Abstract
Background:
Myogenesis is a dynamic process involving temporal changes in the expression of many genes. Lack of dystrophin protein such as in Duchenne muscular dystrophy might alter the natural course of gene expression dynamics during myogenesis.
Objective:
To gain insight into the dynamic temporal changes in protein expression during differentiation of normal and dystrophin deficient myoblasts to myotubes.
Method:
A super SILAC spike-in strategy in combination and LC-MS/MS was used for temporal proteome profiling of normal and dystrophin deficient myoblasts during differentiation. The acquired data was analyzed using Proteome Discoverer 2.2. and data clustering using R to define significant temporal changes in protein expression.
Results:
sFour major temporal protein clusters that showed sequential dynamic expression profiles during myogenesis of normal myoblasts were identified. Clusters 1 and 2, consisting mainly of proteins involved mRNA splicing and processing expression, were elevated at days 0 and 0.5 of differentiation then gradually decreased by day 7 of differentiation, then remained lower thereafter. Cluster 3 consisted of proteins involved contractile muscle and actomyosin organization. They increased in their expression reaching maximum at day 7 of differentiation then stabilized thereafter. Cluster 4 consisting of proteins involved in skeletal muscle development glucogenesis and extracellular remodeling had a lower expression during myoblast stage then gradually increased in their expression to reach a maximum at days 11–15 of differentiation. Lack of dystrophin expression in DMD muscle myoblast caused major alteration in temporal expression of proteins involved in cell adhesion, cytoskeleton, and organelle organization as well as the ubiquitination machinery.
Conclusion:
Time series proteome profiling using super SILAC strategy is a powerful method to assess temporal changes in protein expression during myogenesis and to define the downstream consequences of lack of dystrophin on these temporal protein expressions. Key alterations were identified in dystrophin deficient myoblast differentiation compared to normal myoblasts. These alterations could be an attractive therapeutic target.
Keywords
INTRODUCTION
Skeletal muscle constitutes a large portion of the human body and plays a major role in supporting body posture and movement coordination. Its dynamic malleability and plasticity make it resilient to the daily physical demands and even to minor injuries [1, 2]. Muscle fibers are continuously degenerating and regenerating in the body during developmental stage and even after reaching development. The degeneration/regeneration process is driven by satellite cells through myogenesis. Myogenesis is initiated after the arrest in satellite cell proliferation cycle and switching from single myoblast to fusion of several mono-nucleated myoblasts into multi-nucleated myotube that form a muscle fiber. The entire process involves complex and sequential temporal changes in the expression of certain genes including the switch genes for initiating myogenesis (e.g. expression of myogenic regulatory transcription factors dictating the myoblast to switch from proliferation behavior to differentiation [3, 4]) followed by the expression of key structural and regulatory proteins necessary for muscle fiber organization and function such as sarcomere, sarcoplasmic, contractile and extracellular matrix proteins. Immortalized C2C12 muscle cell line has been extensively used to elucidate molecular mechanisms driving myogenesis and myotube maturation [5–7] Valuable information on changes in protein expression were thus identified using a proteomic profiling approach during differentiation of this immortalized C2C12 cell line [5]. A recent study used triple stable isotope labeling by amino acid in cell culture (SILAC) strategy to profile the proteome of primary human myoblast during differentiation and started shedding light on the complex dynamic changes in protein expression during myogenesis [8]. The authors identified 243 regulated proteins that were clustered into proteins involved in cell differentiation, RNA metabolism, cell adhesion and cytoskeleton organization. Understanding the molecular dynamics involved in myogenesis and muscle fiber formation might help determine therapeutic targets for certain muscle diseases.
A muscle fiber is exquisitely organized multinucleated myocyte that interact with the extracellular matrix via the dystrophin associated protein complex and confer proper function of skeletal muscle [9, 10]. Dystrophin localizes in sarcolemma and is necessary to maintain the structure and the interaction of the sarcolemma with the extracellular matrix during muscle fibers contraction and relaxation. Lack of dystrophin expression might alter the normal temporal changes in protein expression during myogenesis leading to disorganized muscle fiber and disorganized extracellular environment. To better understand the role of dystrophin in myogenesis and myotube maturation we implemented a robust time series proteome profiling experiment using super SILAC strategy [11] and compared temporal changes of protein expression in normal versus dystrophin deficient human muscle cells.
MATERIAL AND METHODS
Human myoblast cell culture and differentiation
Immortalized normal and DMD human myoblast cells were a gift from Dr. Vincent Mouly (Institute of myology, Paris France) [12]. Normal healthy myoblasts were originally isolated from muscle biopsies obtained from semitendinosus muscle of a 25-year-old healthy donor with no genetic mutation in the dystrophin gene while dystrophin deficient myoblasts were isolated from the quadriceps of a 2-year-old DMD patient with a confirmed deletion of exon 48–50 in the dystrophin gene. Isolated myoblasts were immortalized by co-transfection with hTERT and CDK-4 cDNA using retroviral vectors [12]. The overall workflow of muscle cell culture and SILAC strategy are depicted in Fig. 1. First, a batch of SILAC labelled normal myoblast was generated by growing the cells in porcine gelatin coated T-75 cm2 flasks for 5 passages (∼7 cell doublings). The complete growth media was made of Lys- and Arg-free high glucose DMEM supplemented with PromoCell Growth Supplement (PromoCell, Heidelberg, Germany), 1% Penicillin-Streptomycin (P/S), heavy isotopes of Lysine and Arginine –13C6-Arg and 13C6,15N2-Lys (Thermo Fisher Scientific, Waltham, MA) and final fetal bovine serum (FBS) concentration of 20%. Incubation and expansion of myoblasts was performed at 37°C in a humid atmosphere containing 5% CO2 and trypsinized using Trypsin-EDTA (MilliporeTM) to at 70% confluency for further passaging. Around 1×106 myoblast cells were seeded in each T-75 cm2 flask and at each passage for proliferation and differentiation. Twelve flasks (at passage 5) were harvested at the myoblast stage after a 24-hour incubation in serum-free growth media. The remaining 12 flasks of myoblasts were differentiated in SILAC DMEM supplemented with PromoCell Differentiation supplement (PromoCell, Heidelberg, Germany), 2% FBS 1% P/S and heavy isotopes of Lysine and Arginine. The resulting multinucleated myotubes were harvested at day 14 after a 24-hour incubation in serum-free differentiation media and processed for total protein extraction as described below.
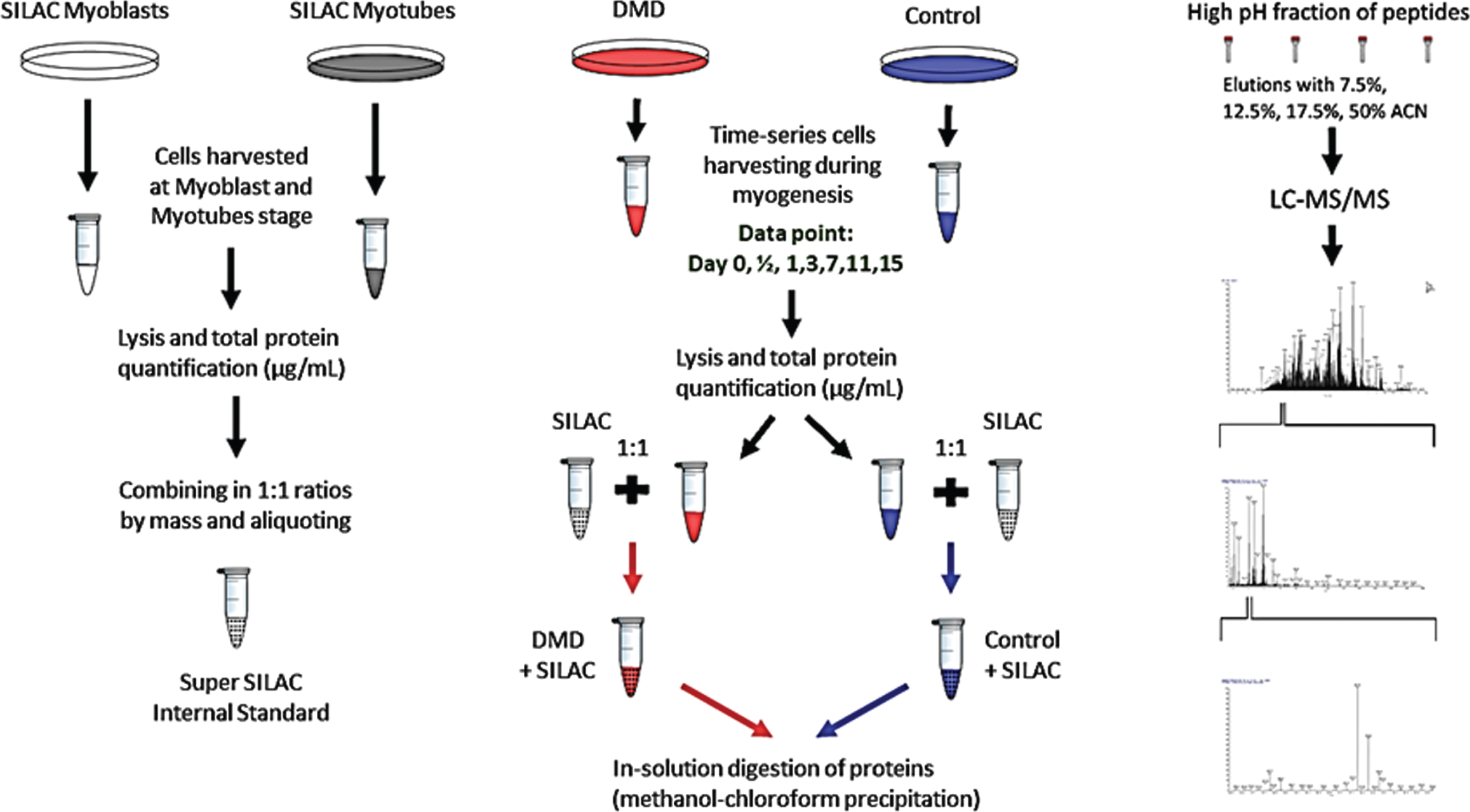
Experimental workflow for temporal proteome profiling of normal and dystrophin deficient human myoblast. Left panel showing preparation of super SILAC myoblast and myotube standard protein extracts prepared from cultured normal myoblast and myotubes incubated in SILAC media (see detail in method section). SILAC protein extracts were prepared from both myoblast and well differentiated myotubes. Then mixed at 1 : 1 ratio to for use as spik-in standard for temporal proteome profiling during differentiation of normal and dystrophin deficient myoblast. Center panel showing culture and differentiation of dystrophin and normal myotube procedure and sample preparation for quantitative proteome profiling using SILAC and LC-MS/MS analysis. At indicated times (0, 0.5, 3, 7, 11 and 15 days after differentiation, cell were harvested, Right panel showing the LC-MS/MS analysis workflow.
Similarly, a set of normal and DMD hSkMCs were cultured in regular unlabeled high glucose DMEM supplemented with PromoCell Growth Supplement, 20% FBS and 1% P/S. These myoblasts were incubated for 14 days in differentiation media containing the same DMEM supplemented with PromoCell Differentiation supplement, 2% FBS 1% P/S but without SILAC. The myotubes were harvested at seven time-points –day 0 (myoblast stage at 90% confluency, days 1/2, 1, 3, 7, 11 and 15, following the last 24-hour incubation in serum-free differentiation media respectively. All the harvested cell pellets, whether SILAC labelled or unlabeled, were stored in PBS at –80°C until lysis.
Cell lysis and protein extraction
The SILAC labelled myoblasts and myotubes (harvested at day 0 and day 14 respectively), and unlabeled myoblast and myotube cells (harvested at seven time-points) were lysed using RIPA buffer (Sigma, MO) by vigorous vortexing followed by10 minutes centrifugation at 14,000 rpm at 4°C. The supernatants containing myotube and myoblast protein extracts were collected, and the protein concentration was determined in each sample set using the Pierce™ BCA Protein Assay Kit. Protein extracts were stored into 100μg workable aliquots and stored at –80°C until needed. SILAC labeled myotube and myoblast extracts were mixed in 1 : 1 ratio to be used as internal standard for temporal proteome profiling during differentiation of normal and DMD myoblast (see below).
Sample preparation for mass spectrometry analysis
The SILAC internal standard was made by combining the SILAC myoblasts and myotubes extracts at 1:1 ratio, w/w. Aliquots containing 50μg of SILAC labeled proteins were mixed with 50μg aliquots of unlabeled protein extracts prepared from the 14 individual samples of DMD and normal cell lysates harvested at 7 different time-points of differentiation (day 0, 0.5, 1, 3, 7, 11, and 15). Each spiked sample consisted of 100μg of total protein concentration diluted with RIPA buffer to bring the volume up to 100μl. Then 300μl of water, followed by 400μl of methanol and 100μl of chloroform were added to the protein solution to delipidate the protein extracts. The samples were then vigorously vortexed and centrifuged at 14,000×g for 5 min. A thin protein layer was formed above the chloroform. The water and methanol above the protein layer was removed by aspiration using pipette tip and the chloroform below the protein layer was removed by slow and careful decanting technique. The protein pellet was washed twice with 400μl of methanol, followed by gentle vortexing and centrifugation 14,000×g for 5 min. The remaining methanol was air dried by leaving the tube open for 1 to 2 minutes. The protein pellet was then resuspended in 100μl of 6 M urea and incubated for 5 min at room temperature. The samples were reduced by adding 5μl of 200 mM DTT solution in 25 mM ammonium bicarbonate and incubated for 1 hour at 37°C. For alkylation of proteins, 20μl of 200 mM iodoacetamide solution in 25 mM ammonium bicarbonate was added and the tubes were incubated for 1 hour at room temperature in dark. Then 20μl of DTT solution was added into each tube and incubated for 10 min at room temperature to quench excess iodoacetamide. Finally, 755μl of 25 mM ammonium bicarbonate was added to dilute the urea in the samples before trypsinization step. The diluted protein samples were incubated overnight with 20μl of 0.10 ug/ul trypsin solution (Promega grade) at 37°C using a shaker at 550 rpm. The reaction was then stopped by adding 50μl of 2.5% TFA before conducting fractionation. Only 1/3rd amount of each of these tryptic digests were used for LC-MS/MS analysis and the remaining was stored at –80°C for future tests. Each sample containing complex peptide mixture was fractionated using the PierceTM High-pH Reversed-Phase Peptide Fractionation columns. Peptide fractionation was performed using sequential elution by solutions containing 7.5%, 12.5%, 17.5%, 50% of Acetonitrile in 0.1% Triethylamine. The resulting elutes were vacuum-dried and stored at –20°C until analysis.
Liquid chromatography tandem mass spectrophotometry (LC-MS/MS) analysis
The dried peptide samples were resuspended in 30μl of 0.1% formic acid, and 7μl of this peptide solution from each SILAC spiked sample was injected onto a Q Exactive™ HF-X Hybrid Quadrupole-Orbitrap™ Mass Spectrometer connected to a Dionex UltiMate 3000 RS UPLC system. A two-hour gradient was run using a mobile phase A of 0.1% formic acid in water, and a mobile phase B of 0.1% formic acid in 80% acetonitrile. An EASY-Spray™ LC Column (75μm diameter, 50 cm length, pore size 100 Å, particle size 2μm; Thermo Scientific) column was first equilibrated with 98% mobile phase A and 2% mobile phase B for 8 minutes, then 5–30% B gradient for 87 minutes, 30–100% B for 5 minutes, 100–100% B for 5 minutes, 100–2% B for 5 minute and 2% B for 10 minutes. Data-dependent acquisition was performed at 120,000 resolutions for a scan mass range of 375–1600 m/z, with one full MS scan followed by 20 MS/MS scans. DDA acquisition was done at 27 eV collision energy, 300 V ion spray and 35°C column temperature.
Protein identification was performed against a human proteome (SwissProt TaxID = 9606, v2017-07-05, ID: UP000002254, last edit: May 16, 2019) containing 25,496 proteins identification) using Sequest algorithm Proteome Discoverer package version 2.2 (Thermo Fisher Scientific, Waltham, MA). Three parallel Sequest nodes were utilized each containing a different static modification of Label: 13C (6)/+6.020Da (R), Label: 13C(6)15N(2)/+8.014Da (K), or no modification. Peptides were matched using methionine oxidation, carbamidomethylation and acetylation dynamic modifications at a 10-ppm precursor mass tolerance and 0.05 Da fragment mass tolerance. Only unique peptides were considered for the quantification purposes, and the ratios of light peptide/heavy peptide were calculated using area of precursor abundance embedded into Proteome Discoverer 2.2 tool. Peptide identification was performed using a strict FDR cut offs (0.01). To correct for SILAC spiking mixing ratio errors, a median normalization was performed for the entire generated data in each sample (supplemental Figure S1).
Data clustering and statistical analysis
For temporal clustering analysis, log2 transformed and median normalized SILAC ratios of 1,756 proteins across all 7 time points were included in the analysis. Clustering was performed for DMD and normal differentiating cells separately. Temporal SILAC ratios of each protein were normalized relative to the maximum SILAC ratio value obtained for that protein across the 7 time points. This was followed by unsupervised clustering in longclust [13] using R [14]. Model-based clustering based on mixture of t-distributions was used. As a result, clusters were represented by a matrix that contains probabilities for each data element (protein) to belong to a specific cluster group. This method enabled clustering of similar longitudinal trajectories together. To simplify data interpretation, we have focused on the top 50 proteins that belong to each cluster. This was based on selecting protein trajectories with highest probabilities to belong to a specific cluster.
To compare temporal proteome profiles of normal and DMD myoblast during differentiation, a third-degree polynomial model was used to investigate the association of time, group (normal and DMD), and their possible interaction on the log2 transformed SILAC ratios. The coefficient of interaction represents the difference of SILAC ratio interaction with time between the two groups DMD and normal differentiating myotubes. If the interaction is significant this indicates that the group (DMD or normal cells) is associated with different trajectories of protein over time. The coefficient of group represents the difference in the mean protein levels (SILAC ratios) between DMD and normal cells. For data interpretation we have reported and discussed the p values of the interaction coefficient. The interaction p value for all 1756 proteins were adjusted for multiple testing using BH method [15].
To define functionality of proteins grouped in each cluster, we used the Database for Annotation, Visualization, and Integrated Discovery (DAVID) tool [16]. The Uniprot accession IDs for each cluster were uploaded into the functional annotation tool on DAVID website and “gene ontology” biological process was selected from the summary result. Groups less than 5 were removed, some manual curation was conducted using Uniprot to group closely related biological function.
Western blot analysis
To confirm the presence of dystrophin in SILAC labelled and unlabeled myotubes, we performed western blotting on all the sample lysates. Aliquots containing 25μg of total proteins from each sample were separated on a NuPAGETM 3–8% tris-acetate protein gel at 200V for 1 hour. The proteins were transferred from the gel to a NovexTM nitrocellulose membrane overnight using NuPAGETM transfer buffer at 0.17Amp for 17 hours. Membranes were stained with Ponsceau stain to confirm efficient protein transfer. The membranes were cut and incubated with polyclonal rabbit antibody against dystrophin (ab15277, Abcam, MA, USA) and monoclonal rabbit against vinculin (SPM227, Abcam, MA, USA) as the loading control. Primary antibodies were used at a concentration of 1 : 300 for dystrophin and 1 : 10,000 for vinculin, and incubated for 3 hours. Donkey monoclonal anti-rabbit secondary antibody was used at a dilution of 1 : 4000 concentration and incubated for an hour at room temperature on a shaker. ECL reagent was used to image the membranes.
To confirm some of the data generated by temporal proteome profiling using SILAC strategy above, a subset of key identified proteins was selected for western blot analysis. These include, proliferating cell nuclear antigen (PCNA), ubiquitin-like protein ISG15 (ISG15), epidermal growth factor receptor (EGFR), disintegrin and metalloproteinase domain-containing protein 17 (ADMA17), signal transducer and activator of transcription 1-alpha/beta (STAT1), tubulin-beta 3 chain (TUBB3), calponin-3 (CNN3), and vinculin used as loading control. Aliquots containing 30μg total proteins from each sample were loaded on a NuPAGE™ 3–8% Tris-Acetate Protein Gels, 1.5 mm, 10-wells for analysis of proteins with higher molecular weight (>80 kDa) or on a Novex™ 16%, Tricine, 1.0 mm, Mini Protein Gel, 10-well for analysis of proteins with low molecular weight (<80 kDa). Electrophoresis and protein transfer was performed as above. Membranes were stained with Ponceau stain to confirm efficient protein transfer then were cut according to the molecular weight for each protein and incubated with the following mouse monoclonal antibodies from Santa Cruz: anti-PCNA (S-56, dilution 1 : 100), anti-EFGR (sc-373746, dilution 1 : 100), anti-ISG15 (sc-166755, dilution 1 : 100), anti-TUBB3 (sc-80005, dilution 1 : 200), anti-ADAM17 (sc-390859, dilution 1 : 100), anti-STAT1 (sc-464, dilution 1 : 200) and a rabbit monoclonal anti-vinculin from Abcam (ab129002, dilution 1 : 1000). Donkey monoclonal anti-rabbit and sheep monoclonal anti-mouse were used as secondary antibodies at dilution of 1 : 3000. Membranes were incubated in shaker and processed as above.
RESULTS
Temporal proteome profiling during myogenesis of normal and DMD human myoblasts
Differentiation of normal and DMD (e.g. dystrophin deficient) human myoblasts into myotubes was monitored by microscopy. Myotubes started to clearly form by day 7 of differentiation and then completely matured by day 11–15 of differentiation (Fig. 2A). There was no major phenotypic difference between normal and dystrophin deficient myotubes (Data not shown but observed under the microscope). Dystrophin protein expression, an essential protein in skeletal muscle [17], started being detectable by western blot in normal differentiating myoblast by day 0 of differentiation and gradually increased in its expression. It was most intense at day 3, 7, 11 of differentiation then slightly decreased by day 15 of differentiation (Fig. 2B). Dystrophin was not detectable in DMD myoblasts and myotubes at any time during differentiation (Fig. 2C).
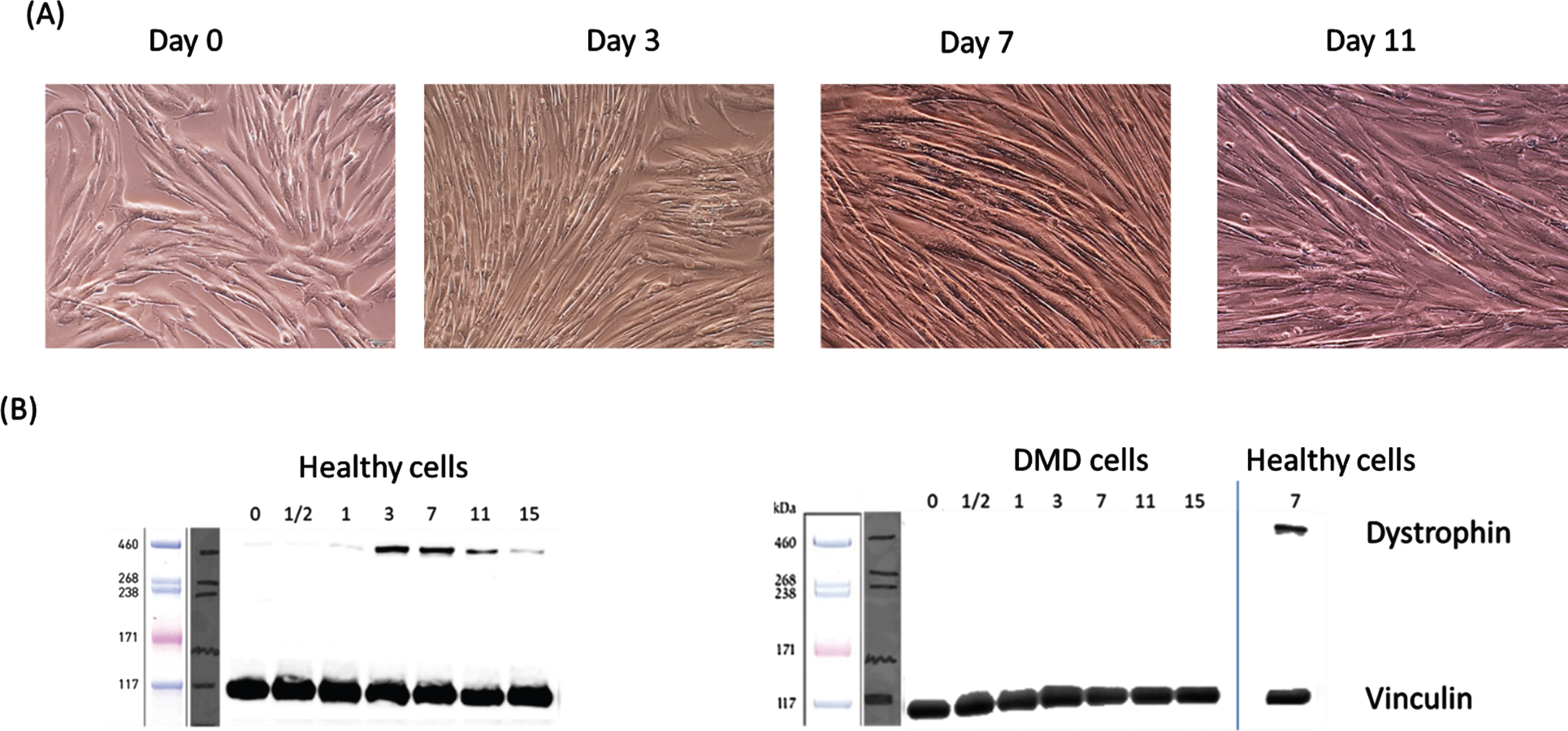
Temporal expression of dystrophin protein during differentiation of immortalized human myoblasts. Confluent DMD and normal myoblast were switched to differentiation media as outlined in the experimental method. At day 0, 0.5, 1, 3, 7, 11 and of differentiation an aliquot of harvested cells were extracted and processed for dystrophin protein expression using western blot. (A) shows confluent myoblasts entering differentiation at day 0 then formation of multinucleated and fused myotubes starting at day 3. Fully mature myotubes are shown at day 11. (B) show temporal expression of dystrophin protein during differentiation of normal (control) and dystrophin deficient myoblasts (DMD). Dystrophin was detected only in normal and not in DMD differentiating myoblast.
Normal and DMD myotubes were harvested at days 0. 0.5, 1, 3, 7, 11 and 15 of differentiation and processed for proteome profiling using Super SILAC spiking strategy as described in the method section. Temporal changes in the expression levels of individual proteins were determined by measuring the relative intensity ratios of selected peptides from a given protein in experimental cells to the homologous peptides of the same protein from the SILAC spiked-in reference. A total of 7,441 proteins were thus identified and quantified in all samples at all data points of differentiating normal and DMD myoblasts combined. However, only 1,756 proteins had quantifiable values across all data points and in both normal and DMD muscle cells.
Clustering of these 1756 proteins based on their longitudinal trajectories revealed 4 major clusters in normal myotubes (Fig. 3A). Further functional grouping using DAVID tool grouped the proteins in each cluster based on their function and biological processes (Fig. 3B) and revealed the following temporal protein expression changes and classification.
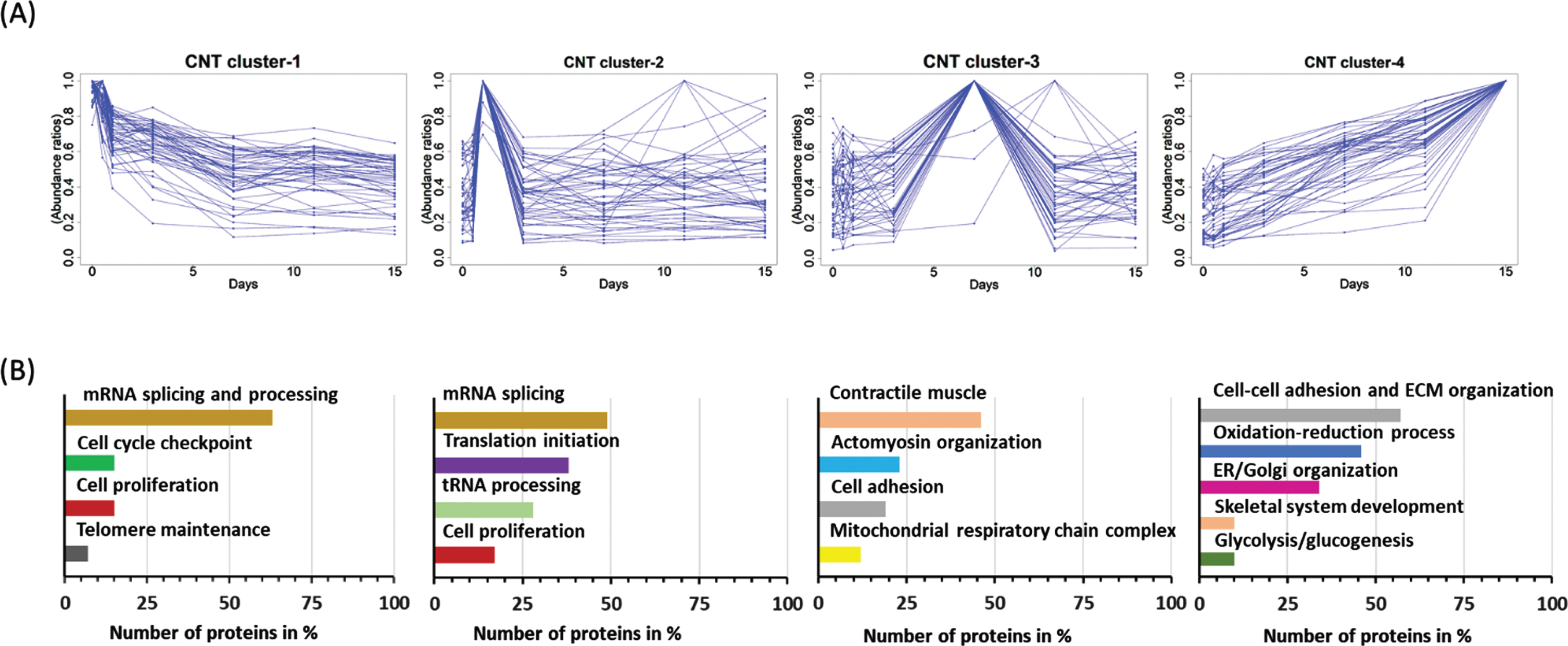
Temporal proteome profiling and data clustering during differentiation of normal myoblast. Proteome profiling was performed at day 0, 0.5, 1, 3, 7, 11 and 15 of differentiation of normal myoblasts using SILAC strategy and LC-MS/MS analysis. (A) Overall data was clustered based on pattern and dynamics of expression. Four distinguishable clusters were thus generated. (A) show temporal expression of top 50 proteins in each cluster. (B) shown GO enrichment analysis using DAVID tool of protein grouped under each cluster.
Top 50 proteins grouped in each of the clusters are listed under supplemental Table S1 and with their respective GO annotation and DAVID classification and we are only summarizing key findings here. Cluster 1 consisted mainly of proteins involved in mRNA splicing and processing, telomere maintenance as well as few proteins involved in regulation of cell proliferation. These proteins were elevated at the myoblast stage then gradually decreased to day 7 of differentiation then remained decreased thereafter. Supplemental Figure S2A shows heatmap of top 50 proteins from cluster 1 that were elevated at the myoblast stage then decreased in function of differentiation. For example, tissue–type plasminogen activator (PLAT), plasminogen activator inhibitor 1 (PAI-1) and telomerase reverse transcriptase (TERT) were approximately 7-fold more expressed at day 0 of differentiation (e.g., myoblast stage entering fusion) relative to day 11 and 15 of differentiation when myotubes were fully formed. Proliferating cell nuclear antigen (PCNA) and epidermal growth factor receptor (EGFR) was elevated by 3 to 4 folds elevated at days 0 and 0.5 of differentiation relative to days 11 and 15 of differentiation. Cluster 2 consisted mainly of additional proteins involved in mRNA splicing and processing (49 proteins) as well as proteins involved in translation initiation machinery, regulation of cell proliferation and apoptosis. Supplemental Figure S2B shows heatmap of exemplar proteins such as heterogeneous nuclear ribonucleoprotein U (HNRNPU) involved nuclear chromatin organization, regulation of telomeres and mRNA alternative splicing. This protein was elevated at day 0 and 0.5 of differentiation then gradually decreased in function of differentiation. Epidermal growth factor receptor kinase substrate 8 (EPS8) involved in actine dynamic regulation during exist from cell cycle [18] and the translation initiation factor 3 subunit F involved in translation machinery also showed similar patterns as HNRNPU. Cluster 3 consisted of protein that exhibited lower expression levels at day 0 of differentiation, gradually increased to reach a maximum expression at day 7 of differentiation then lightly decreased and remained stable thereafter. Most proteins grouped under cluster 3 consisted of proteins involved in the regulation of myosin and actin cytoskeleton organization, muscle contractile apparatus, and few proteins involved in cell adhesion and mitochondria morphogenesis. Supplemental Figure S2C shows heatmap temporal expression of exemplar proteins including myosin regulatory light polypeptide 9 (MYL9), myosin regulatory light chain 12B (MYL12B), xin actin-binding repeat-containing protein 1 (XIRP1), tropomodulin 3 (TMOD3), muscle contractile proteins such as myosin light chain 1/3, (MYL1) myosin light chain 9 (MYL9), myosin heavy chain 9 (MYH9) and tropomyosin 2 beta (TPM2) and some mitochondria morphogenesis proteins such as polyribonucleotide nucleotidyltransferase (PNPT1) [19]. Finally, cluster 4 comprised the largest number of regulated proteins whose expression was low at the myoblast stage then gradually increased to reach maximum expression at day 11 and 15 of differentiation. Proteins grouped under this cluster consisted mainly of protein involved in cell-cell adhesion, extracellular matrix organization, the oxidation-reduction processes, actine and actomyosin organization, sarcomere organization, proteasome-mediated ubiquitin-dependent protein catabolic process, protein folding and glycolytic/glucogenesis process. Supplemental Figure S2D shows heatmap temporal expression of representative proteins in this group including ubiquitin/ISG15-conjugating enzyme E2 L6 (UBE2L6), actine stabilization such as drebrin-like protein, collagen type I alpha 1 chain (COL1A1), collagen type XII alpha 1 chain (COL12A1), lami-nin subunit alpha–5, prolyl 4-hydroxylase subunit alpha 1(P4HA1) and caveolae–associated protein 1 (CAVIN1) just to highlight few.
Lack of dystrophin expression in DMD derived myoblast perturbed temporal expression of some key regulated proteins identified in differentiation of normal myoblasts
Although dystrophin deficient DMD derived myoblasts did not exhibit major phenotypic alteration during myogenesis, major alterations occurred in the temporal expression of some proteins. Comparative temporal proteome profiling identified 162 proteins whose temporal expression was significantly altered (adjusted p-value for multiple testing < 0.05) in differentiating dystrophin deficient myoblasts compared to differentiating normal myoblasts (see supplemental Table S2 for the complete list). Of these, 99 proteins had a declining expression while the remaining 63 proteins had and increasing expression in function of differentiation of DMD myoblast compared to normal control myoblast. Most of these temporal expression alterations observed during differentiation of dystrophin deficient myoblasts occurred in proteins belonging to cluster 4 and that are associated with the final myotube maturation and organization. Figure 4 shows heatmap of the top 50 proteins that were altered in their temporal expression (34 decreased and 16 increased) during differentiation of dystrophin deficient myoblasts relative to normal myoblast. Functional grouping further grouped these proteins into cell adhesion proteins, focal cell adhesion (extracellular matrix organization, some mitochondria associated proteins. Figure 5 shows examples of temporal expression of some relevant proteins to myoblast differentiation and maturation such as those involved in cell-ECM interaction [20] such as laminins (e.g. LAMA5, LAMC1), caveolae associated protein 1 (CAVIN1), COL1A1 and ADAM17 (Fig. 5A). A significant decline was also observed for few proteins associated with mitochondria, such as ATP synthase subunit d (ATP5PD), NADH dehydrogenase [ubiquinone] iron-sulfur protein 2 (NDUFS2), calcium-binding mitochondrial carrier protein SCaMC-1 (SLC25A24) and glutamate dehydrogenase 1 (GLUD1) (Fig. 5B). Another significant decline in the temporal protein expression during differentiation of DMD myoblast compared to normal myoblast included a group of proteins involved in the ubiquitination such as Ubiquitin–like protein ISG15 (ISG15), E3 ubiquitin–protein ligase DTX3L (DTX3L), Ubiquitin/ISG15-conjugating enzyme E2 L6 (UBE2L6), just to cite few and a group of the proteasome complex such as proteasome subunit beta type–8 (PSMB8), proteasome activator complex subunit 1 (PSME1), and a large group of proteins involved in the interferon pathway such as interferon-induced protein with tetratricopeptide repeats 1 (IFIT1), signal transducer and activator of transcription 1–alpha/beta (STAT1), probable ATP–dependent RNA helicase DDX58 (DDX58), 2′-5′-oligoadenylate synthase 3 (OAS3). Figure 3C shows an example from each category. These proteins and additional ones (Fig. 4 and Supplemental Table S2) declined in their temporal expression levels during differentiation of dystrophin deficient myotubes compared to normal myotubes and their potential role in differentiation.
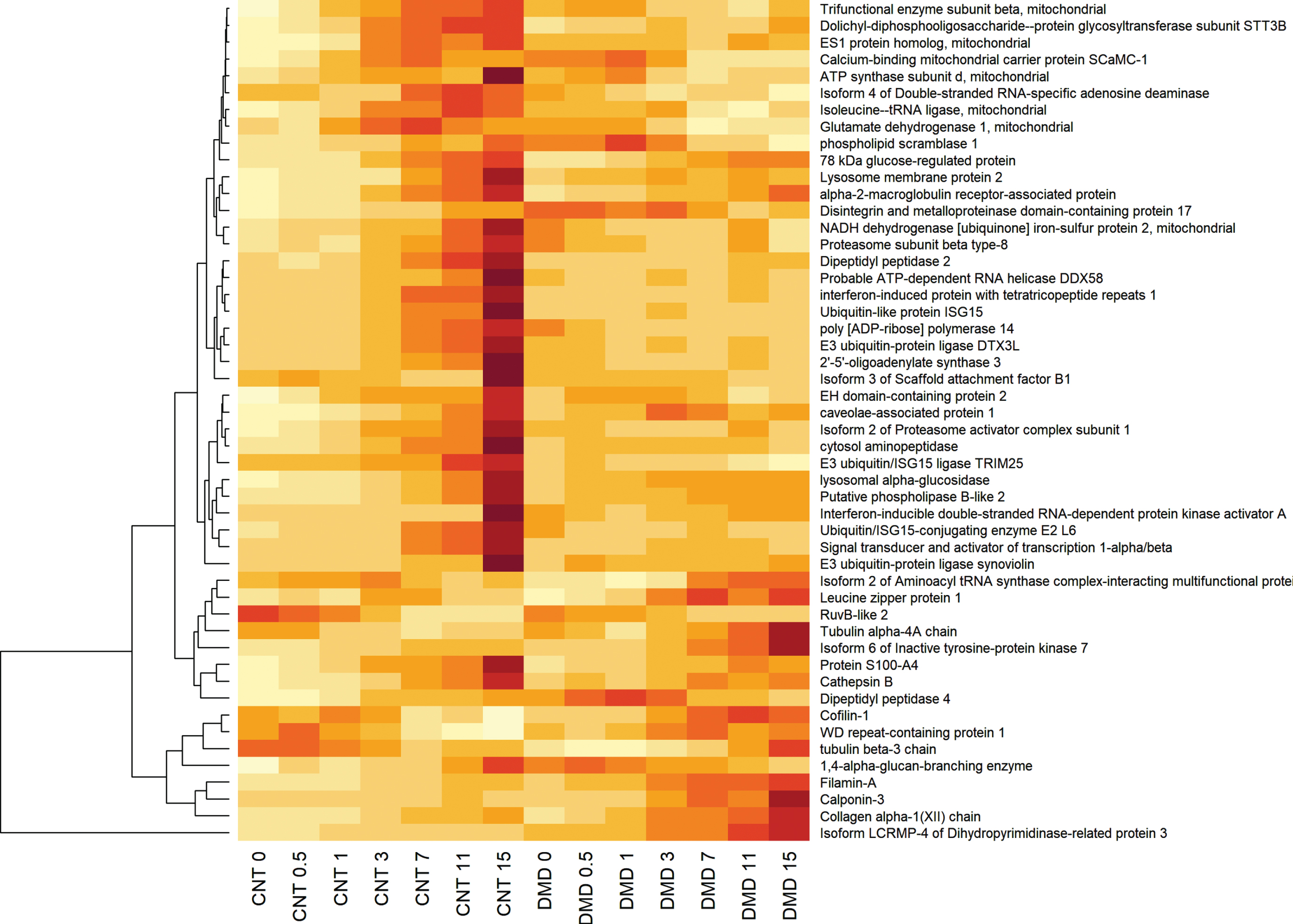
Hierarchical clustering of the top 50 proteins that were altered in their temporal expression during differentiation of dystrophin deficient myoblasts relative to normal myoblast. Yellow color gradient means decreased expression while red gradient means increased expression and represent expression levels of each protein at day 0, 0.5, 1, 3, 7, 11 and 15 of differentiation of normal (CNT) and dystrophin deficient (DMD) myoblasts.
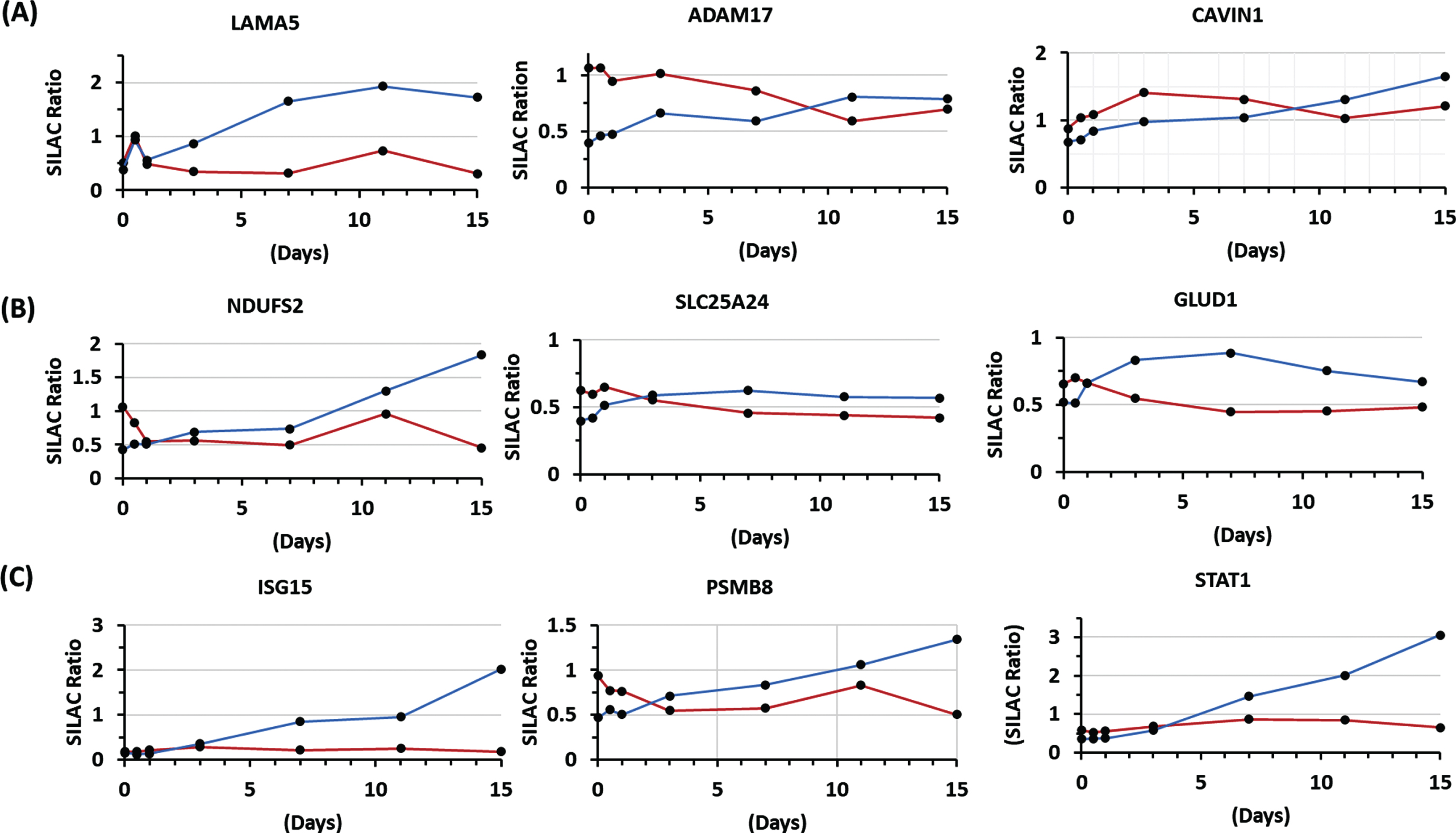
Exemplar graphs showing differential decrease in the temporal expression of some proteins during differentiation of dystrophin deficient (red line) and normal (blue line) myoblasts. (A) Example of proteins involved in cell-ECM adhesion (LAMA5: Laminin alpha 5, ADAM17: Disintegrin and metalloproteinase domain-containing protein 17, CAVIN1: Caveolae-associated protein 1). (B) Example of mitochondria associated proteins (NDUFS2: NADH dehydrogenase [ubiquinone] iron-sulfur protein 2, SLC25A24: Calcium-binding mitochondrial carrier protein SCaMC-1, GLUD1: Glutamate dehydrogenase 1), (C) Examples of a protein involved in ubiquitination (ISG15: Ubiquitin-like protein ISG15), a protein belonging to the proteasome complex (PSMB8: Proteasome subunit beta type-8) and a protein involved in the interferon pathway (STAT1: Signal transducer and activator of transcription 1-alpha/beta).
Interestingly temporal expression of another group of proteins increased during differentiation of DMD myotubes compared to normal myotubes. Figure 6 shows few examples such the differential increased expression of COL12A1, alkaline phosphatase tissue type (ALP), transgelin, dihydropyrimidinase-related protein 3 (DPYSL3), calponin-3 (CNN3) and beta class tubulins. The complete list of proteins whose temporal expression differentially increased during differentiation of DMD relative to normal myoblast is provided under Supplemental Table S2) and the significance of this increase is discussed below for some proteins.
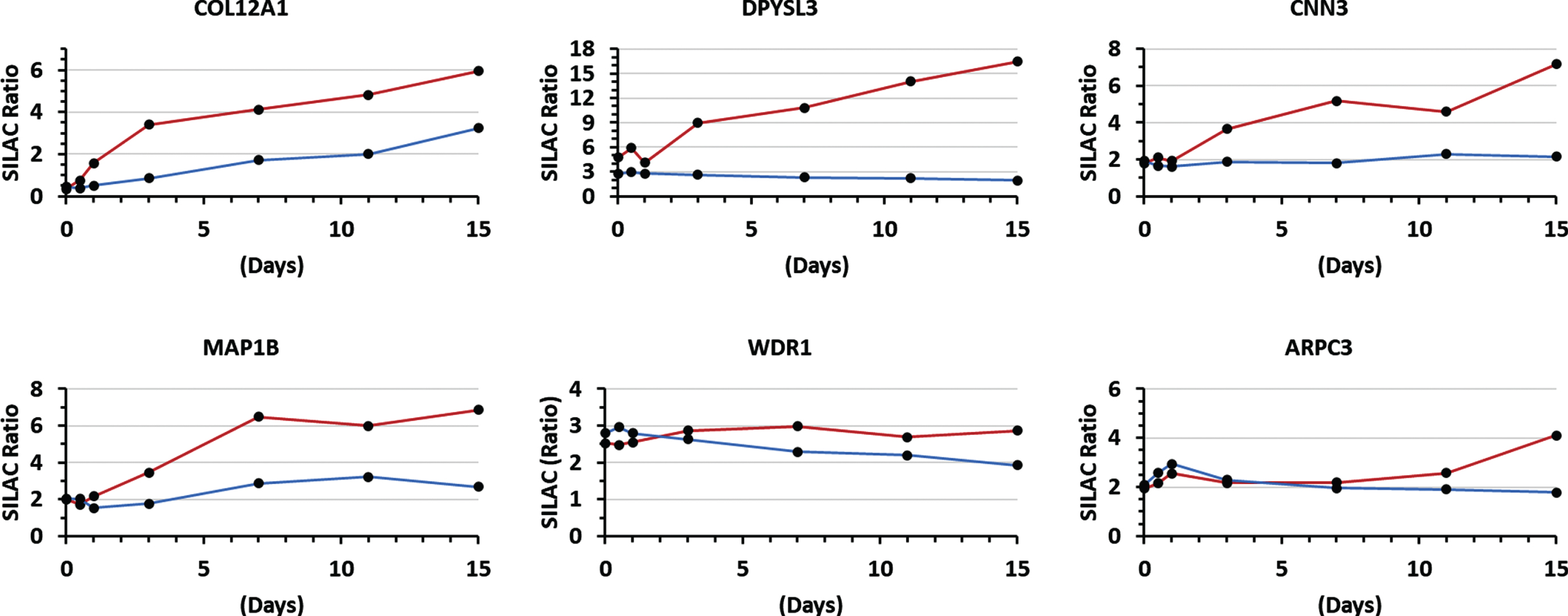
Exemplar graphs showing differential increase in the temporal expression of some proteins during differentiation of dystrophin deficient (red line) and normal (blue line) myoblasts. Examples are shown for an ECM protein (COL12A1: collagen alpha-1(XII) chain) and proteins involved in cytoskeleton remodeling and microtubules organization (DPYSL3: dihydropyrimidinase-related protein 3, CNN3: calponin-3, MAP1B: microtubule-associated protein 1B, WDR1: WD repeat-containing protein and ARPC3: actin-related protein 2/3 complex subunit 3.
To confirm some of the temporal proteome profiling data, western blot analyses were performed for some key proteins for which there was a good quality antibody available. Figure 7 shows western blot data for as set of selected proteins including PCNA. ISG15, STAT1, CNN3, TUBB3 and ADAMS17. PCNA clearly decreased during differentiation of both normal and dystrophin deficient cells while ISG15 increased at the late stage of differentiation of normal myoblasts and remained undetectable in DMD myoblast across all data points. CNN3 slightly increased in both normal and DMD myoblast during differentiation but decreased in normal myoblast while in continued to increase in DMD myoblast at day 15 of differentiation. STAT1 increased overtime during differentiation of normal myoblasts but decreased overtime during differentiation of DMD myoblasts. TUBB3, gradually increased during differentiation of DMD myoblasts but remained evenly expressed during differentiation of normal myoblasts. Finally, ADAM17 decreased overtime during differentiation of both normal and dystrophin deficient myoblast. This western blot data agrees with our temporal proteome profiling data further demonstrating the robustness of SILAC strategy to measure changes in protein levels during differentiation of normal and dystrophin deficient myoblasts. Other tested proteins such as EGFR and DPYSL3 did not validate due to nonspecific antibody binding during western blot.
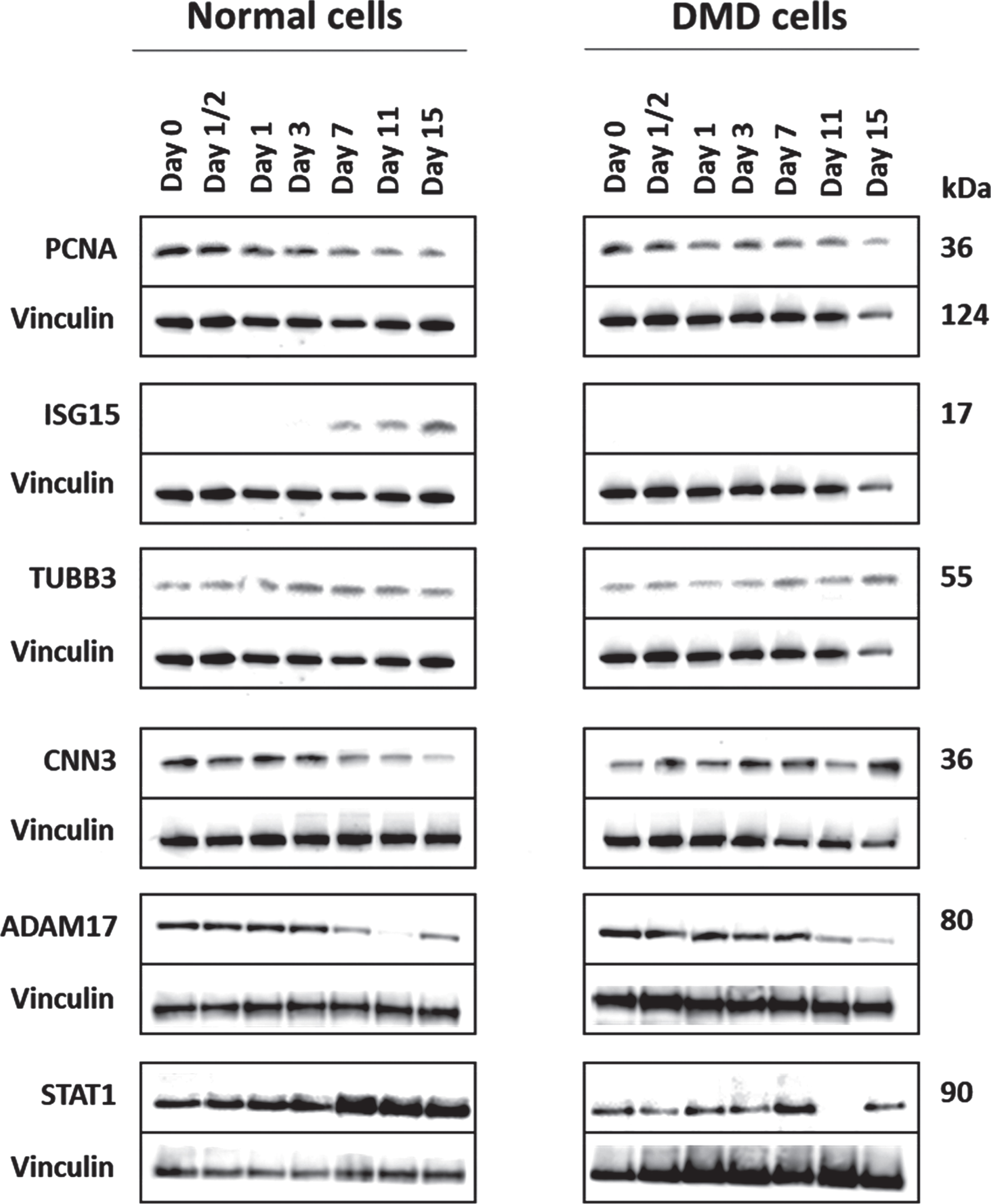
Representative western blots confirming temporal changes in the expression of some key proteins in normal and dystrophin deficient myoblasts during differentiation. PCNA (proliferating cell nuclear antigen); ISG15 (ubiquitin-like protein ISG15), TUBB3 (tubulin beta-3 chain), CNN3 (calponin-3), ADAM17 (disintegrin and metalloproteinase domain-containing protein 17), STAT1 (signal transducer and activator of transcription 1-alpha/beta). Aliquots containing 30μg of each sample were used for the western blot analysis and vinculin was used as a loading control. This analysis was performed on replicate samples.
DISCUSSION
Differentiation of myoblasts into myotubes involves complex and sequential changes in protein expression. Understanding the dynamics and changes of protein expression that occurs during myoblast differentiation into myotubes might help identify potential pathway that are important in muscle regeneration under healthy and diseased conditions. Earlier proteome profiling study used C2C12 cell line to define temporal changes in protein expression during differentiation into myotubes. This provided useful insights about protein expression dynamics involved in myogenesis and identified key changes in protein expression associated with myogenesis. Although C2C12 cell line has been criticized as not being the ideal cell model to study skeletal muscle biology, it provided useful insights about protein expression dynamics and changes associated with myogenesis [5] that agree with a more recent study that used triple SILAC strategy to study temporal protein expression changes associated with differentiation of primary human myoblast into myotubes [8]. In this study we used human myoblast cells lines that were derived from muscle biopsies of a healthy subject and a DMD patient and immortalized by transduction with both human telomerase reverse transcriptase and cyclin dependent kinase 4 expressing vectors [12]. Both normal and DMD derived myoblasts formed well defined multinucleated myotubes by day 7 of differentiation. Normal myoblasts but not DMD derived myoblasts expressed dystrophin protein as revealed by western blot and proteomic analysis. Interestingly although DMD derived myoblast lacked dystrophin expression, they were not phenotypically different from that of normal myoblast expressing dystrophin. However, several differences at the proteome levels were identified and are described below. Using super SILAC strategy [11] in combination with LC-MS/MS analysis and clustering analysis enabled unambiguous characterization of temporal changes in protein expression during differentiation of both normal and dystrophin deficient immortalized human myoblast. Here we will first define temporal changes in protein expression in normal myotubes then examine the effect of lack of expression of dystrophin on these proteome profiles in DMD derived myoblasts. Temporal proteome profiling of normal myoblast in function of differentiation and cluster analysis identified 1,057 protein that clustered into 4 major sequential clusters based on their temporal expression. Because of the large number of regulated proteins (see supplemental table S1) we will only focus on most relevant pathways.
Proteins involved in myoblast cell cycle exit and differentiation
Perhaps the most striking changes in temporal protein expression were observed in the first and second cluster when myoblasts started switching from proliferating to differentiating state. These two-cluster consisted mainly of proteins involved in mRNA splicing and processing, regulation of cell proliferation, and in translation initiation machinery in agreement with previous study [8]. Interestingly the largest group of proteins belonging to clusters 1 and 2 (61 proteins) were those involved in mRNA splicing and processing. Extensive studies have shown the importance of mRNA regulation and processing during myogenesis and muscle development [21, 22], and will not be further discussed here due to the lengthy list of proteins. The second largest group of proteins belonging to clusters 1 and 2 consisted of proteins involved in the translation initiation machinery such as eukaryotic translation initiation factor 3 subunits that were previously reported to be involved in muscle myogenesis and differentiation [23, 24], and several other ribosomal protein subunits and will also not be discussed here further here. We will rather focus on some key proteins involved in myoblast differentiation initiation and myotubes organization. Some of the early changes included a sharp decrease in the expression of EGFR, PCNA, proliferation-associated protein 2G4 (PA2G4) and EPS8. These findings agree with earlier studies demonstrating that the decline in PCNA expression preceded expression of developmental sarcomeric myosin during differentiation of satellite cells isolated from rat muscle fibers [25]. The decline in PA2G4 was reported to play a role in the regulation of ErbB3 binding protein-1 that controls proliferation and differentiation of muscle satellite cells [26]. A more recent study demonstrated the role of ESP8 in the progress of myoblast fusion [27]. These proteins are all involved in positive regulation of cell proliferation and their decreased expression agrees with myoblast cell proliferation arrest. Additional proteins that belonged to cluster 1 and 2 and decreased in their expression in function of differentiation included PAI-1 and TERT. Our data agrees with previous studies supporting the role of PLAT in myogenesis [28, 29]. PLAT is a protease that activates plasmin, a potent fibrinolysis factor. This process is required for myogenesis and muscle fiber regeneration [30]. TERT, although it was used to immortalize the studied myoblasts herein and was overexpressed in these cells, it still exerted its cell cycle exit expression pattern. It was elevated at day zero of differentiation then gradually decreased during differentiation of both normal and DMD myoblasts, in agreement with earlier report about the role of telomerase in cell cycle exit and differentiation of myoblasts [31, 32].
Proteins involved in the organization of cytoskeleton, the contractile machinery and cell adhesion
This class of proteins were grouped under Cluster 3 and were expressed at low levels at the myo-blast stage (day 0 of differentiation) then gradually increased in their expression to reach maximum expression at day 7 of differentiation. Then after day 7 of differentiation some of these proteins decreased while others plateaued. Functional annotation using DAVID revealed three major classes of proteins. Those involved in actin and cytoskeleton organization such as tropomodulin-3 (TMOD3), an actin capping factor that blocks actin elongation and depolymerization and stabilizes it [33]. This phenomenon is required for the development of striated muscle [34]. Myosin-9 (MYH9), involved in actomyosin structural organization and was reported by other as being essential for cytoskeleton organization and myoblast fusion [35]. MYH9 sharply increased in its expression at day 7 then decreased thereafter of differentiation further supporting its role in myoblast fusion and myogenesis. Myosin regulatory light polypeptide 9 (MYL) followed same temporal expression pattern in function of differentiation as MYH9. Although the exact role of MYL9 in myogenesis remain to be elucidate a recent study showed that its expression in skeletal muscle dramatically increased in myostatin-knockout pigs further supporting its role in muscle development [36]. Other muscle specific proteins such as tripartite motif-containing protein 72 (TRIM72), and contractile machinery such as tropomyosin’s (TPM2, TPM4), troponins (TNNI1 and TNNT2), myosin light chain 4 (MYL4), myosin light polypeptide 6 (MYL6), myosin regulatory light chain 12B (MYL12B) and myosin phosphatase Rho–interacting protein (MIRP) also increased in their expression levels at day 7 of differentiation further supporting the development of typical muscle myotubes. TRIM72 gradually increased from day 0 to day 7 of differentiation then sharply decreased at day 11 and day 15 during myotubes formation. TRIM72 is a muscle specific protein that facilitate plasma membrane repair and other function [37] but was also shown to act as negative feedback regulator of myogenesis [38], hence its temporal expression pattern in function of myoblast differentiation (sharp increase up to day 3 and 7 then sharp decrease to day 11 and 15 of differentiation).
Nexilin (NEXN), torsin-1A (TRO1A) and galec-tin-3-binding protein (LG3BP) were also grouped under cluster 3. All three proteins are involved in actin organization and cell adhesion [39]. Their exact role in myoblast differentiation and myotube organization remains to be further studied. Nevertheless, NEXN has been shown to play a role in caping actin in cardiac muscle cells and might also interact with actin in skeletal muscle cells [40].
Proteins involved in myotubes organization and homeostasis
This class of proteins were grouped under cluster 4. They exhibited the lowest level of expression at day 0 of differentiation then gradually increased in function of differentiation to reach maximum expression at the end of differentiation. A large portion of cluster 4 proteins consisted of protein involved in oxidation-reduction processes and metabolism (49 proteins). The increase of these protein at the end stage of differentiation is most likely due to the contractile activity of myotubes leading to release of superoxide which in turn requires increased proteins and enzymes to maintain redox homeostasis of myotubes in culture [41]. The second major class of proteins whose expression levels increased at the myotube stages consisted of proteins involved in cell adhesion and extracellular matrix organization (48 proteins). These proteins are required to maintain cell-cell adhesion and interaction of myotube’s sarcolemma with extracellular matrix. Some key proteins under this category included integrin alpha-11 (ITGA11), cadherin-13 (CDH13), talin-2, laminins (LAMA5, LAMB1, LAMB2, LAMC1), CAVIN1, SAFB, delta-sarcoglycan, fibrillin-1, transmembrane glycoprotein NMB (GPNMB) and several collagens (COL8A1, COL5A2, COL5A1, COL1A1, COL3A1, COL1A2, COL6A1, COL12A1). While most of these cell adhesion proteins are well known to contribute to ECM organization and myotube integrity [42], Other reports have demonstrated the role of less common proteins in myotube maturation such as SAFB, talin-2, and CAVIN1. Caveolin-cavin system has been reported to play a role in sarcolemmal organization and muscle fiber integrity in zebra fish as a model [43]. Knockdown of CAVIN1 expression resulted in a major disruption of the sarcomere organization in the muscle fibers of zebra fish. Thus, increased expression of CAVIN1 and other caveolae associated protein of normal myotubes supports their sarcolemma organization and overall maturation. Talin-1 and 2, that connect the integrins to actin cytoskeleton, have been shown to also play a major role in sarcomere assembly and organization [44] and C2C12 was reported to act as positive regulator of differentiation of C2C12 myoblasts [45]. The third major group of proteins whose expression increased at myotube stages included those involved in vesicle trafficking and ER to Golgi transport (23 proteins total), The increase in the expression of this class of proteins suggests sarcoplasmic reticulum development and myotube maturation [46–48]. Another interesting group of proteins found in cluster 4 and whose expression increased at myotube stage consisted of proteins involved in endocytosis (e.g. EH domain–containing protein 2, sorting nexins, clathrin interactor 1, NPC intracellular cholesterol transporter 1, early endosome antigen 1), protein turnover (e.g., cathepsins B, D and Z, collagen catabolism, some proteosomes and ubiquitination machinery), and glycolysis/glucogenesis (e.g. fructose-bisphosphate aldolase C, triosephosphate isomerase, glyceraldehyde-3-phosphate dehydrogenase, phosphoglucomutase-1, L-lactate dehydrogenase A, glucose-6-phosphate isomerase, ADP-dependent glucokinase). Altogether, the increased expression of all these cluster 4 proteins at day 11 and 15 of differentiation suggest mature and functional myotubes resembling muscle fibers biochemically.
Lack of dystrophin expression altered key proteins involved in myotubes proper differentiation and maturation
Although dystrophin deficient myotubes looked like normal myotubes under the microscope, alterations in temporal expression of several proteins were detected (n = 151) in DMD myoblast during differentiation. Most of these altered expressions (> 80%) belonged to proteins under cluster 3 and 4 suggesting that they occurred at the myotube maturation stage. This agrees with a study showing some perturbations at different stages of differentiation [49]. Perhaps some of the most relevant alteration in temporal expression of proteins during maturation of dystrophin deficient myotubes is the significant decrease in the expression of protein in volved in cell-extracellular matrix adhesion and cell-cell junction such as laminins (especially LAMA5 and LAMB2), caveolae-associated protein 1 and 2 (CAVIN1 and 2), Integrin-linked protein kinase, desmoplakin, ADAM17 [50]. Laminins are well known for connecting muscle cell sarcolemma to the extracellular environment via interaction with a-dystroglycan (a-DG) of the dystrophin glycoprotein complex [9, 17]. This decreased expression of laminin could be a feedback regulatory expression due the lack of dystrophin. This hypothesis is supported by a study in the special issues showing that laminin alpha 2 turnover was increased in mdx mice compared to wild type mice [51] (see Novak et al. under this special issue) and by a more recent temporal proteome profiling study conducted on skeletal muscle of wild type and mdx mouse model [52].
Caveolins (CAVIN1 and 2) have been shown to be enriched in the sarcolemma fraction of rat skeletal muscle [53] and they have been also reported to be implicated in muscular dystrophy [54]. Furthermore, a recent study has shown that lack of expression of CAVIN1 in a mouse model caused some type of muscular dystrophy same as in human with secondary CAVIN1 deficiency [55, 56]. However, the decrease of CAVIN1, due to lack of dystrophin expression is novel and remain to be further investigated. ADAM17 also known as TACE is involved in the processing of the membrane-bound TBF-alpha precursor to release soluble precursor but has been also shown to play a role in the mechanotransduction of MAPK during myogenesis [50]. Its decreased temporal expression in DMD myoblast during differentiation could be driven by the lack of dystrophin expression but further studies are needed to support this hypothesis.
Significant decrease in several mitochondrial associated proteins (e.g, NADH ubiquinones, calcium-binding mitochondrial carrier protein SCaMC-1, ES1 protein homolog, trifunctional enzyme subunit beta, ATP synthase subunit d, Glutamate dehydrogenase 1, adenylate kinase 2, phosphoenolpyruvate carboxykinase, proton/calcium exchanger protein) was also observed at late-stage differentiation of dystrophin deficient myotubes. These alterations in mitochondrial associated proteins could explain the impaired mitochondrial oxidative phosphorylation reported in skeletal muscle of mdx mouse model for DMD [57]. The decrease in several proteins involved in the interferon pathway in dystrophin deficient myoblast during differentiation, is novel to this study and remain to be carefully examined although a study conducted on C2C12 demonstrated the role of interferon gamma in protection of C2C12 cells from apoptosis [58].
Interestingly temporal expression of few other proteins increased during differentiation of DMD myotubes compared to normal myotubes. These included a temporal increase in the expression of COL12A1, alkaline phosphatase tissue type (ALP), transgelin (TAGLN), actin-related protein 2/3 complex subunit 3 (ARPC3) thin filament binding protein calponine-3 (CCN3), dihydropyrimidinase-related protein 3 (DPYSL3), microtubule-associated protein 1B (MAP1B) and beta class tubulins (e.g. TUBB3). Although the exact significance of the increase temporal expression of these proteins in the absence of dystrophin remain to be further examined, few of these proteins such as CNN3 and DPYSL3, have been shown to interact with actin and are involved in cell muscle development [59–61]. Furthermore, a recent study has shown that knockdown of CNN3 expression impairs proliferation and differentiation of C2C12 into myotubes [62]. Another study demonstrated that ALP was expressed and localized specifically to undifferentiated type 2C fibers in skeletal muscle of DMD patients [63] further supporting our observation that ALP also increased in its expression in dystrophin deficient myoblast during differentiation could be related to a biochemical downstream consequence of lack of expression of dystrophin. Nevertheless, the exact role of the increased expression of these actine and ECM interacting proteins remain to be carefully examined. It could be a compensatory mechanism to maintain the myotube integrity in the absence of dystrophin or a disorganized cytoskeleton. Either way, further studies are needed to validate these finding on skeletal muscle tissues from mouse models and from human specimens and further study the mechanistic aspect of these alterations using in vitro and in vivo experiments.
A major limitation of our study is the use of one normal and one DMD cell line. Analysis of additional normal and dystrophin deficient cell lines is important to fully interpret the data in a meaningful way. Nevertheless, a subset of generated data using these two cell lines agrees with previous studies conducted on C2C12 [5, 64], other primary cell cultures of human and animal model origins [8, 65] and more recently on human pluripotent stem cell models for myogenesis in the presence and absence of dystrophin [66]. Interestingly, some key proteins (e.g. LAMA5, LAMB2, prelamin A/C, COL6A1) that were decreased in dystrophin deficient cells relative to normal cells at the myotube stage agree with a more recent temporal proteome profiling performed on skeletal muscle of wild type and dystrophic mouse model at different ages [52]. Further evaluations using muscle biopsies from DMD and age matched healthy controls are ongoing in our laboratory to validate some of the key proteins identified in this pilot study.
Footnotes
ACKNOWLEDGMENTS
This work was partially supported by the US Department of Defense CDMRP program grant Award # W81XWH-16-1-0572 and the NIH center grant P50HD090254. The authors would like to thank the Decker Foundation for supporting the purchase of the state-of-the-art mass spectrometry instrument.
CONCLUSION
Using super SILAC strategy, useful temporal proteome profiles associated with myoblast differentiation were generated. These profiles were sequential and provided insight about molecular changes associated with myoblast cell cycle exit and myotube maturation in normal and dystrophin deficient cells. Dystrophin deficiency perturbed the temporal expressions of some proteins mainly at the late stage of differentiation indicating the importance of dystrophin protein in the maturation of myotubes. Those alterations could prove useful therapeutic target in DMD. However, validation studies using an in vivo system are needed to confirm our in vitro generated data herein.
AUTHOR CONTRIBUTIONS
This experimental design and rationale for this study was conceptualized by MVG and YH. Mass spectrometry analysis was conducted by EHC. Statistical modeling and analyses were performed by SMT. Manuscript was written by MVG and YH and revised by SMT and EHC; final manuscript was reviewed and approved by all authors prior to submission.
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
