Abstract
Huntington’s disease (HD) (OMIM 143100) is caused by an expanded CAG repeat tract in the HTT gene. The inherited CAG length is known to expand further in somatic and germline cells in HD subjects. Age at onset of the disease is inversely correlated with the inherited CAG length, but is further modulated by a series of genetic modifiers which are most likely to act on the CAG repeat in
Keywords
BACKGROUND
Huntington’s disease (HD) is one of > 50 diseases caused by expanded short tandem repeats [1, 2]. In those diseases where the repeat is coding, as in HD, the repeat unit is usually CAG and this is translated to a homopolymeric glutamine tract in the encoded protein. There are nine such diseases, often referred to collectively as the polyglutamine diseases. The sections of these proteins containing expanded glutamine form cellular aggregates [3]. The polyglutamine diseases have disease-causing expansion lengths that are much shorter than those in diseases where the repeats causing the expansion are not translated [4, 5], implying a possible constraint on length at the level of the protein. While somatic expansion is critical in reaching the intracellular pathogenic CAG length threshold, the subsequent events leading to cell dysfunction and death have not been conclusively defined (Fig. 1). Much attention has focused on the expanded glutamine tract in the protein but it has never been conclusively proven that this elicits toxicity in cells in human disease, and the genetic evidence implicates CAG length rather than polyglutamine length as critical in HD pathogenesis [6–8]. Other potential pathogenic mechanisms that cannot be precluded include RNA-based toxicity as in myotonic dystrophy (OMIM 160900) [9], RAN translation [10] and aberrant exon 1 splicing [11]: all of these mechanisms would also be exacerbated by somatic expansion of the repeat in individual cells. Recent evidence of neurodevelopmental effects in HD [12], and early phenotypes in peripheral blood mononuclear cells [13, 14], may indicate other pathways impacted by the unexpanded CAG length, but the genetic evidence in HD subjects very clearly points to somatic expansion as likely to be important in disease manifestation.
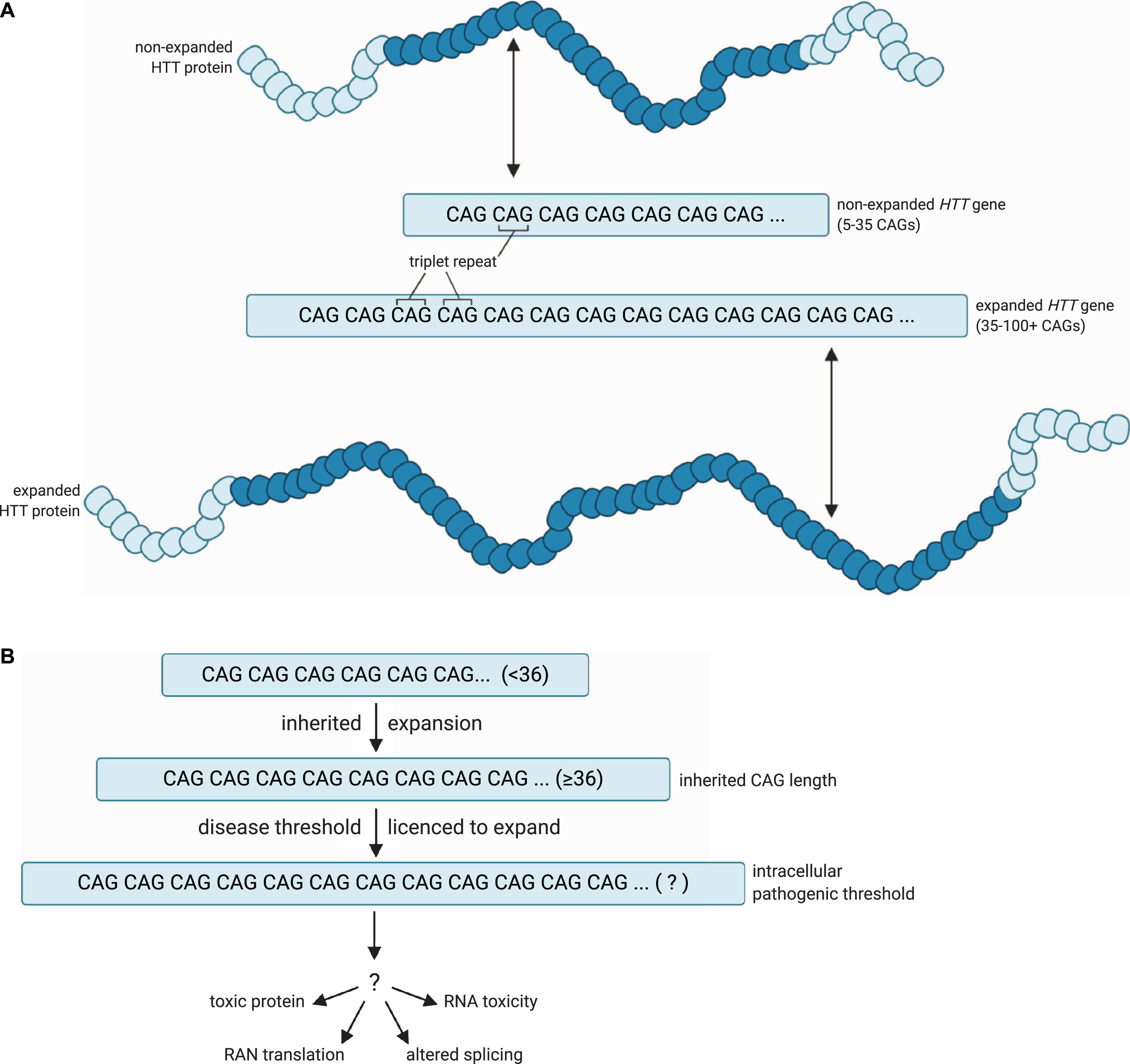
A model for the pathogenic threshold in HD. A) HD pathogenesis is largely determined by an expanded cytosine-adenine-guanine (CAG) trinucleotide repeat within exon 1 of the huntingtin (
In HD, age at onset of disease is largely determined by the length of the CAG tract [15–17]. More recently, however, age at onset has been shown to be modulated by a series of genetic modifiers whose discovery has revolutionised the way we think about HD pathogenesis [7, 18– 20]. Two types of genetic modifier revealed so far have provided evidence that has made us rethink our notions of HD pathogenesis. First, some of the encoded proteins translated from these modifiers act directly on DNA and are most likely to exert their effect at the level of the mutated expanded DNA, through modulating the length of the CAG tract in both somatic and germline cells [18–21], and indeed, this has been shown experimentally in cell culture [22, 23]. Second, the exact sequence at the
A TWO-STAGE HYPOTHESIS OF DISEASE PATHOGENESIS
We know that further expansion of the
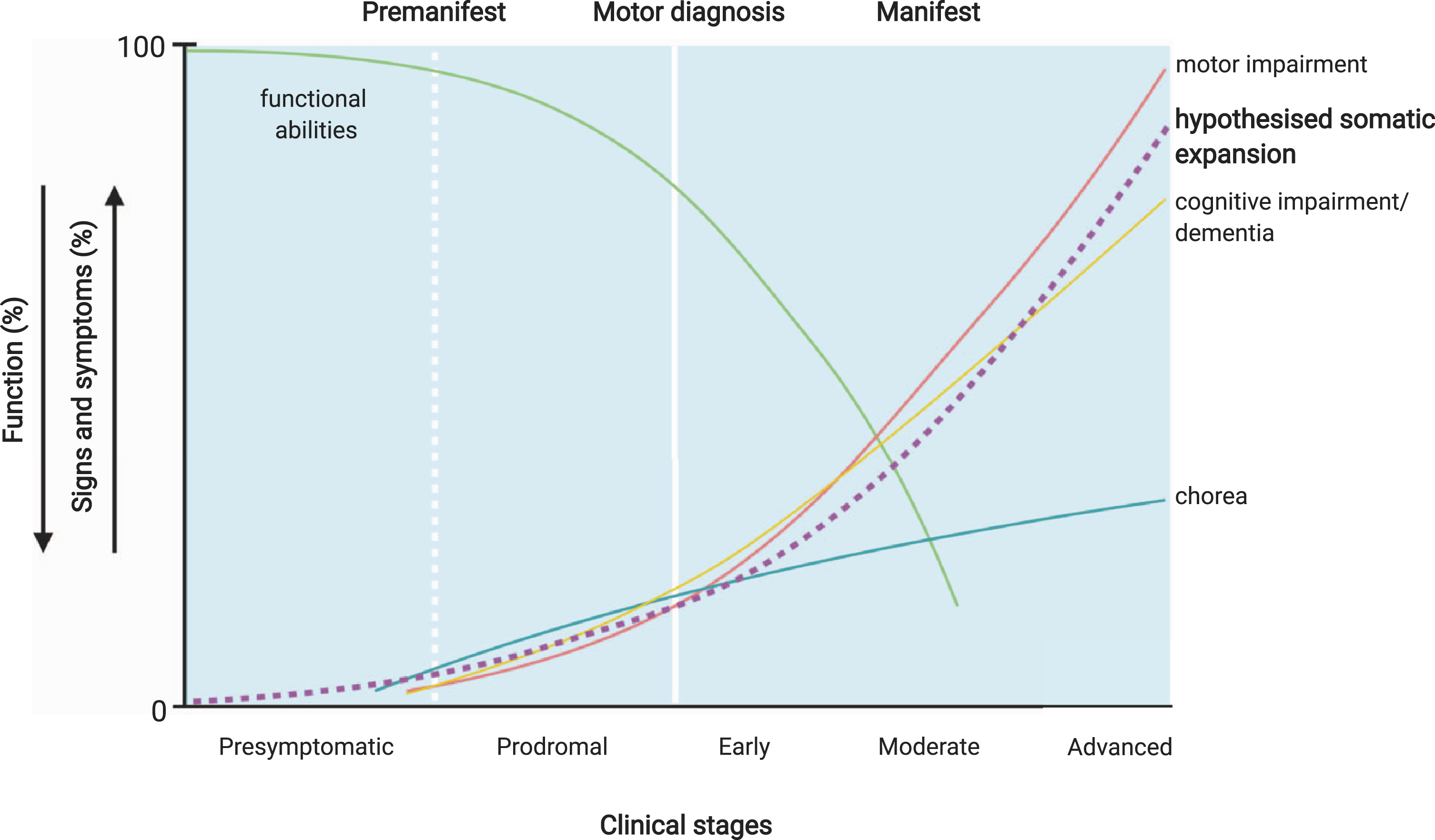
Potential relationship of CAG tract expansion and clinical Huntington’s disease events. The premanifest period of the disease may reflect the presence of a proportion of disease-relevant cells with sufficient somatic expansion to induce neuronal dysfunction, but too few to manifest overt clinical symptoms. Premanifest HD includes a presymptomatic period where no signs or symptoms are present, and prodromal HD, characterised by the onset of subtle signs and symptoms, which may be the result of the
The
There are some potential clues to the intracellular pathogenic threshold. We might be able to improve the definition of the edge of the pathogenic thresholds using data from mouse models. In mouse models the repeat is normally expanded to 100 CAG or more in order to induce a disease-like phenotype in the short-lived mouse [46, 47]. Even in the presence of tracts of over 100 CAG in their
EVIDENCE FROM HD ANIMAL MODELS
There are many animal models of HD generated in a number of different ways (Table 1). They can be divided into those expressing transgenes with a truncated section of human
Animal models of Huntington’s disease with up to 100 CAG repeats
NSE, neuron-specific enolase; CMV, cytomegalovirus; PrP, prion gene promoter. Not reported means no data were available. No means somatic expansion was investigated and not seen.
Although there are multiple rodent models which have been deployed to help us understand the biology of HD and begin the search for therapies, many are limited in their ability to inform us of the effects of genetic modifiers of disease, as they often present with repeats well above the presumed intracellular pathogenic threshold and a severe phenotype. The most useful are those with relatively short repeats (Table 1) though they have differences in their genetic manipulations that make straightforward inferences about the threshold for intracellular pathogenesis complex. They all have either full length human
An added complication is that very long repeats appear less pathogenic than shorter disease-causing repeats, and more prone to contraction than expansion [88], though it is not clear why [89, 90]. The earliest onset and most deleterious phenotypes are seen around 150 CAG with longer CAG tracts giving later phenotypic changes [89–91] though it should be noted that in mice with an inherited ∼150 CAG there is also somatic expansion and the repeat length in the susceptible cells is likely to be longer than 150 CAG. Very long repeat tracts form unusual DNA structures [2] that can inhibit transcription or translation of
A number of models, still encoding glutamine but using a mixed CAACAG rather than a pure CAG tract, can help to establish a window for a pathogenic repeat length. The mixed CAACAG stabilises the repeat tract [78], preventing germline and somatic expansion.
The BAC HD model with 97 glutamines encoded by a mixed CAACAG tract fulfils this criteria— the mixed CAACAG tract prevents both germline and somatic expansion in mice but is still pathogenic (Table 1) [78]. These mice have 5 copies of the transgene integrated into their genome and express the BAC HD
Given the BAC HD line with a stable tract of 97 glutamine-encoding codons has a phenotype [78] this sets an upper bound to the likely intracellular pathogenic length (Table 1). The HdhQ92 mouse with a human exon 1 pure CAG tract knocked into mouse
There are additional limitations in extrapolation from mice and other animal models to people [101]. Expression levels of the gene and protein are not necessarily at endogenous levels. Genetically the most accurate animal models are those with long CAGs knocked into their mouse
While both people with HD and the animal models of disease have development of phenotypic changes over time, animals do not have an age at onset of manifest disease, as at clinical diagnosis in humans. In both people and models the changes seen depend on what phenotypes are examined and how they are measured [38, 105]. The differences in disease manifestation in people are not reflected in mice, because laboratory mice are much less genetically diverse and live in a more uniform environment. Genetic variation in HD subjects influences the presentation of many non-motor symptoms for instance [106]. Most HD mouse models, despite possession of a repeat length that would give juvenile HD with its different clinical presentation, show a similar motor phenotype (though this may be an artefact of how this is measured) (Table 1) [47]. They also display very little frank neurodegeneration, though they often have smaller and lighter brains than their wildtype counterparts [46]. A series of matched knock-in lines with identical glutamine encoding stretches in
EVIDENCE FROM OTHER DISEASES
Repeat sequences are common in the genome and biologically functional [107] and there is a growing list of diseases caused by expanded repeated sequences in DNA [1, 108]. A series of neurodegenerative diseases are caused by expanded CAG sections in their coding sequence, invariably translated to a polyglutamine tract [109]. These diseases have some striking similarities: the repeat threshold at which disease is caused is in most cases a similar length [4, 48], they show a strong relationship of repeat length with age at onset of disease, many show somatic and germline expansion of their causative repeat [31, 110– 112] and they have similar genetic modifiers of their ages at onset [113] (Table 2). This implies that the underlying events leading to expansion of the CAG tracts in these diseases might have common mechanisms that can be used to inform all of these diseases, though the molecular pathogenic events downstream of the CAG tract may be specific to each disease.
Evidence from human CAG-repeat disorders
Table 2 shows the diseases caused by expanded CAG tracts where the repeat is definitely or likely to be translated to a polyglutamine tract in the cognate protein— it is perhaps of relevance that most of the polyglutamine protein products have a role in DNA repair [33, 114]. Only spinocerebellar ataxia 6 (SCA6, OMIM 183086) shows no evidence of somatic expansion of the CAG tract, though there is genetic anticipation in families, implicating germline expansion [115–117]. SCA6 may therefore be an exception, not requiring intracellular somatic expansion to elicit pathogenesis. The CAG tract disease range is shorter than in the other diseases, and the repeat occurs in
SCA1 disease-causing expanded CAG tracts are 39 CAGs or more with no interruption, or 45– 81 with interruptions. Lack of interruptions gives earlier disease onset [135] and in uninterrupted alleles there is a strong length correlation with age at onset [127]. The interruptions are CAT, encoding histidine rather than glutamine, and the later onset of disease was assumed to be mediated by the resulting change in the protein [126], but it appears more likely to be mediated at the level of DNA by the somatic expansion widely seen in this disease [123, 179]. The pathology of SCA1 is concentrated in the cerebellum with a characteristic early and severe degeneration of the Purkinje cells [4] although recent evidence shows that subjects have widespread degeneration in deep cerebellar structures and the brainstem as well as cerebral pathology [180]. In postmortem SCA1 human brain, the highest levels of somatic expansion are not seen in the cerebellar regions and the Purkinje cells most affected in the disease [129], though at the end stage of disease the earliest affected cells may have been lost. Additionally, Purkinje cells are low in number compared with other cerebellar neurons [181], and thus rare, large expansions in these cells are likely to be underestimated when looking at whole cerebellar tissue. However, elegant work in mice has shown that it is likely to be protein interactions, particularly with capicua, that drive cell-specific intracellular pathogenesis in the Purkinje cells [182, 183]. Nevertheless, somatic expansion may drive other pathogenic events in SCA1: a similar genetic modification signal was seen in SCA1 as in HD, implying that age at onset is at least partly modulated by similar events in both diseases [113].
SCA2 is more complicated. Most CAG tract alleles have CAA interruptions, but may also be interrupted by CCG, encoding glycine. Pure CAG tracts over 34 CAG cause the ataxic phenotype of SCA2 [5, 145], but interrupted alleles in what would normally be considered the long normal or low SCA2 range (see Table 2), give a Parkinsonian or amyotrophic lateral sclerosis phenotype [138, 184]. No evidence of somatic expansion has been seen in the phenotypes associated with interruptions [149, 150] but it is seen in SCA2 [140].
SCA3 is perhaps the most interesting and informative of the SCAs with respect to the CAG length pathogenic threshold. Normal alleles may have repeat lengths up to 44 CAG, whereas disease-associated alleles range from 52– 75 CAG, with most disease alleles harbouring repeat lengths of over 60 CAG (Table 2) [48]. There is a window where no repeat lengths have been reported between the normal and disease ranges in SCA3 as in DRPLA, SCA12 and SCA17. The CAG tract is usually interrupted by two CAAs and there does not seem to be an association between the presence of interruptions and phenotype. Notably the somatic mosaicism observed is of the order of a few repeats even in the presence of CAG tracts of 70– 80, and expansions are more prevalent in peripheral tissues than in nervous tissue [153, 154]. Though these analyses are in relatively few brains and do not use techniques that would reveal individual large expansions, nevertheless this appears to be a more stable CAG repeat tract than in HD or SCA1 for instance, especially given the CAG tract length. This provides a repeat tract length for neurodegeneration of a minimum of 60 CAG in SCA3.
SCA17 is caused by an expanded mixed CAA/CAG tract in
There are limitations to extrapolating from other diseases. They have different pathologies and different susceptible cell types. Notably in most of these diseases regional pathology and somatic expansion are not correlated, but relatively few subjects have been analysed in anatomical detail and only one study conducted at the single cell level. This study, measuring somatic expansion in single cells in DRPLA, compared somatic mosaicism in cerebellar structures in early versus late onset patients [173]. Higher rates of expansion were more evident in late onset case than early onset cases, though this may well be a function of age [88, 187]. The frequency of expansions was highest in glial cells, with Purkinje cells lower and granular cells lower again. Relative levels of expression of the cognate genes in the most susceptible cells are not known, but are assumed to underlie differential spatial pathogenesis [121, 188], and transcription appears to be important in promoting somatic repeat tract length changes [85– 87, 189– 192]. Finally, surviving cells that are examined in post-mortem human brain may be resistant to the ongoing toxicity mechanisms and therefore uninformative about the intracellular pathogenic repeat length threshold.
MOUSE MODELS IN OTHER REPEAT DISORDERS
There are multiple mouse models of each non-HD polyglutamine repeat disorder, most of which have not had somatic expansion of the repeat surveyed systematically (Table 3). Most alleles were cloned from patients as transgenes or knocked into the endogenous mouse genes, and often required longer CAG repeat lengths than in humans to evoke a phenotype. As in HD animal models, transgenic mouse models of these diseases often demonstrate severe early-onset neuropathology and behavioural syndromes whilst knock-in mouse models tend to show milder late-onset phenotypes that perhaps parallel the disease more accurately, but are slower to produce phenotypes. Consistent with animal models of HD, animal models of other triplet repeat disorders tend to show increased disease phenotype as CAG repeat length increases, though this is influenced by the promoter used, transgene copy number and resultant transgene expression. Cemal et al. [193] generated a series of eight YAC SCA3 models and found that disease severity increased both with an increased CAG repeat tract length and an increased transgene copy number such that an animal with 72 CAG and one copy of the transgene developed symptoms later than an animal with 67 CAG repeats and two copies of the transgene.
Animal models in other repeat disorders
FL, full length;
In some cases, allelic series have been ‘naturally’ generated through intergenerational expansions or contractions following extensive breeding [196– 198, 225]. These models allow us to explore the effect of CAG repeat length in a well-controlled system. One such system is a series of transgenic DRPLA mouse models carrying 76, 96, 113 and 129 CAG, whose motor deficits and cognition worsen with CAG repeat length and age. High levels of somatic expansion were observed in the cortex, liver and kidney of the Q76 mice [196], and although no behavioural phenotype was initially reported in the Q76 after 64 weeks, they showed reduced survival and body weight when compared with non-transgenic littermates [197] as well as neuronal intranuclear accumulation [196]. Again, repeat instability is likely to occur in all models, but was only examined in Q76 animals.
Genomic context is an important driver of repeat instability in these models. Early studies of independent transgenic mouse models of SBMA with 45
Marked repeat instability has been observed in a transgenic mouse model of SCA3, CMVMJD94, which carries 94 CAG repeats [225]. Expansion was observed in multiple tissues, but within the brain mosaicism was most notable in the pontine nuclei, substantia nigra and striatum. Somatic instability correlated well with neuronal atrophy and gliosis in the pontine nuclei and substantia nigra, but pathological involvement was not seen in the striatum [225]. Another mouse model of SCA3, Ki91, and a mouse model of SCA1, Sca1154Q/2Q, also demonstrate similar tissue-specific patterns of repeat expansions, with notable expansions in the striatum [207, 224]. This extends to other repeats— the same tissue distribution of expansion is seen in models of myotonic dystrophy [187]. These data suggest that whilst repeat instability is not associated with cerebellar neuronal vulnerability in models of SCA, it is likely that repeat instability in areas other than the cerebellum might contribute to disease pathogenesis [259]. Intergenerational instability has been observed in numerous models of SCA3 despite the interrupted CAG tract; this could be due to the presence of a long uninterrupted stretch of CAG at the 3’ end of the tract [222, 227]. These findings suggest that somatic instability is occurring in these model systems.
Some models have allowed us to examine the substantial effect that inheriting only 1 or 2 CAG additional repeats may have on phenotype [225]. SCA3 mice with 83 CAG repeats did not demonstrate behavioural differences, yet SCA3 mice with 94 CAG repeats and similar expression levels demonstrated rotarod deficits and behavioural abnormalities from 16 weeks. It was concluded that the threshold for disease in this model was between 84– 94 CAG repeats. Analysis of data from two cohorts of Q94 also revealed an inverse correlation between the length of the CAG repeat tract and the time spent on the rotarod [225].
Whilst animal models have been invaluable in examining pathogenesis in these diseases, as in HD models, to date, it has been difficult to show directly that somatic expansions are causative to neuronal dysfunction, earlier age at onset and faster disease progression. Interpretation of results is difficult when repeat sequence and length are not clearly defined or have not been examined. Many of the issues that arise in the HD animal models also arise in animal models of other repeat disorders and for many of the same reasons. However, the conclusion from human CAG repeat disorders, and also the corresponding mouse models, would indicate that a repeat length of less than 100 CAGs is toxic to cells— at the shorter end of that estimated by Kaplan et al. [34]. Exactly where the intracellular pathogenic threshold falls remains unclear, but the evidence would place it at over 60 CAG. The question remains whether it is possible to define the intracellular pathogenic threshold more accurately.
WHAT EVIDENCE DO WE NEED TO REFINE OUR DEFINITION OF THE INTRACELLULAR PATHOGENIC THRESHOLD?
The parameters used to establish the CAG-length threshold for HD pathogenesis by Kaplan et al. [34] included the CAG size threshold for disease to arise, the subject’s inherited repeat length as measured in blood, and their current age: these data are available. However, they also require a measure of the cell group critical portion— of the most susceptible cell population(s), what proportion have died, or are dysfunctional, at onset of clinical disease? The final unknown, for HD and the other repeat diseases, is the basal expansion rate of the repeat over time. In HD, the cell group critical portion can be estimated from previous work that showed around half of the most susceptible D2R-expressing medium spiny neurons in the striatum have been lost at onset [260–262]. This parameter could likely be estimated in living subjects from imaging data, as recent well-standardised structural imaging and clinical data has been collected in prospective studies in both manifest and premanifest subjects [38, 263].
The basal expansion rate of the repeat in the most susceptible cells is much more difficult to measure or to derive from existing data. Given the likely stochastic nature of the allele expansion process and the data available in human brain which indicates very long repeats in some cells [24], this will be hard to estimate. However, the very long repeats could be rare events and indeed, could be protective in those surviving cells, as such repeat lengths are seen to reduce phenotype severity and delay onset in mice [90]. The most useful data are likely to come from single cell approaches in a combination of human and mouse brain. It would be ideal if all the data we needed could be derived from human brain, but this is unlikely to be sufficient as human postmortem brain is at the end stage of disease, and the only cells that can be surveyed are those that have survived. These are likely not representative of those that died earlier, and they may well themselves have been dysfunctional at death. Nevertheless, given this is likely a stochastic process there might be surviving cells at different points in the pathological trajectory that could be used in single cell experiments to define the pathogenic CAG tract length threshold. There are methods to sequence and size the
Mouse brain is likely to offer a clearer picture of the dynamics of the pathological process, as tissues can be taken across the lifespan of the mouse and can be processed immediately to generate high quality single cell data. One major disadvantage of most HD mouse models is that they show little frank neurodegeneration, and in this respect do not recapitulate the human disease, but rather display neuronal dysfunction. However, for some analyses this is an advantage. Current data indicates that HD cellular dysfunction can be measured by single cell RNA-seq [266–268], though the disconnect between behavioural and gene expression changes observed by Landles et al. [96] may make this difficult to interpret. The barrier here is gaining a measure of
Using the age at onset genetic modifier data obtain-ed in people might help to establish the pathogenic threshold. The effect sizes and directions of the known modifiers can be used to construct a polygenic risk score, which here consists of the sum of all known modifier alleles, weighted by the effect of each allele on onset [272]. This score can be used to predict somatic expansion in individuals without requiring expansion to be measured directly, thereby greatly increasing sample size, and may be incorporated into the Kaplan model [34]. This assumes that age at onset is a surrogate for measuring somatic expansion: Ciosi et al. [6] showed that individuals with higher blood DNA
Given the recent interest in targeting somatic expansion of the expanded CAG in
CONFLICT OF INTEREST
LJ is a member of the Scientific Advisory Boards of LoQus23 Therapeutics and Triplet Therapeutics. JD, SP, NR and PH have no conflicts of interest.
