Abstract
The histopathological image classification method, based on deep learning, can be used to assist pathologists in cancer recognition in colon histopathology. The popularization of automatic and accurate histopathological image classification methods in this way is of great significance. However, smaller medical institutions with limited medical resources may lack colon histopathology image training sets with reliable labeled information; thus they may be unable to meet the needs of deep learning for many labeled training samples. Therefore, in this paper, the colon histopathological image set with rich label information from a certain medical institution is taken as the source domain; the colon histopathological image set from a smaller medical institution with limited medical resources is taken as the target domain. Considering the potential differences between histopathological images obtained by different institutions, this paper proposes a classification learning framework, namely unsupervised domain adaptation with local structure preservation for colon histopathological image classification, which can learn an adaptive classifier by performing distribution alignment and preserving intra-domain local structure to predict the labels of the colon histopathological images from institutions with lower medical resources. Extensive experiments demonstrate that the proposed framework shows significant improvement in accuracy and specificity of colon histopathological images without reliable labeled information compared to models without unsupervised domain adaptation. Specifically, in an affiliated hospital in Fuyang City, Anhui Province, the classification accuracy of benign and malignant colon histopathological images reaches 96.21%. The results of comparative experiments also show promising classification performance of our method in comparison with other unsupervised domain adaptation methods.
Keywords
Introduction
Colon cancer is common in malignant tumors across the world [1, 2]. The early diagnosis and treatment of colon cancer is vital to patient health and quality of life. Currently, Pathological examination is the ‘gold standard’ in cancer diagnosis depending on the manual qualitative analysis of professional pathologists, which is time consuming, empirical and prone to misdiagnosis. Computer-aided automatic histopathological image classification is an effective tool to assist pathologists in colon histological image diagnosis. With the development of slice digitization technology and artificial intelligence technology, deep learning methods have been applied to histopathological image analysis [3]. Deep neural networks possess strong ability in automatic feature extraction, complex model construction, and efficient feature expression [4, 5]. Moreover, deep learning methods can extract features of pixel-level original data, level by level, from bottom to top. At present, research on the use of deep learning methods to realize the automatic classification and recognition of colon pathological canceration has attracted extensive attention [6–9].
However, for smaller medical institutions with lower medical resources, there are many challenges to using deep learning methods for classifying and recognizing colon histopathological images. For example, there are smaller colon histopathological image samples and fewer cases for smaller medical institutions. Furthermore, there is a shortage of senior pathological experts in smaller institutions to accurately label samples. Therefore, small medical institutions lack efficient labeled samples to train deep learning models. Although some institutions have disclosed labeled pathological image datasets of colon tissues for training the classification and recognition model of pathological canceration of colon tissue, differential pathological section-making processes, staining schemes and scanning instruments may lead to divergence in the imaging of pathological sections obtained by different institutions. That is, the data distributions are different. When we take the labeled colon histopathological image set from a certain institution and the colon histopathological image set without any labeled information from another institution with limited medical resources as the source colon histopathological images (SCHI) and the target colon histopathological images (TCHI), the classification performance of the deep models directly trained on SCHI will be degraded in TCHI. In other words, although deep neural networks can extract deep abstract features of the colon histopathological images, the performance of the cross-domain classification from SCHI to TCHI is still poor, mainly due to the distribution difference between them. Therefore, it is significant to adapt their distributions and explore useful information in SCHI for improving the classification accuracy of TCHI.
Domain adaptation can deal with cross-domain classification problems by transferring useful knowledge from a source domain to a different but related target domain [10, 11]. When there is no labeled information available in the target domain, corresponding domain adaptation is referred to as unsupervised domain adaptation (UDA), which is more challenging. It is assumed that the target domain
UDA methods contain deep UDA methods and non-deep UDA methods. Deep UDA methods can be directly conducted on the original image data of the source domain and target domain. This kind of methods uses deep neural network to learn the adaptive features. However, it needs to re-train the network and tune many parameters, which is both complex and expensive. Non-deep UDA methods mainly contain feature-based UDA and classifier-based UDA which learn domain adaptation features and directly obtain an adaptive classifier, respectively, by reducing the distribution differences between domains. It should be noted that non-deep UDA methods can be applied to the deep features of the source domain and target domain to improve the cross-domain classification performance [15]. Feature-based UDA includes subspace alignment [16–19], Wasserstein based feature matching [20, 21] and maximum mean difference (MMD) based common feature learning [11, 23]. However, these methods only aim to learn domain adaptation features; they must connect traditional classifiers to complete the classification task. Classifier-based UDA directly trains an adaptive classifier by adding domain adaptation regularization term to a classification model. Dual-level adaptive and discriminative [24] facilitates joint classifier learning in a semi-supervised manner ignoring the geometric structure information in the source and target domains. Adaptation regularization based transfer learning (ARTL) [25] pertained to classifier-based UDA can reduce the difference between domains and improve the discrimination of the classifier by adding inter-domain distribution adaptation regularization term. On this basis, to preserve the internal information of the domains, innovative methods of ARTL are proposed [12, 26]. However, these classifier-based UDA methods maintain the manifold consistency of the source and target domains. Yet the differences between domains may lead to different local manifold structures. Forcibly maintaining the manifold consistency in different domains will weaken the classifier discrimination.
Taking the above challenges into account, we propose an unsupervised domain adaptation with local structure preservation (UDALSP) method. It constructs a structure risk-minimization classification model with distribution adaptation regularization and intra-domain graph regularization to align the distributions and preserve the intra-domain local structure, respectively. The proposed UDALSP pertains to Classifier-based UDA and does not require re-training the network and tuning many parameters like Deep UDA methods, nor does it require connecting traditional classifiers like feature-based UDA. It is derived from ARTL, but distinctly different from ARTL. While reducing the distribution differences between domains, our UDALSP preserves the intra-domain local geometric structure by constructing an intra-domain graph regularization instead of maintaining the manifold consistency of the source and target domains, and learns an adaptive classifier. Adjacent samples following the same distribution usually share the same category; thus preserving the intra-domain local structures of the two domains plays a key role in the classifier discrimination, which can enhance the classifier discriminant. Finally, based on the proposed UDALSP and deep learning, a cross-domain classification framework is constructed for colon histopathological.
The contributions of this paper are as follows: An intra-domain graph regularization is presented. By adding it to the distribution adaptation regularized classifier, the proposed method UDALSP learns an adaptive classifier with intra-domain structure preservation which can enhance the classifier discrimination. UDA is used to address the problem of cross-domain colon histopathological images classification. Specifically, a new classification learning framework is proposed to mine the useful knowledge of SCHI, which is used to solve the classification problem of TCHI. Experiments demonstrate that differences in colonic histopathological images from different institutions make the classification model trained on SCHI performs poor for TCHI. Moreover, the performance of our UDALSP and the proposed classification framework in dealing with the classification of colon pathological images without any label information are verified.
Related works
ResNet [27] is a famous deep convolutional neural network (CNN) architecture which can be pre-trained on the large natural scene image database ImageNet and has been increasingly used to extract the deep features. However, the feature representations of histopathological images directly learned by the pre-trained architecture may be insufficiently discriminative, thus affecting the classification accuracy of histopathological images [28]. Because of the transferable nature of deep CNN parameters and limited labels for retraining the parameters, pre-trained deep CNN with parameter fine-tuning has been widely used in histopathological image classification and recognition [29, 30]. Therefore, the above problems can be addressed through the fine-tuning strategy avoiding training a deep CNN from scratch. Based on pre-trained ResNet and labeled histopathological image training set, the literatures [31, 32] adopt the fine-tuning strategy to quickly and efficiently re-train the ResNet-50, which performs well in histopathological image classification and successfully extracts discriminative deep features of these images. Pre-trained ResNet-50 with parameter fine-tuning has been well used in histopathological image classification and feature extraction [33–35]. In this paper we use Pre-trained ResNet-50 to extract the deep features of SCHI and TCHI to train a classifier. Due to cross-domain discrepancy, the distribution of SCHI and TCHI in the deep feature space is still different, which will result in poor performance of the cross-domain classification from SCHI to TCHI. Therefore, we attempt to use UDA to learn an adaptive classifier in the deep feature space, which can perform well on TCHI.
Distribution alignment is a common strategy for unsupervised domain adaptation. Gretton et al [36] propose MMD as a measurement method for distribution difference, which can be used in marginal distribution alignment. Pan et al [11] propose transfer component analysis (TCA), which embeds the minimization of the inter-domain MMD into a dimensionality reduction framework, to learn low-dimensional shared subspace of two different domains and reduce the marginal distribution difference between them. Tian et al. [37] adopt K-nearest neighbors and MMD to achieve local and global distribution alignment. In order to jointly align the inter-domain marginal and conditional distributions, Long et al. [22] propose joint distribution adaptation (JDA) using another measurement method class-wise MMD (CMMD) to jointly reduce the inter-domain marginal and conditional distribution difference. Li et al. [38] propose discriminative transfer feature learning based on robust-center (DTFLRC), which establishes CMMD with robust-centers to align the source and target domains. These methods aim to find a shared feature representation for the source and target domains by aligning their distributions. However, they cannot remove the difference between domains and still require traditional classification methods to complete the target classification task in the feature space with cross-domain discrepancy. Our UDALSP aligns the inter-domain distribution through a classifier learning procedure by minimizing inter-domain CMMD distance, and directly obtains an adaptive classifier for the target domain.
Local structure preservation aims to preserve the local geometric structure of given data. Due to the fact that adjacent samples following the same distribution usually share the same category, local structure preservation has been considered in many classification and clustering studies. The literature [39, 40] utilizes graph regularization for preserving the local manifold structure during the clustering process. ARTL [25] jointly minimizes graph regularization, structural risk function and inter-domain CMMD distance to preserve the local manifold consistency underlying the whole data in source and target domains and align their distributions, so as to learn an adaptive classifier for cross-domain classification. Based on ARTL, manifold embedded distribution alignment (MDDA) [41] preserves the local manifold consistency in a manifold embedded feature space, where the cross-domain difference still exists. Since adjacent samples following different distributions might belong to different categories, these UDA methods of using graph regularization to preserve the local geometric structure of the whole data ignore the inter-domain distribution difference and will weaken the adaptive classifier discrimination. Different from these methods, our UDALSP uses intra-domain graph regularization to preserve the intra-domain local manifold structure in the source and target domains respectively, thereby enhancing the discriminant structure of the classifier.
Unsupervised domain adaptation with local structure preservation
Notations
We denote the label space of the source and target samples
Main idea
The main idea of UDLSP is to learn an adaptive classifier f for cross-domain classification. UDLSP has two fundamental steps: 1) distribution alignment, which jointly reduces inter-domain marginal and conditional distribution difference by minimizing the inter-domain CMMD distance; and 2) intra-domain local structure preservation, which preserves the intra-domain local manifold structure in the source and target domains, respectively. Integrating the two steps with the principle of structural risk minimization on the source samples, an optimization problem is formulated as
According to the representer theorem [42], f can be written as
Using the square loss function,
where
We use CMMD [22] to estimate the distance between the source and target distributions. By minimizing the distance, the distributions can be aligned so that the generalization performance of the obtained f can be improved on the target domain. Based on the inter-domain distribution distance,
where
Graph Laplacian regularization is often used to preserve the local geometric structures of given samples [25, 43]. On the basis of it, we construct the intra-domain graph regularization to preserve the respective local structures inside the two domains, so as to enhance the discriminant structure of the obtained f.
Let
where,
Let
Integrating formulas (3), (4) and (9), we rewrite the optimization problem in (1) as
UDALSP algorithm is summarized in Algorithm 1.
Given
Deep feature extraction
We use the pre-trained ResNet-50 with fine-tuning to extract the deep features of SCHI and TCHI, as shown in Fig. 1. First, pre-train ResNet-50 on ImageNet to obtain the initial parameters, and then re-train ResNet-50 on SCHI to fine-tune the parameters. Because the ImageNet dataset consists of 1000 categories, while the histopathological image dataset involved in this section consists of only 2 categories, therefore when we fine-tune the parameters, it is necessary to replace the 2048×1000 fully connected layer of the last layer with the 2048×2 fully connected layer.
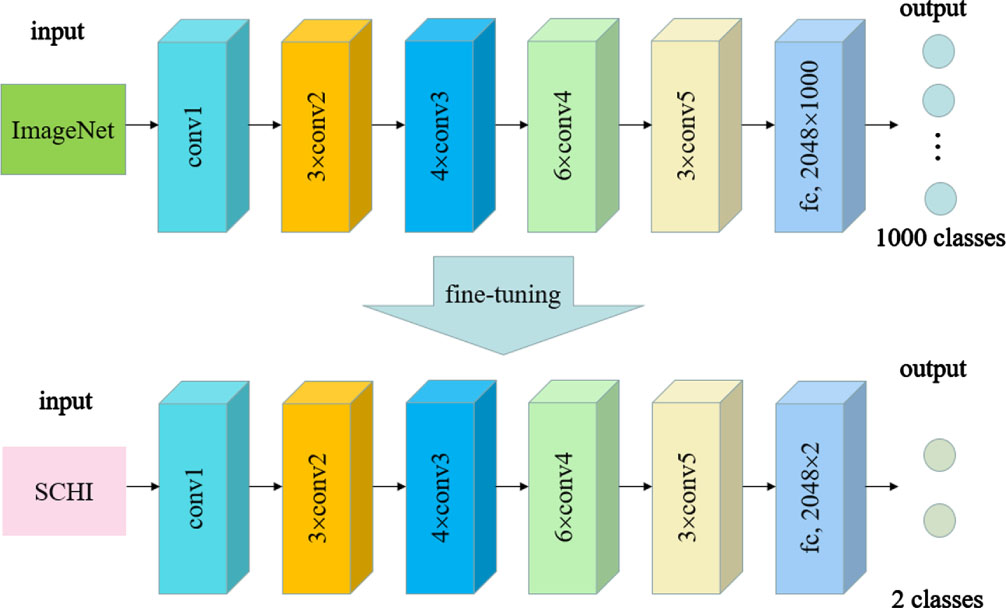
Fine-tuned ResNet-50.
The fine-tuned ResNet-50 can not only realize the classification of histopathological images; it can also learn their 2048-dimensional deep feature space when the last fully connected layer is removed. In view of the difference between TCHI and SCHI, the fine-tuned ResNet-50 trained on SCHI may perform poorly on TCHI. Therefore, in our work, the histopathological images in SCHI and TCHI are input into the fine-tuned ResNet-50. By removing the last fully connected layer, we can obtain the deep features of these images. As the most discriminative fully connected layer is removed, the high-dimensional deep features extracted by the model are more likely to characterize the general properties of colon histopathological images from different institutions.
Cross-domain colon histopathological image classification based on UDALSP goes through two stages, as is shown in Fig. 2. In the first stage, the deep features of SCHI and TCHI are extracted. Colon histopathological images in SCHI and TCHI are scaled to 224×224×3; the pre-trained ResNet-50 parameters are fine-tuned by using the samples in SCHI to obtain the fine-tuned ResNet-50. We then remove the last fully connected layer of the fine-tuned ResNet-50 to obtain a feature extraction model, denoted as Feature-extract. The colon histopathological images in SCHI and TCHI are input into the Feature-extract model to obtain their high-dimensional deep feature representations
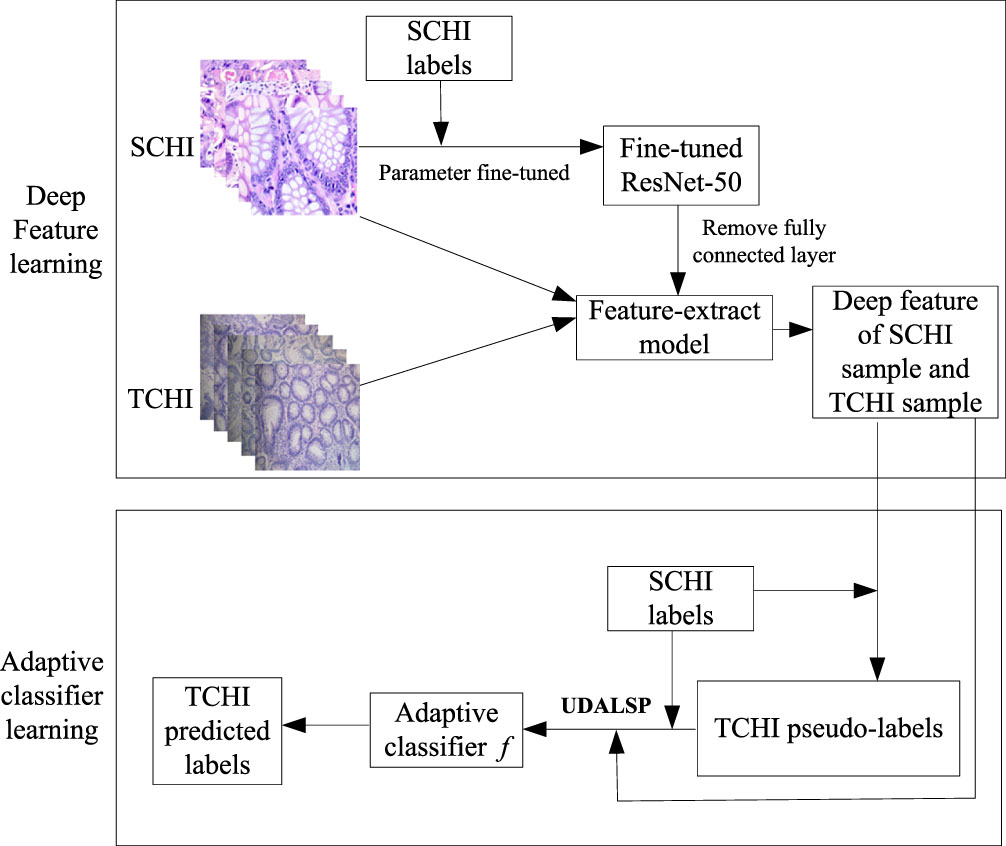
Framework of cross-domain colon histopathological image classification based on UDALSP.
Data set
We collected a small set of colon histopathological images from the pathology department of an affiliated hospital in Fuyang City, Anhui Province as TCHI, which consists of 317 hematoxylin and eosin (H&E) stained colon histopathological images of 2304×1728 pixels, obtained by a Digi Retina 16 HD camera connected to an Olympus BX53 microscope. This study has been approved by the ethics committee of that hospital. In order to verify the performance of the proposed classification frame, three senior pathologists are invited to label these images. Among them, 182 are benign, and the remaining 135 are malignant. Another colon histopathological imageset was taken as SCHI, which was produced through rotating, flipping and cropping the original colon histopathological images from James A. Haley Veterans Hospital in Tampa, Florida obtained by a Leica Microscope MC190 HD camera connected to an Olympus BX41 microscope [44]. The imageset consists of 10000 labeled colon histopathological images of 768×768 pixels, of which 5000 are benign and the remaining 5000 are malignant. The exemplary images in TCHI and SCHI are shown in Fig. 3. We can observe that there are obvious differences in the data distributions of TCHI and SCHI.
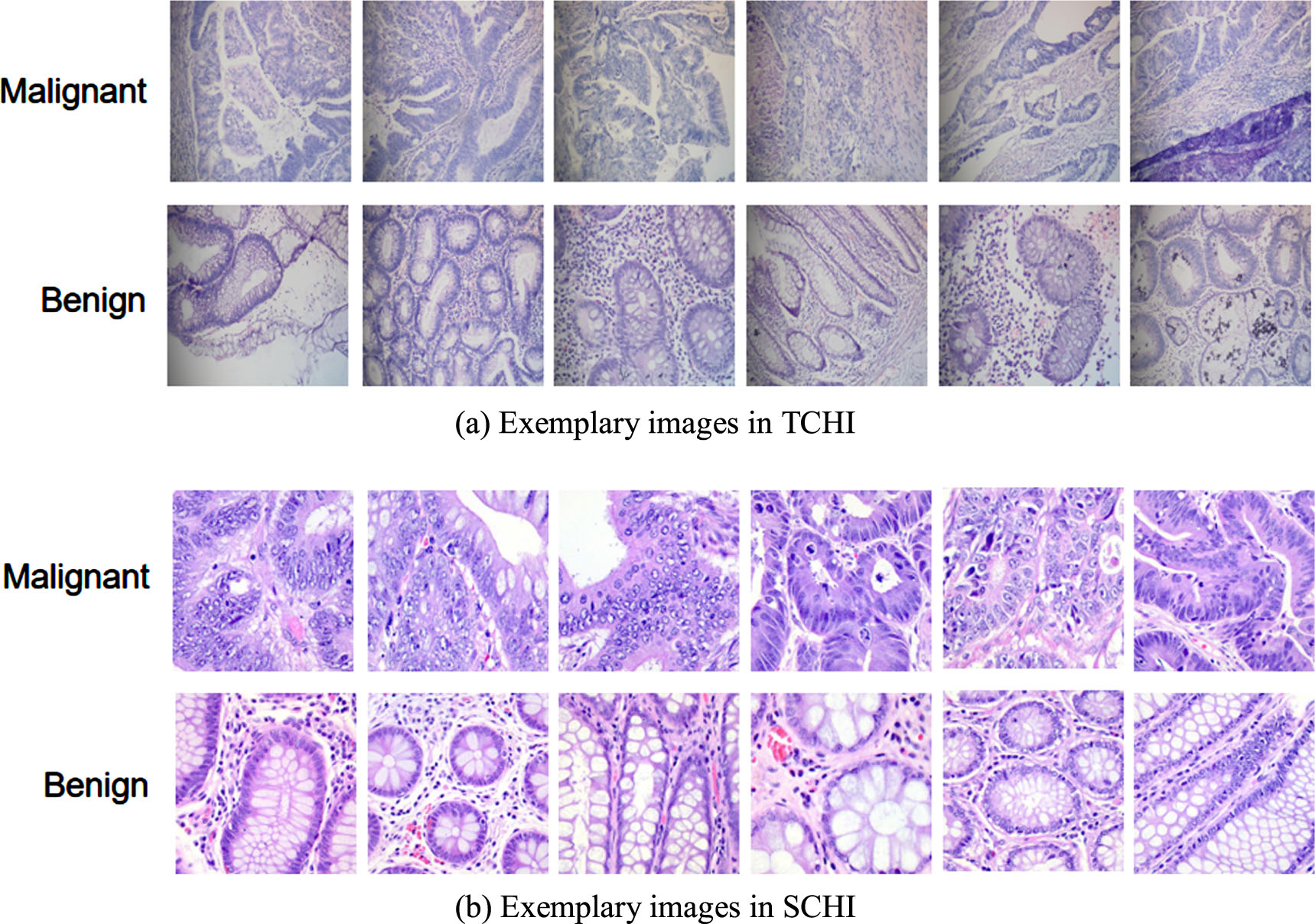
Image samples from TCHI and SCHI.
In the deep feature extraction stage, 80% of the labeled images in SCHI are used to fine-tune the ResNet-50 parameters. To improve the generalization performance of the model, the brightness and contrast of the 80% images are changed randomly, and they are further divided randomly in the ratio of 8:2 to obtain the training set and validation set, respectively. The remaining 20% is used as the test set to test the performance of the fine-tuned ResNet-50 on colon pathological images, in which the number ratio of benign and malignant pathological images is 1:1. The learning rate is set to 0.001, the batch size is set to 32, Epoch is set to 60, the optimizer is set as the stochastic gradient descent with momentum, and the momentum coefficient is set to 0.09. These hyper-parameters are selected using cross-validation. In the stage of adaptive classifier learning, due to the absence of label samples in the target domain, the optimal parameters of the proposed UDALSP cannot be tuned using cross validation. We estimate the optimal parameters using the empirical searching strategy. Since UDALSP can be considered as a variant of ARTL and shares the same type of parameters with it, we keep these parameters consistent with ARTL, that is, λ = 0.1, α = 10, β = 1, p = 10, and the kernel function set as Gaussian kernel function.
As in [11, 38], we take the classification accuracy on the target samples
In medical images classification problems, specificity, and sensitivity are also commonly utilized as evaluation metrics. In addition, the confusion matrix can intuitively evaluate the classification performance on medical images. Therefore, this paper also uses the specificity, sensitivity and confusion matrix of the predictions on TCHI, to evaluate the performance of the proposed classification learning framework on cross-domain colon histopathological image classification. The special calculation formulas are as follows.
Using the proposed UDALSP-based classification framework, the cross-domain colon histopathological image classification task SCHI⟶TCHI is conducted, and the confusion matrix of the predictions on TCHI is shown in Fig. 4 with TN: 171, TP: 134, FN: 1 and FP: 11. Among 182 benign colon histopathological images, 171 are accurately predicted and 11 are misclassified; among 135 malignant histopathological images, 134 are accurately predicted and 1 is misclassified. Overall, 305 of the 317 colon histopathological images in TCHI are accurately predicted, and only 12 are misclassified, with the classification accuracy: 96.21%, specificity: 93.96%, sensitivity: 99.26%.
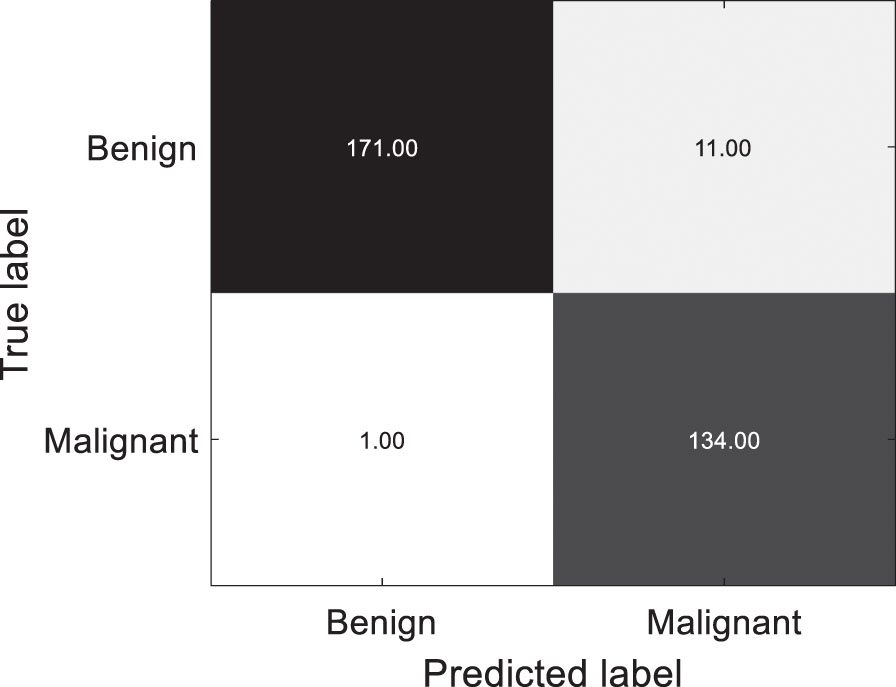
Confusion matrix of the predictions on TCHI.
By analyzing the stability of fine-tuned ResNet-50 and its classification performance in SCHI test set and TCHI, the effectiveness of the proposed classification framework for SCHI⟶TCHI is verified. First, in the process of fine tuning the pre-trained ResNet-50, the classification accuracy curve and cross-entropy loss curve of the training set and the validation set are presented in Fig. 5. Figure 5(a) shows that the classification accuracy of the training set and the validation set stabilizes within 60 epochs, and finally close to 100%; Fig. 5(b) shows that the cross-entropy loss of the training set and the validation set can also be stable within 60 epochs, and finally close to 0. Secondly, the fine-tuned ResNet-50 is used to classify 2000 colon histopathological images in the test set. We find only one image is misclassified; the classification accuracy reaches 99.95%, indicating that the fine-tuned ResNet-50 has good generalization performance on the SCHI. Finally, 317 colon histopathological images in TCHI are directly classified by using the fine-tuned ResNet-50 (without UDA). Among them, 179 of the images are misclassified, the classification accuracy is only 43.53% with specificity: 2.20%, sensitivity: 99.26%. Although the sensitivity is similar to our proposed method, the accuracy and specificity are much smaller, indicating that the generalization performance of the fine-tuned ResNet-50 trained on SCHI is degraded on TCHI. This phenomenon is caused by the difference between SCHI and TCHI. The classification framework based on UDALSP improves the accuracy and specificity by 52.68% and 91.76%, respectively, demonstrating its effectiveness.
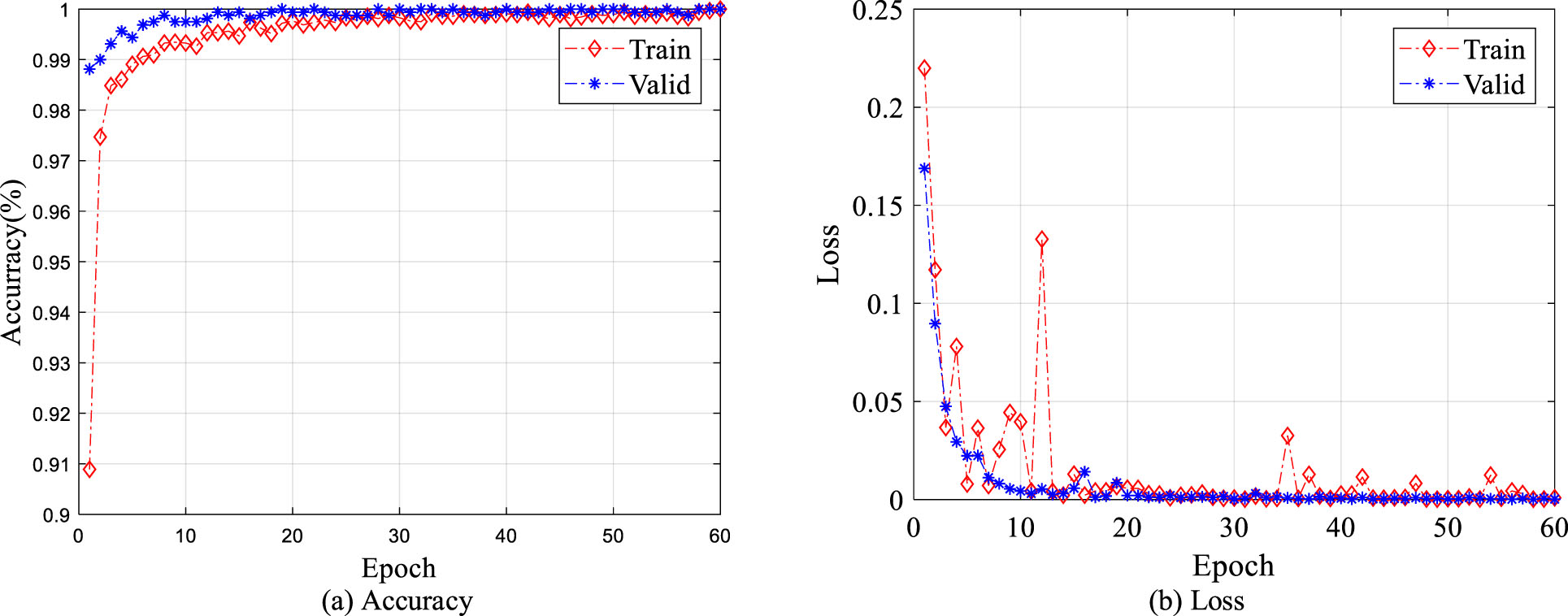
Classification accuracy and cross-entropy loss of training set and validation set.
To further verify the effectiveness of the proposed classification framework, the widely used Macenko method [45] is used to normalize the colon pathological images in SCHI and TCHI. In the process of training the fine-tuned ResNet-50 on SCHI, the classification accuracy curve and cross-entropy loss curve of the training set and validation set are presented in Fig. 6. Figure 6(a) and 6(b) show that the classification accuracy and the loss of the training set and the validation set are still able to converge within 60 epochs. The classification accuracy of the fine-tuned ResNet-50 on the test set reaches 99.90%, indicating that the fine-tuned ResNet-50 continues to show better classification performance on SCHI. However, when using the fine-tuned ResNet-50 to classify the colon histopathological images in TCHI, only 140 of the 317 images are correctly classified, and the classification accuracy is only 44.16% with specificity: 2.75%, sensitivity: 100%. Therefore, stain normalization does not significantly improve the generalization performance of the fine-tuned ResNet-50 on TCHI. The main reason is that although stain normalization can reduce the color difference between pathological images in SCHI and TCHI to a certain extent, it cannot effectively alleviate the differences between histopathological images from different institutions caused by different data preparation procedures, including sectioning, staining, tissue collection.
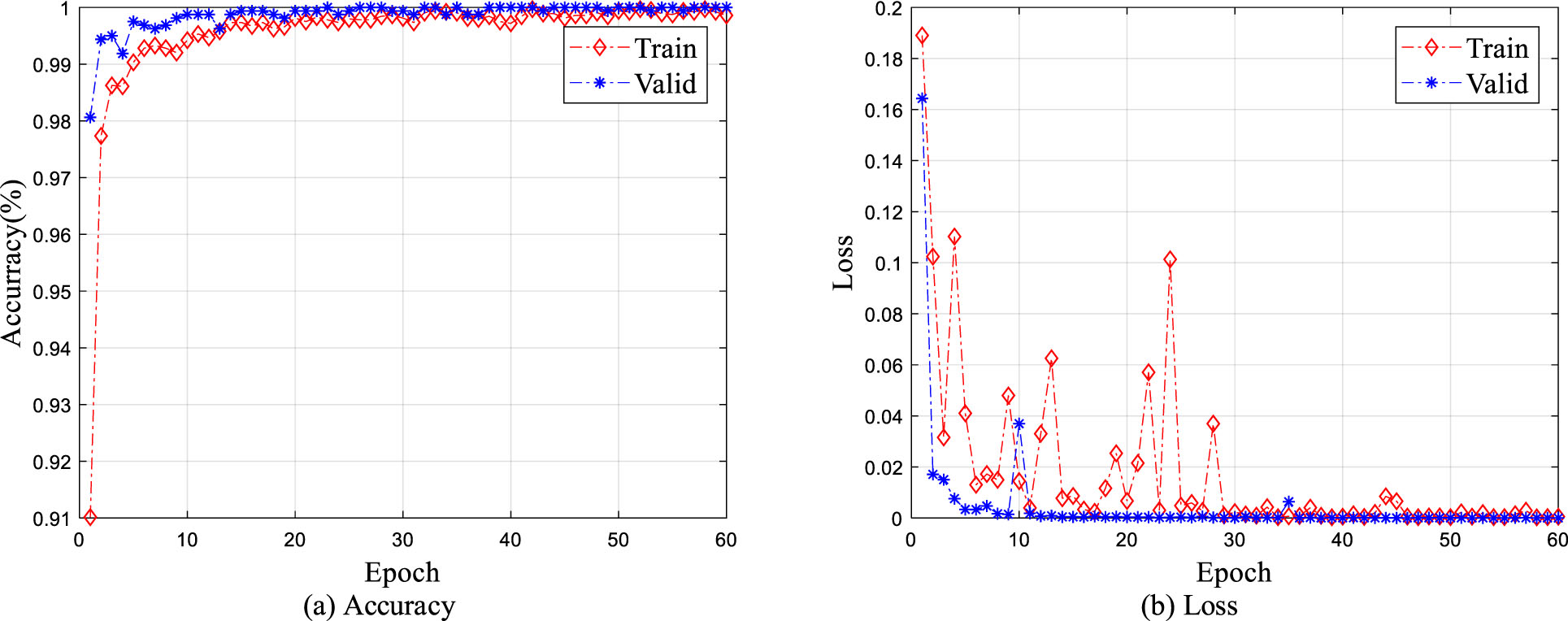
Classification accuracy and cross-entropy loss of stain normalized training set and validation set.
Additionally, we use our classification framework to conduct cross-domain classification task SCHI⟶TCHI with stain normalization. The confusion matrix of the predictions on TCHI is shown in Fig. 7 with TN: 171, TP: 133, FN: 2 and FP: 11. Among 182 benign colon histopathological images, 171 are accurately predicted and 11 are misclassified; Among 135 histopathological images of malignant colons, 133 are accurately predicted and 2 are misclassified. Overall, the accuracy is 95.90% with specificity: 93.96%, sensitivity: 98.52%. The classification framework based on UDALSP improves the accuracy and specificity by 51.74% and 91.21%, respectively, indicating that our method performs well on the cross-domain classification regardless of whether colon pathological images are stain normalized or not.
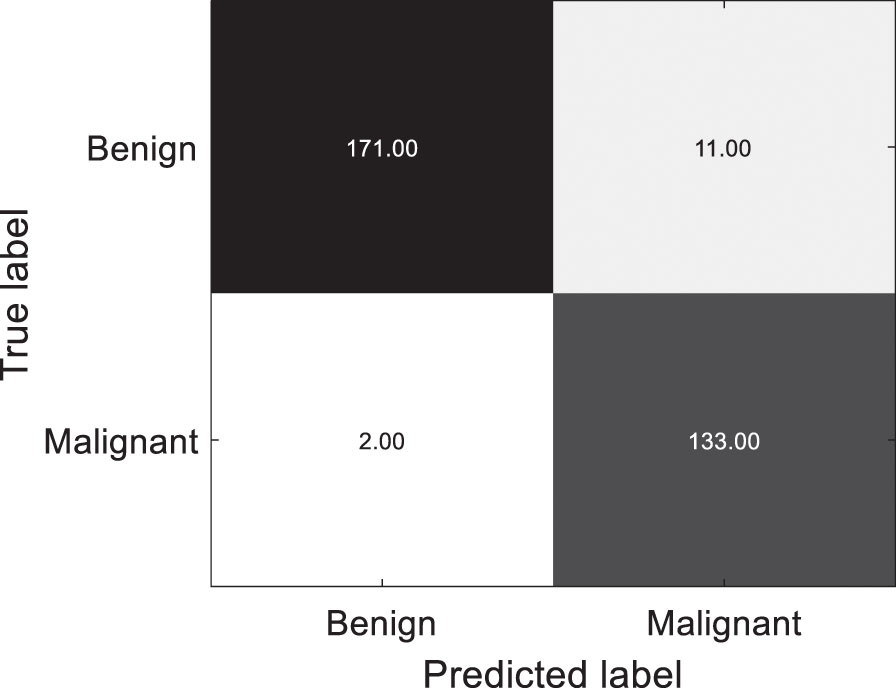
Confusion matrix of predictions on stain normalized TCHI.
We compare the performance of the proposed UDALSP with several state-of-the-art UDA methods on cross-domain colon histopathological image classification from SCHI to TCHI. The comparison methods include eight non-deep UDA methods: TCA [11], JDA [22], geodesic flow kernel (GFK) [19], group-lasso regularized optimal transport (OT-GL) [21], joint geometrical and statistical (JGSA) [46], ARTL [25], manifold embedded distribution alignment (MDDA) [41], probability-based graph embedding cross-domain and class discriminative feature learning framework (PGCD) [47], and four deep UDA methods: deep adaptation network (DAN) [48], joint adaptation network (JAN) [49], conditional domain adversarial network (CDAN) [50], domain conditioned adaptation network (DCAN) [51]. We use the parameters recommended by the original papers for all the compared methods. In view of the instability of deep UDA methods, they are performed 10 times on SCHI⟶TCHI and their mean accuracies are shown in Fig. 8. For the non-deep UDA methods, the experiments are conducted based on the deep features extracted by the fine-tuned ResNet-50 like our proposed classification framework.
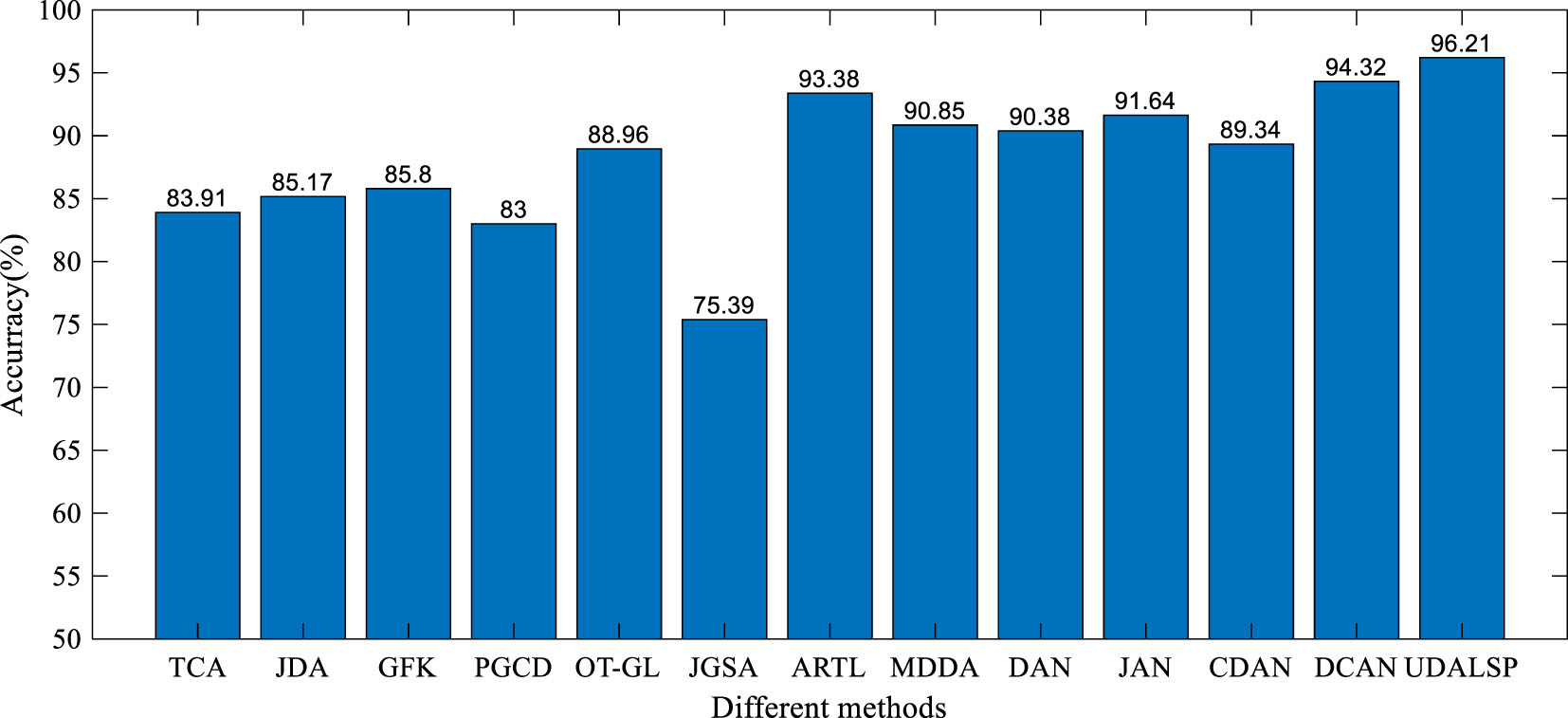
Accuracy of different methods on SCHI⟶ TCH.
As can be seen from Fig. 8, the classification accuracy of UDALSP on SCHI⟶TCHI reaches the highest. TCA and JDA mine the statistical information of SCHI and TCHI to adapt their distributions. GFK, PGCD and OT-GL mine the geometric structure information of SCHI and TCHI to match their features. JGSA aims to align the features of SCHI and TCHI geometrically and statistically. However, these feature-based UDA methods can only reduce the cross-domain discrepancy but cannot remove it, and ultimately require a traditional classifier to complete classification task. ARTL and MDDA directly learn an adaptive classifier, where the distributions of SCHI and TCHI are adapted and the manifold consistency of the two domains is enhanced. The classification accuracy of the two methods on SCHI⟶TCHI is 93.38% and 90.85% respectively, which is higher than most of the feature-based UDA methods, and higher than that of DAN and CDAN. Our UDALSP also directly learns an adaptive classifier, and due to space limitations, we only compare its sensitivity and specificity with the other two classifier-based UDA methods (i.e. ARTL and MDDA). As shown in Fig. 9, there is no significant difference in sensitivity among the 3 methods, but our UDALSP has higher accuracy and specificity than ARTL and MDDA. The main reason is that ARTL and MDDA ignore the difference of local geometric structure between SCHI and TCHI, while UDALSP can preserve the respective local geometric structure of the two domains in the process of classifier learning to enhance the classifier discrimination. The classification accuracy of UDALSP is as high as 96.21%, which is not only higher than the eight non-deep UDA methods, but also higher than the four deep UDA methods.
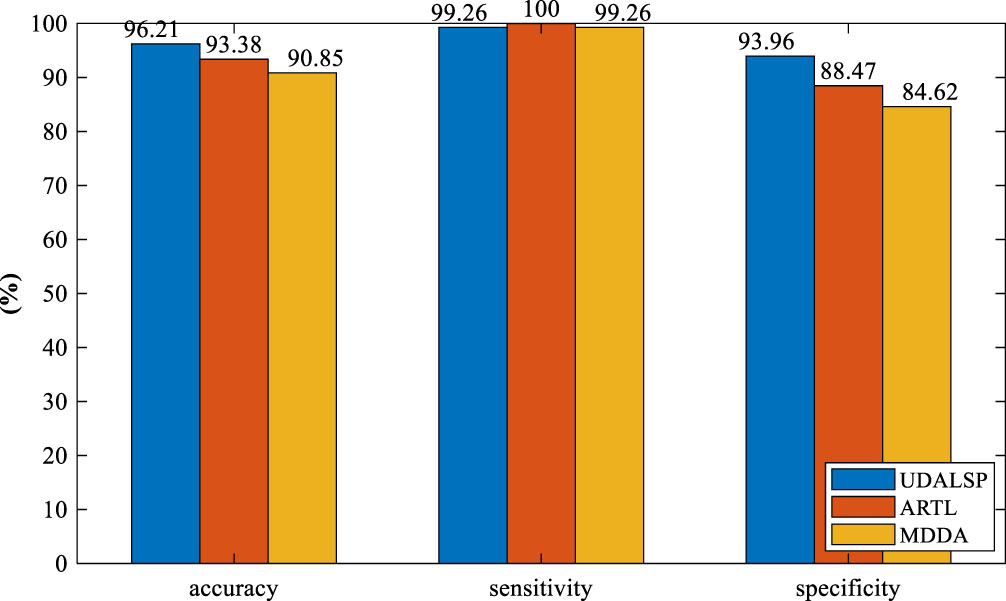
Accuracy, sensitivity, specificity of different methods on SCHI⟶TCHI.
To further verify the superiority of our UDALSP over its peers, we conduct comparative experiments on the benchmark cross-domain dataset ImageCLEF-DA [49], which contains 12 common categories shared by the following three domains: Caltech-256 (C), Pascal VOC 2012 (P) and ImageNet ILSVRC 2012 (I). Accordingly, 6 cross-domain tasks can be constructed, i.e., I⟶P, I⟶C, C⟶P, C⟶I, P⟶C, P⟶I. There are 50 images in each category and 600 images in each domain. For the non-deep UDA methods, including our UDALSP, the experiments are conducted based on the deep features extracted by the fine-tuned ResNet-50 trained on the source domain. Since this dataset is not a medical image set, we only use the classification accuracy on it to evaluate UDALSP and the compared methods on the 6 cross-domain tasks I⟶P, I⟶C, C⟶P, C⟶I, P⟶C, P⟶I. It can be observed in Table 1 that our UDALSP outperform the feature-based UDA methods (i.e. TCA, JDA, GFK, OT-GL, JGSA and PGCD) on most tasks (4/6). This is because the feature-based UDA methods only learn a latent shared feature space to reduce cross-domain difference, and these methods also need to train a traditional classifier in the feature space where the difference still exists. In addition, UDALSP preserving the intra-domain local geometric structure performs better than ARTL and MDDA on most tasks (4/6). The main reason is that the classifier-based UDA methods (i.e. ARTL and MDDA) maintain the manifold consistency of the source and target domains ignoring the inter-domain distribution difference, which will degrade the performance of the adaptive classifier. However, for the tasks I⟶P and C⟶P, UDALSP is beaten by PGCD, MDDA and ARTL as reported in Table 1. A possible explanation is that the deep features of the target samples extracted by the fine-tuned ResNet-50 have weak discriminability and the intra-domain structure preservation of our UDALSP has a negative impact. Nevertheless, UDALSP provides best results on most tasks and achieves the highest average accuracy of 89.2%, 1.4% higher than the deep UDA methods.
Accuracy (%) on ImageCLEF-DA
We also evaluate the parameter sensitivity of UDALSP with respect to p, α, β and λ. The accuracy of UDALSP with varying values of nearest neighbors p is presented in Fig. 10(a), which indicates a wide range p ∈ [4, 50] for optimal parameter values. From Fig. 10(b), we observe that α with too-high values undermines model performance, and the wide range α ∈ [100, 104] is the best choice. Figure 10(c) shows the accuracy of UDALSP with different values of β, indicating that UDALSP can achieve a robust performance with regard to a wide range β ∈ [0.01, 5] and the best choice is β ∈ [0.1, 5]. For the parameter λ, a wide range of λ ∈ [10-3, 1] is the best as presented in Fig. 10 (d).
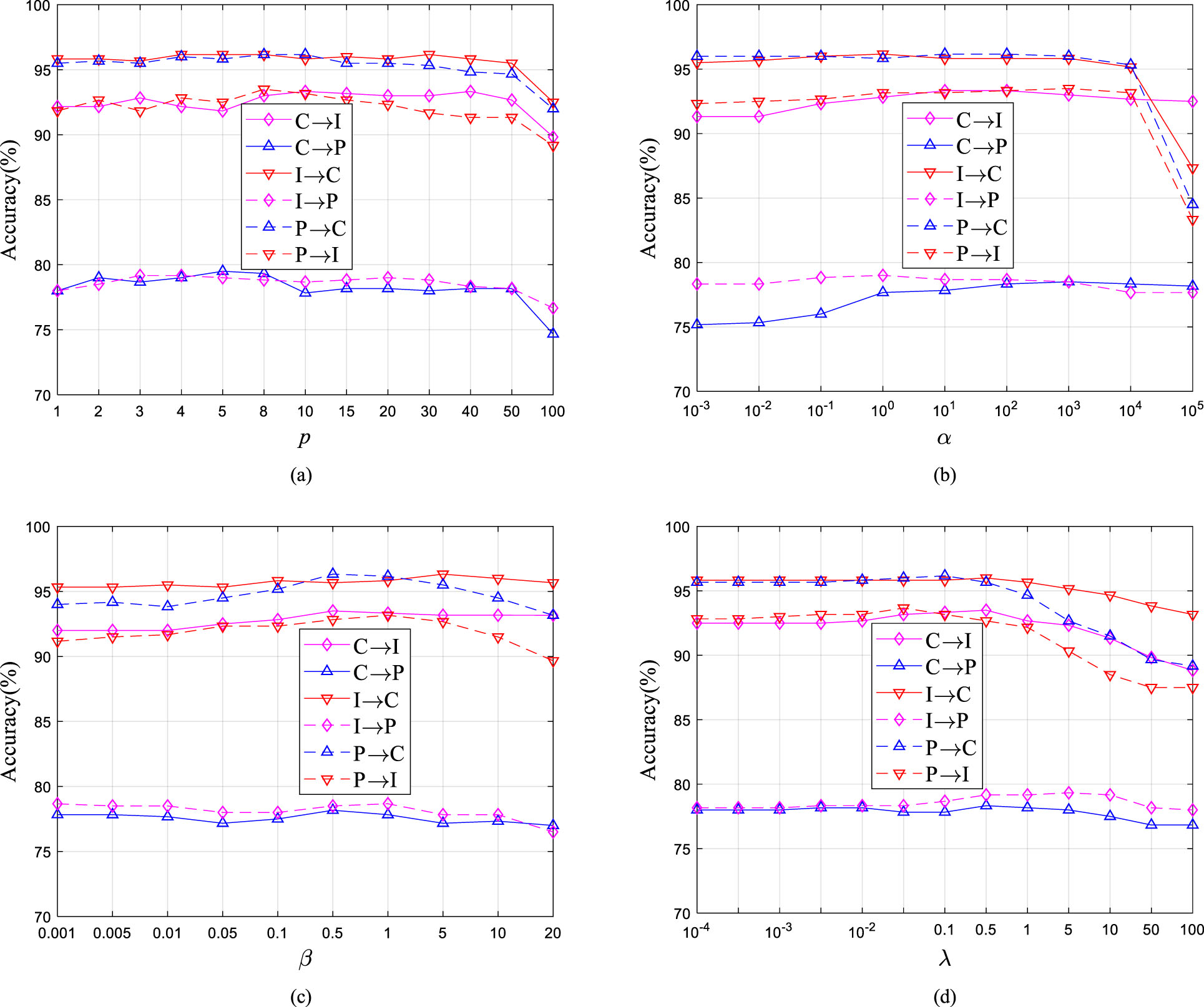
Parameter sensitivity of UDALSP, (a)-(d) shows the classification accuracy with regard to p, α, β, λ.
This paper proposes a classification learning framework based on UDALSP for cross-domain histopathological image classification from SCHI (i.e., source domain) to TCHI (i.e., target domain), which is significant for medical institutions to assist their pathologists in colon histopathological image diagnosis, especially for smaller medical institutions with lower medical resources. First, the fine-tuned ResNet-50 is trained on SCHI, to extract the deep features of the colon histopathological images from SCHI and TCHI. Then, UDALSP is used to adapt the two domains, and learns an adaptive classifier for the target domain to predict the labels in the target domain. Comprehensive experiments are conducted on a cross-domain colon histopathological image classification task. The results show that the proposed classification framework based on UDALSP performs well on cross-domain colon histopathological image classification with accuracy and specificity improvement of 52.68% and 91.76%, respectively. In addition, the proposed UDALSP is compared with the present state-of-the-art UDA methods and it can outperform these methods on both colon histopathological image set and the benchmark UDA dataset, which demonstrate the superiority of UDALSP with intra-domain local structure preservation.
However, UDALSP cannot deal with negative transfer incurred in poorly discriminative feature spaces where adjacent samples following the same distribution might belong to different categories. In the future, we intend to devise a more general UDA model to compensate for the challenge and extend our work to domain generalization problem for histopathological image classification.
Footnotes
Acknowledgments
The work was supported in part by the National Nature Science Foundation of China (No. 71901001), the Major Special Science and Technology Project of Anhui Province, China (No. 201903a05020020), the Key Research Project of Natural Science in Colleges and Universities of Anhui Province, China (Nos. KJ2021A1253, KJ2021A1251), and the Startup fund for doctoral scientific research, Fuyang Normal University, China (No. 2022KYQD0010).
