Abstract
Background:
Exon skipping strategies in Duchenne muscular dystrophy (DMD) have largely been directed toward altering splicing of exons flanking out-of-frame deletions, with the goal of restoring an open mRNA reading frame that leads to production of an internally deleted but partially functional dystrophin protein.
Objective:
We sought to apply exon skipping to duplication mutations, assuming that the inherently limited efficiency of antisense oligonucleotide-induced exon skipping would more frequently skip a single copy of a duplicated exon, rather than both and result in significant amounts of wild-type
Methods:
We tested this hypothesis in fibroblast cell lines derived from patients with a variety of single or multiple exon duplications that have been modified to allow transdifferentiation into a myogenic lineage.
Results:
Using a variety of 2’O-methyl antisense oligonucleotides, significant skipping was induced for each duplication leading to a wild-type transcript as a major mRNA product.
Conclusions:
This study provides another proof of concept for the feasibility of therapeutic skipping in patients carrying exon duplications in order to express wild-type, full-length mRNA, although careful evaluation of the skipping efficiency should be performed as some exons are easier to skip than others. Such a personalized strategy is expected to be highly beneficial for this subset of DMD patients, compared to inducing expression of an internally-deleted dystrophin.
INTRODUCTION
Duchenne muscular dystrophy (DMD) is the most common inherited disease of muscle, affecting 1 in 5,266 newborn males [1]. This devastating disorder occurs due to mutations in the
It is usually expected that exon skipping in patients with out-of-frame deletions results in a slower disease progression, replacing a DMD (out-of-frame) mRNA with a Becker muscular dystrophy (BMD, in-frame) mRNA. However, this may be insufficient to significantly modify the disease course in some cases, because the resulting internally-deleted BMD-like protein isoforms could have altered stability or molecular function. Deletions of one or more exons account for around 65% of all dystrophinopathy (DMD or BMD) mutations, whereas duplications account for between 6 and 11% in several large cohorts [3, 6]. Although duplication mutations are less common than deletions, they are a reasonable target for therapy, as skipping of one copy of a single duplicated exon should result in expression of a full-length dystrophin molecule, an attractive therapeutic goal [7, 8]. Several groups have explored this approach. In one study in cultured muscle cells from patients with duplications of exon 44 or exon 45, skipping exon 44 was too efficient, leading to the skipping of both exons and another out-of-frame transcript, whereas skipping of exon 45 was not too efficient and did not require skipping of an additional exon [7]. Another study compared skipping of isolated exon 2 duplications versus duplications of exons 2–7; unsurprisingly, efficient multiexon skipping was more difficult to achieve [8].
At first glance, this therapeutic strategy seems challenging because skipping of both copies of an out-of-frame duplicated exon would result in an out-of-frame deletion transcript, resulting in no clinical benefit (albeit with no clinical harm, either). However, such highly efficient exon skipping has not been demonstrated in human trials to date [4, 9]. Here we sought to test the feasibility of single duplication exon skipping in patient-derived cell lines using 2’O-methyl phosphorothioate antisense oligonucleotides (AONs). Our data demonstrate in additional cell lines that single exon skipping is titratable, and that high levels of wild-type (WT) transcript can be reached in cell culture, implying that a route toward duplication skipping therapy exists.
MATERIALS AND METHODS
Establishment of FibroMyoD cell lines
Following informed consent obtained under an Institutional Review Board-approved protocol, a skin punch biopsy was performed under local anesthesia. Primary human cells fibroblasts were isolated from biopsies by enzymatic dissociation with collagenase 1A (Sigma- Aldrich, St. Louis, MO, USA). Cells were expanded in the proliferation medium composed of DMEM 1 g/L glucose (Gibco), 20% fetal bovine serum (FBS; PAA Laboratories GmbH, Pasching, Austria), 100 mg/mL penicillin, 100 mg/mL streptomycin (Biowest, Nuaille, France), and L-glutamine (Gibco, Thermo Fisher Scientific, Waltham, MA, USA). Cells were immortalized using a lentiviral hTERT/puromycin selection construct, and subsequently infected with a tetracycline-responsive MyoD expression lentiviral construct with a hygromycin selection marker (derived from a plasmid gift from the laboratory of V. Mouly, Paris) [10]. For lentiviral infection, primary fibroblasts were plated into a 12-well plate and then infected when they reached approximately 30% confluence. The following day 1 ml of proliferative media was added. One to two days later, cells were split into 6-well plates and grown until reaching 70% confluency, at which point proliferative media was complemented with 400 μg/ml of hygromycin and 1 μg/ml of puromycin, and the cells (FibroMyoD cells) were maintained under selection for 12 days.
Myoblast/myotube induction
FibroMyoD cells were plated on laminin-coated plates and cultured to 70% confluency, at which time the media was switched to SkBM-2 complete media (Lonza, Basel, Switzerland) with the addition of doxycycline (4 μg/ml; Sigma) to induce differentiation to myoblasts. At 90–95% confluency (2–3 days later), the media was changed to Skeletal Muscle Cell Differentiation Medium (C-23061, Promocells, Heidelberg, Germany) with doxycycline 4 μg/ml, in which cultures were maintained.
Antisense oligonucleotides transfection and RNA extraction
Antisense oligonucleotides transfection were performed at the myoblast stage, at 90% confluence and prior to myotube differentiation. 400 μl ml/well of pre-warmed OptiMEM (Thermo Fisher Scientific) media was added to the myoblasts and the plate was transferred to a 5% CO2 incubator for 1 h. Various concentration of AONs were diluted in OptiMEM, and transfection using Lipofectamine 2000™ was performed according to the manufacturer. Briefly, both AON and Lipofectamine solution were mixed separately gently by pipetting. Both solutions were combined and incubated for 10–15 min at room temperature to allow complex formation, and the mix was carefully dropped onto the 6 well plate. The following morning, differentiation medium was added to each well, and 3 days later cells were collected and RNA extraction was performed using a standard Trizol protocol (Thermo Fisher Scientific).
RT-PCR and transcript quantification
Reverse transcription was performed using the protocol provided by the manufacturer with a longer incubation time for the reverse transcriptase process (45 min instead of 30 min) (Maxima K1671, Thermo Fisher Scientific). 1 μg of RNA was reverse transcribed for 10 min at 25°C followed by 45 min at 50°C and the reaction was stopped by heating at 85°C for 5 minutes. PCR (and, in some instances, a second round of nested PCR) was performed using primers pairs and cycling condition (see Table 2). Agarose gels were imaged and band intensities calculated using the gel analysis function of ImageJ software, providing a semi-quantitative analysis of the relative efficiency in generating the skipped transcript. Western blot was performed on protein extracts from all cell lines, but dystrophin protein signal could not reliably be obtained; consistent with prior reports in forced-myotube cultures [7, 8], RNA analysis only is presented.
AON, antisense oligonucleotide. Relative location in exon represents the position relative to the first nucleotide of each exon
AON, antisense oligonucleotide. Relative location in exon represents the position relative to the first nucleotide of each exon
RESULTS
2’O-methyl antisense oligonucleotides (AONs) that target motifs predicted to be involved in splicing-specific exons (Table 1) were transfected into patient-derived cell lines with exon duplications. They were previously tested and optimized in previous published pre-screen performed by the group of Dr. Wilton [11]. In order to test for restoration of a WT transcript, RNA was isolated for RT-PCR analysis of the relevant exon context (Fig. 1); DNA sequencing confirmed the identity and size of all shortened transcript products and established that exon junctions were conserved (Supplemental Figures 1 and 2). Exon skipping efficiency was estimated by densitometry analysis of gel images comparing induced dystrophin mRNAs to the full-length WT transcript after
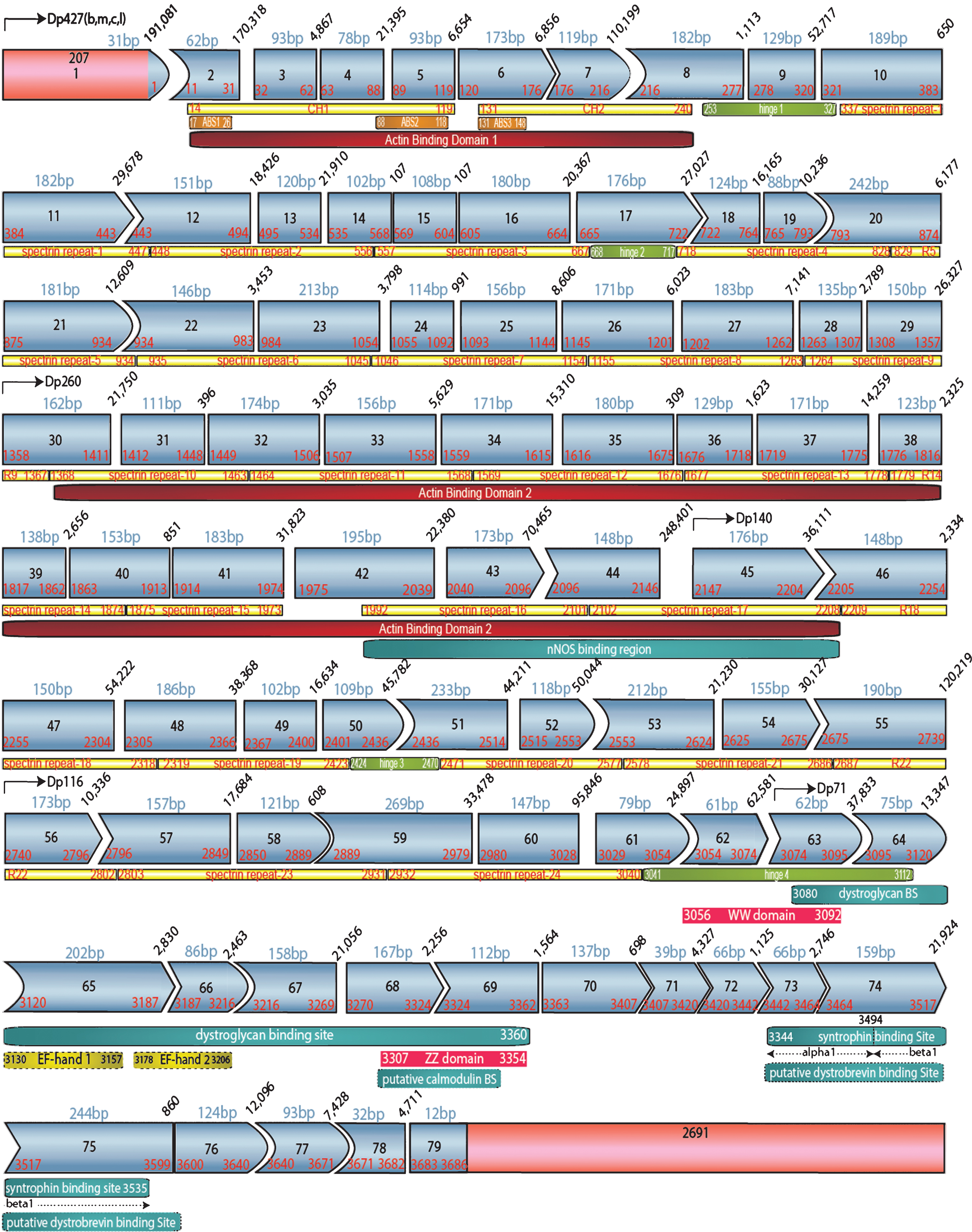
Human
As shown in Fig. 2, in most cases AONs were able to induce skipping of targeted dystrophin exons although the relative efficiencies of skipping varied, presumably reflecting that skipping is more easily induced in some exons than in others. For example, skipping of exon 21 to a WT transcript occurred robustly at AON concentrations of 50 mM, with restoration of 100% WT transcript at the higher dose of 150 mM, without evidence of the complete exon 21 exclusion that would result in a non-therapeutic out-of-frame transcript (Fig. 2a).
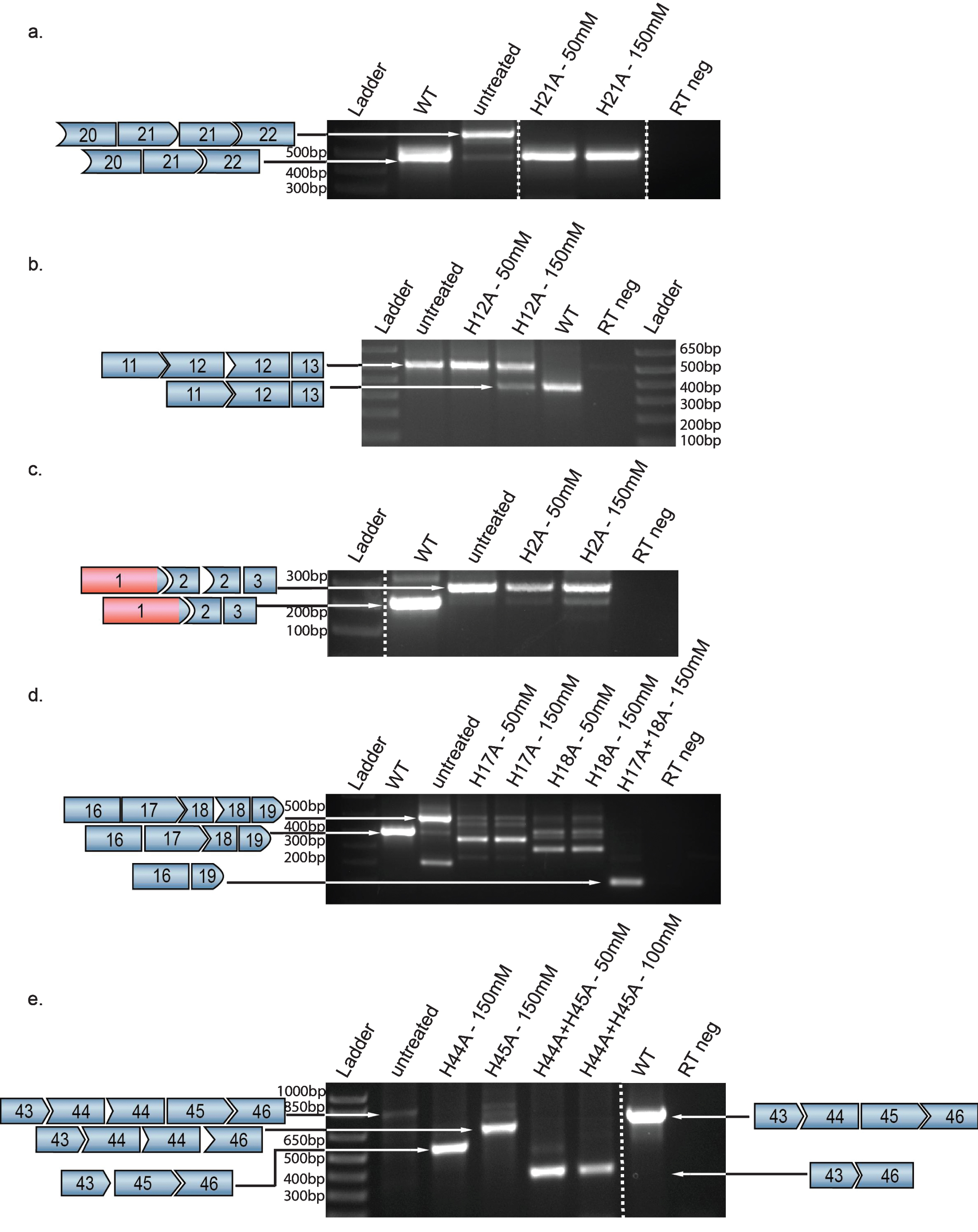
In contrast, skipping of exon 12 shows an evident dose response gradient between 50 mM and 150 mM (Fig. 2b). Of note is the response seen in cells carrying a duplication of exon 2 – the most common single exon duplication in DMD patients [6, 13] – where a dose response is seen but the highest concentration used (150 mM) only resulted in 18% of the total transcript being WT (Fig. 2c).
Very highly efficient skipping that leads to complete exclusion of an exon might prove difficult to titrate therapeutically in boys with DMD. However, results with cells from patients with duplications of exon 18 or exon 44 suggest an alternate approach. In the case of exon 18 duplication, robust skipping leads to an exon 18-deleted (Del18) out-of-frame transcript (70% Del18) even at 50 mM (Fig. 2d). In this case, co-delivery of AONs targeting exons 17 and 18 results an in-frame transcript (Del17-18) as 92% of total transcript. Similarly, in a cell line from a patient with an exon 44 duplication, complete exclusion of exon 44 was robustly induced, leading to a nonfunctional Del44 transcript (81% Del44 at 150 mM; Fig. 2e). Combined delivery with an exon 45-targeted AON (at 50 mM each) results in robust expression of an in-frame Del44-45 in-frame transcript, which cohort studies suggest would be highly protective [14, 15]. Western blot was performed on protein extracts from all cell lines, but dystrophin protein signal could not reliably be obtained. Since it takes usually 10 days of transdifferentiation of wild type fibromyoD to naturally express dystrophin, several round of transfection were performed to sustain efficient skipping. However, this mediated toxicity.
DISCUSSION
With marketing approval of the phosphorodiamidate morpholino oligomer (PMO)
Skipping of only one copy of a single exon duplication might, in principle, be even more beneficial than exon skipping in the setting of deletions, as it could restore a WT gene transcript rather than BMD-like mRNA. The resulting WT dystrophin protein can reasonably be expected to have greater stability than that resulting from treatment of deletion mutations, resulting in a greater accumulation and sustenance of dystrophin protein expression. Exon duplications account for 6–11% of
One theoretical concern is that single exon duplications might be “over-skipped”, resulting in an out-of-frame deletion transcript. This seems unlikely to occur
Beyond this special case, our data demonstrate that for many duplicated exons, single exon skipping is titratable to result in high levels of WT transcript, and suggest that such AON treatment might be personalized based upon the exon involved. Among exons studied here, exon 21 duplication seems to be within the “sweet spot” of the titratable spectrum, with robust skipping of only one copy even at relatively low concentration of AON. In contrast is exon 12, which demonstrates a very faint WT band in the absence of treatment (Fig. 2b), suggesting some native low-level skipping or PCR artifact. However, although a dosage effect can be seen with the portion of WT transcript clearly increasing, the effect is modest (to reach 18% WT).
In contrast to exon 21, treatment with an AON targeting the out-of-frame duplicated exon 18 for excision resulted, even at low levels, in over-skipping and thus another out-of-frame transcript. As demonstrated in Fig. 2d, a potential solution is combination therapy targeting both exons 17 and 18, which efficiently excises both exons, restoring the reading frame in a resulting Del17-18 transcript. Skipping of the duplicated exon 44 was similarly highly efficient even at 50 mM (data not shown) while 150 mM induced overskipping to a deletion 44 transcript (Fig. 2e). Notably, a previous study showed that targeting exon 44 resulted in overskipping whereas the targeting of exon 45 resulted in single exon skipping, suggesting that the efficacy is dependent on the duplicated exon itself, perhaps due to differing dependence between exons on regulatory elements for proper inclusion in the mRNA [7] or reflect significance of the individual break-points. In this case, simultaneous skipping of exon 43 [22] or exon 45 restores the reading frame. We decided to remove both exons 44 and 45 which would likely be highly beneficial, as the paucity of reported patients with a deletion of exons 44–45 (del44-45) suggests an ascertainment bias attributed to a very mild phenotype [14].
The difference between skipping efficiency for all exons may be due to a variety of reasons. Oligomer efficiency can be affected in patients with duplications or mutations that alter pre-mRNA splice motifs or secondary structures [23]. A previous study found that in Dup18 patients, the fact that the donor and acceptor sites were out of context at the duplication junction did not appear to affect oligomer activity [23]. Dystrophin gene mutations can influence the ability of oligomers, designed against normal dystrophin gene transcript, to excise target exons. The specific mechanisms involved remains to be elucidated, and future studies should investigate the effect that duplications of exons and surrounding intronic regions have on pre-mRNA splicing and folding.
A variety of approaches to exon skipping has been in or are approaching clinical trials. In addition to the 2’O-methyl AONs used here, and the PMOs discussed above, another type of compound refered as tricyclo-DNA oligomers (tc-AONs) shows promising results, as systemic delivery promotes a high degree of rescue of dystrophin expression in skeletal muscles, the heart and, to a lesser extent, the brain in the
In summary, we found that AON-induced single exon skipping is quite variable among exons but is in general titratable, and that high levels of WT transcript can be reached with AONs in a cell culture system. In considering human trials, complete skipping of an exonic duplication is highly unlikely to worsen symptoms in a DMD patient, given that the symptoms result from an existing out of frame mRNA that overskipping would also produce. As potential therapeutic products targeting specific mutations are developed, trials in duplicated subjects should be considered.
ACKNOWLEDGMENTS INCLUDING SOURCES OF SUPPORT
This work was supported by internal discretionary funds.
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
