort and IPDGC replication cohort), and the two other Mendelian randomisation studies exploring the association between smoking and PD
PD, Parkinson’s disease; IPDGC, International Parkinson’s disease Genomics Consortium.
Early interventional studies demonstrated beneficial effects of nicotine on motor and cognitive deficits in PD [34–41]. However, placebo-controlled studies largely failed to replicate the protective association between smoking and PD [42–44]. This may be because trials assessed transient changes in PD symptoms post-nicotine in small cohorts. Given the heterogeneity in PD progression, demonstrating disease-modification would necessitate large sample sizes and long follow-up. Trials may also have failed because they examined disease-modifying effects of nicotine on established PD. However, the mechanisms underlying PD risk may be different to those driving disease progression. Our study suggests that smoking influences the former, highlighting the need for prevention trials of candidate disease-modifying therapies in pre-manifest PD.
Mechanisms underlying the effect of smoking on PD risk remain speculative. A disease-modifying effect of nicotine is biologically plausible given evidence that it mitigates MPTP- and 6-hydroxydopamine-induced motor dysfunction in animal models of PD [45]. The apparent protective effect of smoking on risk of PD was not observed when stratifying the MR analysis to a SNP in
In this study we observed evidence for an increase in risk of PD with higher alcohol intake. Although a recent meta-analysis found that alcohol consumption was associated with a lower risk of PD in case-control studies (OR never versus heavy/moderate drinking: 0.74; 95%CI 0.64–0.85), in meta-analysed cohort studies the difference was non-significant [46]. In this study we similarly observed a corresponding discrepancy between the effect of alcohol intake on risk of PD in the observational analysis (OR 0.4) and MR analysis (OR 1.1). The inverse association observed in case-control studies may be driven by reverse causation, whereby lower alcohol consumption in cases reflects a low dopaminergic state or survivor bias. Our finding is consistent with the neurotoxic effect of excess alcohol in human neuropathological studies, and with studies in rodents showing that alcohol increases oxidative damage in nigral cells [47, 48]. Our results suggest that alcohol consumption is unlikely to help prevent PD.
We have previously observed a protective effect of higher BMI on risk of PD in an MR study [49]. Recently, a hypothesis-free MR approach exploring causal associations between exposures and PD observed that most of the top hits related to a protective effect of increased adiposity [50]. The results of our study support these previous observations.
The key strength of MR analyses is the use of genetic variants to explore the nature of associations between exposures and outcomes, thus limiting bias from residual confounding or reverse causation present in observational studies. However, there are general limitations inherent to all MR studies. Firstly, MR analysis assumes linearity, precluding us from identifying non-linear exposure-outcome associations. Secondly, it has been argued that survival bias may distort MR results. We mitigated this in part by employing two-sample design, which has been shown to minimise the impact of survival bias on effect estimates and previous work has shown that the protective effect of higher BMI on risk of PD was not clearly explained by survival bias [49, 51].
Additional specific limitations of our study include that we did not explore whether effects differed between sub-groups (for example between sexes, as previously suggested for the PD-smoking association) [1]. Our phenotypes were constructed using self-reported data derived from online surveys and may therefore suffer from recall bias and desirability bias. Finally, the individuals studied were not a random sample of the general population (Table 2 for cohort demographics), potentially leading to selection bias. For example, non-randomly-sampled participants can lead to biased SNP effect estimates and alter MR results [52]. Exposures and outcomes (excluding the IPDGC data) were self-reported and therefore our SNP effects may be underestimated. Underestimated SNP effects on the exposure may lead to inflated exposure-outcome IVW estimates away from the null. Nonetheless our results are consistent with a previous MR study demonstrating a protective effect of higher BMI on risk of PD with a larger effect size [49]. Independent replication of our findings in other datasets would provide further evidence that our findings are not an artefact of self-selection in the 23andMe cohort, including showing that our MR instrument for ever-never smoking is predictive of amount smoked.
In conclusion, we provide evidence to support a protective effect of smoking and high BMI on PD risk. Conversely, we observed a detrimental effect of alcohol on PD risk. Although a better understanding of the underlying mechanisms, as well as development of safe delivery methods is necessary, such findings help guide the prioritisation of candidate neuroprotective approaches for randomized controlled trials in participants at risk of developing PD.
Footnotes
ACKNOWLEDGMENTS
The work was financially supported by The Michael J. Fox Foundation for Parkinson’s Research (grant MJFF12737), National Institute on Aging (NIA) part of the National Institutes of Health (NIH) and by 23andMe, Inc. This work was supported in part by the Intramural Research Programs of the National Institute on Aging (NIA). The Preventive Neurology Unit at the Wolfson Institute of Preventive Medicine is funded by the Barts Charity. MPJ acknowledges the Isaac Schapera Trust for funding support.
CONFLICT OF INTEREST
The authors have no potential conflicts of interest to report.
