Abstract
Background:
Although Lumipulse assays and conventional ELISA are strongly correlated, the precise relationship between their measured values remains undetermined.
Objective:
To determine the relationship between Lumipulse and ELISA measurement values.
Methods:
Patients who underwent cerebrospinal fluid (CSF) Alzheimer’s disease (AD) biomarker measurements and consented to biobanking between December 2021 and June 2023 were included. The relationship between values measured via Lumipulse assays and conventional ELISA were evaluated by Passing-Bablok analyses for amyloid-β 1-42 (Aβ42), total tau (t-tau), and phospho-tau 181 (p-tau 181). Studies using both assays were systematically searched for in PubMed and summarized after quality assessment.
Results:
Regression line slopes and intercepts were 1.41 (1.23 to 1.60) and –77.8 (–198.4 to 44.5) for Aβ42, 0.94 (0.88 to 1.01) and 98.2 (76.9 to 114.4) for t-tau, and 1.60 (1.43 to 1.75) and –21.1 (–26.9 to –15.6) for p-tau181. Spearman’s correlation coefficients were 0.90, 0.95, and 0.95 for Aβ42, t-tau, and p-tau181, respectively. We identified 13 other studies that included 2,117 patients in total. Aβ42 slope varied among studies, suggesting inter-lab difference of ELISA. The slope and intercept of t-tau were approximately 1 and 0, respectively, suggesting small proportional and systematic differences. Conversely, the p-tau181 slope was significantly higher than 1, distributed between 1.5–2 in most studies, with intercepts significantly lower than 0, suggesting proportional and systematic differences.
Conclusions:
We characterized different relationship between measurement values for each biomarker, which may be useful for understanding the differences in CSF biomarker measurement values on different platforms and for future global harmonization.
Keywords
INTRODUCTION
Alzheimer’s disease (AD), a leading cause of dementia, is characterized by amyloid-β plaques and phosphorylated tau tangles in the brain. Clinical diagnosis is not sufficiently specific [1] and biological confirmation via biomarkers is important [2, 3]. Measurement of cerebrospinal fluid (CSF) biomarkers is one of the main approaches for biological confirmation of the brain pathology [4–8] and various studies have confirmed that measurement of CSF Aβ 1-42 (Aβ42), total tau (t-tau), and tau phosphorylated at threonine 181 (p-tau 181) are useful for AD diagnosis [9]. Previous studies have predominantly used conventional enzyme-linked immunosorbent assays (ELISA) such as INNOTEST (Fujirebio Europe N.V., Ghent, Belgium) to measure Aβ42, t-tau, and p-tau 181, whereas Aβ42/40 ratio is currently recommended over Aβ42 itself [10, 11]. Widespread usage of CSF AD biomarkers is expected until blood-based biomarkers become available in clinical practice, for implementing disease-modifying therapies. However, comparing results measured by different laboratories or analytical platforms is difficult and efforts towards global standardization and quality control are ongoing worldwide [12, 13].
Fully automated immunoassays, including those using the LUMIPULSE system (FUJIREBIO INC., Tokyo, Japan), are advantageous because they reduce inter and intra-laboratory variations via automation [14]. These assays are now receiving approvals from local authorities worldwide and replacing conventional ELISA. Although strong correlations have been repeatedly reported between Lumipulse assays and conventional ELISA [15–25], their measurement values do not seem to be necessarily interchangeable, hampering the seamless understanding of previous and future AD biomarker studies. The relationship between these measurement values could be affected by differences in methodology between institutions/studies, or by the fundamental nature of assay characteristics.
The aim of this study was to elucidate the relationship between CSF AD biomarker values measured using Lumipulse assays and conventional ELISA.
MATERIALS AND METHODS
Participants and settings
All patients who underwent lumbar puncture for CSF biomarker measurement at the Tokyo Metropolitan Institute for Geriatrics and Gerontology between December 2021 and June 2023 were recruited for the Tokyo Medical Biobank. Those who consented to biobanking with sufficient remaining CSF samples were subjected to measurements using both Lumipulse assays and conventional ELISA as quality controls. The measured data from all patients with measurement results from both assays were retrospectively reviewed. This study was approved by the Institutional Review Board of the Tokyo Metropolitan Institute for Geriatrics and Gerontology. The remaining CSF, serum, and plasma samples were stored as part of the Tokyo Medical Biobank for future study.
Sample collection and storage
CSF samples were obtained using a standard lumbar puncture procedure. The first tube was sent for cell counting and routine biochemical testing. Subsequent CSF samples were directly collected in polypropylene low-binding tubes after confirming the absence of visible blood contamination, as recommended [26].
For the measurement of Aβ42, t-tau, and p-tau 181 using Lumipulse assays, CSF was directly collected into 2.5 mL sterile polypropylene low-binding tubes (False bottom tube CSF, Sarstedt AG & Ci. KG, Nümbrecht, Germany, Cat# 63.614.625) and stored at –80°C until measurement. The storage time before biomarker analysis was within eight months. Before amendment of the biobank protocol, CSF samples of 19 patients were directly collected into polypropylene low-binding tubes and aliquoted into 1.5 mL tubes (Proteosave SS 1.5 mL, Sumitomo Bakelite Co., Ltd., Tokyo, Japan, Cat# MS-4202X) and stored at –80°C until measurement. The storage time before biomarker analysis was within eight months. The measurement results of these samples using different handling methods were visually plotted and carefully evaluated to determine the possibility of bias before inclusion in the main analyses.
For the measurement of CSF p-tau 181 via ELISA, CSF was directly collected into an autoclaved 4 mL polypropylene tube (Serum tube, Sumitomo Bakelite Co., Ltd., Cat# MS-4604) as instructed, stored at –20 to –30°C, and collected the same day by a clinical laboratory company—LSI Medience Cooperation (Tokyo, Japan). The storage time before biomarker analysis was within two months.
For the measurement of Aβ42 and t-tau via ELISA, CSF was directly collected into polypropylene low-binding tubes and aliquoted into 1.5 mL tubes (Proteosave SS 1.5 mL, Sumitomo Bakelite Co., Ltd.) and stored at –80°C until measurement. The storage time before biomarker analysis was within 16 months.
Lumipulse assays
Measurements of CSF Aβ42, p-tau 181, and t-tau were conducted at FUJIREBIO INC. using LUMIPULSE G1200 and immunoreaction cartridges/calibration sets: β-amyloid 1-42 (Cat# 230336/260258), pTau 181 (Cat# 230350/260227), and Total Tau (Cat# 230312/260203). The intra-assay coefficient of variation (CV) for Aβ42, p-tau 181, and t-tau were 1.8–3.1, 0.8–3.3, and 2.7–4.4%, respectively, as stated by the manufacturer. Cut-off values for p-tau 181 and t-tau were predetermined at 56.5 and 404 pg/mL, respectively.
ELISA
The measurement of CSF p-tau 181 via ELISA was conducted at Tokiwa Chemical Industries Co., Ltd. (Tokyo, Japan) using the Finoscholar ELISA kit (Nipro Corp., Osaka, Japan), which is identical to the INNOTEST (Fujirebio Europe N.V.) assay [27]. CV was < 10%. The lower limit of quantification and reporting was 25 pg/ml. The cutoff value was set at 50 pg/mL.
The measurements of CSF Aβ42 and t-tau were conducted at Tokyo Metropolitan Institute for Geriatrics and Gerontology using INNOTEST β-amyloid (1-42) (Fujirebio Europe N.V.) and Finoscholar hTau (Nipro Corp.) performed as per manufacturer’s instructions. Intra-laboratory CV for CSF Aβ42 was calculated using the duplicated measurement results of run validation control (RVC) on three different days. The intra-laboratory CV was 7.1% using RVC2 (low concentration) and 10.8% using RVC1 (high concentration). Institutional cutoff values were predetermined at 500 and 300 pg/mL based on previous results [8, 29].
Statistical analysis
Statistical analyses were conducted using MedCalc Statistical Software version 20.218 (MedCalc Software Ltd., Ostend, Belgium; https://www.medcalc.org; 2023). GraphPad Prism version 9 (GraphPad Software, San Diego, CA, USA) was used to generate graphs in the Supplementary Material. Samples with biomarker concentrations below the lower quantification limit were plotted and are provided in the scatter plot but were excluded from the main analyses. Missing data were addressed using a pairwise deletion approach. Categorical variables were presented as percentages and continuous variables were presented as mean±SD after confirming normal distribution. The differences in CSF biomarker measurement values between Lumipulse assays and conventional ELISA were visualized using Bland-Altman difference plots [30]. Regression lines for CSF concentration of Aβ42, t-tau, and p-tau 181 measured via Lumipulse assays and conventional ELISA were evaluated using the Passing-Bablok analysis, which is a robust non-parametric regression analysis assuming error in both x- and y-axis [31] suitable and recommended for method comparison studies in a situation when normal distribution is difficult to assume [32, 33]. Perpendicular residuals were used for the calculations. The median and 95% confidence intervals (95% CI) of the slope and intercept of the regression equation for each biomarker are provided. With sufficient sample size and small residuals, 95% CI of the slope including 1 and 95% CI of the intercept including 0 suggest a non-significant bias [31]. Residuals from the regression lines were also plotted in rank order, and the 95% CI are depicted in the figures. Although not necessarily recommended for assessing the agreement between the two analytical methods, Spearman’s correlation coefficients were also reported with 95% CI for comparison with previous studies.
Sample size estimate
A sufficient sample size for the Passing-Bablok analysis has been reported to be 30–90 when the relevant difference is defined as 10%, the measurement range is large, and sampling is uniform [34]. The Clinical and Laboratory Standards Institute (CLSI) guidelines for comparison of measurement procedures and bias estimation suggest at least 100 for manufacturing validation studies and 40 for those conducted in medical laboratories [32]. An interim analysis was conducted with a sample size of 66 and the sample size was increased to over 100 for p-tau181 based on the observed results.
Systematic review
A systematic review was conducted in compliance with the Preferred Reporting Items for Systematic Reviews and Meta-analysis (PRISMA) 2020 statement [35]. The protocol was registered with PROSPERO before formal screening (CRD 42023406903). Existing literature on the relationship between CSF AD biomarkers between Lumipulse assays and ELISA was screened from the PubMed database from 2017 (when the first Lumipulse AD biomarker assay β-amyloid 1-42 was released) to 2023; searched on April 6, 2023. The terms used were as follows: LUMIPULSE, CLEIA, “Chemiluminescent enzyme immunoassay,” Chemiluminescent AND Alzheimer, Fully-automated AND Alzheimer, and Biomarker AND Commutability. The inclusion criteria for the study were as follows: measurement of CSF AD biomarkers in the same samples using both Lumipulse assays and ELISA. The exclusion criteria were as follows:<10 cases and inability to extract data related to regression or correlation characteristics. Two authors (M.K. and S.K.) independently conducted the literature search and any disagreements were resolved through discussion. After selecting articles based on these criteria, their reference lists were also carefully assessed, and those meeting the criteria were included in the study.
The quality assessment of each study was conducted based on a quality assessment tool specific to biomarker measurement procedure comparison studies (QUABICS) (described in the Supplementary Material), which was modified from a quality assessment tool for cross-sectional studies of biomarker data (BIOCROSS) [36]. The quality of each study was assessed “as a study assessing the relationship between biomarker values measured in two assays” and not the study itself. Many studies provided biomarker correlation or regression data as a part of publication with different major objectives. Information on all studies is provided in tables, and only studies with moderate or high quality based on quality assessment scores were included in the forest plots and meta-analyses.
Primary information extracted from the literature included patient characteristics, measurement conditions, statistical methods, slopes and intercepts of the regression line, and correlation coefficients. The primary outcome measures were the slope and intercept of the regression line. The additional outcome measure was the correlation coefficient. Meta-analyses were conducted using a random-effects model. For the meta-analysis of slopes and intercepts, only studies reporting effective confidence intervals were included and analyzed using STATA/MP 18 (StataCorp LLC, TX, USA). For the meta-analysis of correlation coefficients, studies reporting mean values and sample sizes were included and analyzed using MedCalc version 20.218. Funnel plots were created to assess the possibility of publication bias.
RESULTS
Demographic characteristics of the participants
During this period, 166 patients who underwent lumbar puncture were recruited, and 138 consented for the Tokyo Medical Biobank. Sufficient CSF was unavailable in three patients because of technical difficulties. The remaining 135 patients were included in the study (recruitment rate: 81.3%). The demographic characteristics and availability of biomarker results of the participants are summarized in Table 1. Patients with frontotemporal lobar degeneration (FTLD) included those clinically diagnosed with frontotemporal dementia (FTD, n = 7), corticobasal syndrome (CBS, n = 9), or progressive supranuclear palsy (PSP, n = 3). Patients with other diseases included those clinically diagnosed with multiple system atrophy (n = 4), hydrocephalus (n = 4), amyotrophic lateral sclerosis (n = 2), and neuronal intranuclear inclusion disease (NIID, n = 2).
Demographic characteristics and biomarker result availability of the 135 participants
Demographic characteristics and biomarker result availability of the 135 participants
AD, Alzheimer’s disease; DLB, dementia with Lewy bodies; FTLD, frontotemporal lobar degeneration; MCI, mild cognitive impairment; PD, Parkinson’s disease; n/a, not available.
Relationship between Lumipulse and conventional ELISA measurement values for each biomarker
Based on the scatter plots and Bland-Altman difference plots (Supplementary Figure 1) we decided to include all measured samples, although the results of Aβ42 need to be interpreted with caution for the effect of tube difference. Spearman’s correlation coefficient was 0.90 [95% CI: 0.84 to 0.94] for Aβ42, 0.95 [95% CI: 0.93 to 0.97] for t-tau, and 0.95 [95% CI: 0.93 to 0.97] for p-tau181. Using predetermined cutoffs, the binary results matched between the two assays at 95% for t-tau and 92% for p-tau181. Bland-Altman difference plot showed that the value of Aβ42 and p-tau181 was lower in ELISA in the higher range (Supplementary Figure 1D, F) suggesting proportional differences.
Passing-Bablok regression lines and residual plots are provided in Fig. 1A–F. The Cusum test showed no significant deviation from linearity. The regression line slope for Aβ42 was higher than 1 (1.41 [95% CI: 1.23 to 1.60]), and the 95% CI of intercept included 0 (–77.8 [95% CI: –198.4 to 44.5]) indicating a proportional difference towards lower values measured via ELISA (Fig. 1A, G). The subgroup analysis results differentiating the sampling procedures are shown in Supplementary Figure 2. The 95% CI of the regression line slope for t-tau included 1 (0.94 [95% CI: 0.88 to 1.01]) and the intercept was slightly higher than 0 (98.2 [95% CI: 76.9 to 114.4]), indicating a small systemic difference towards lower values measured via ELISA (Fig. 1B, G). The regression line slope for p-tau181 was significantly higher than 1 (1.60 [95% CI: 1.43 to 1.75]) and the intercept was significantly lower than 0 (–21.1 [95% CI: –26.9 to –15.6]) indicating a proportional and systematic difference (Fig. 1C, G). While the difference plot for Aβ42 and t-tau did not show outliers, two outlier residual values for p-tau181 were observed (Fig. 1D–F).
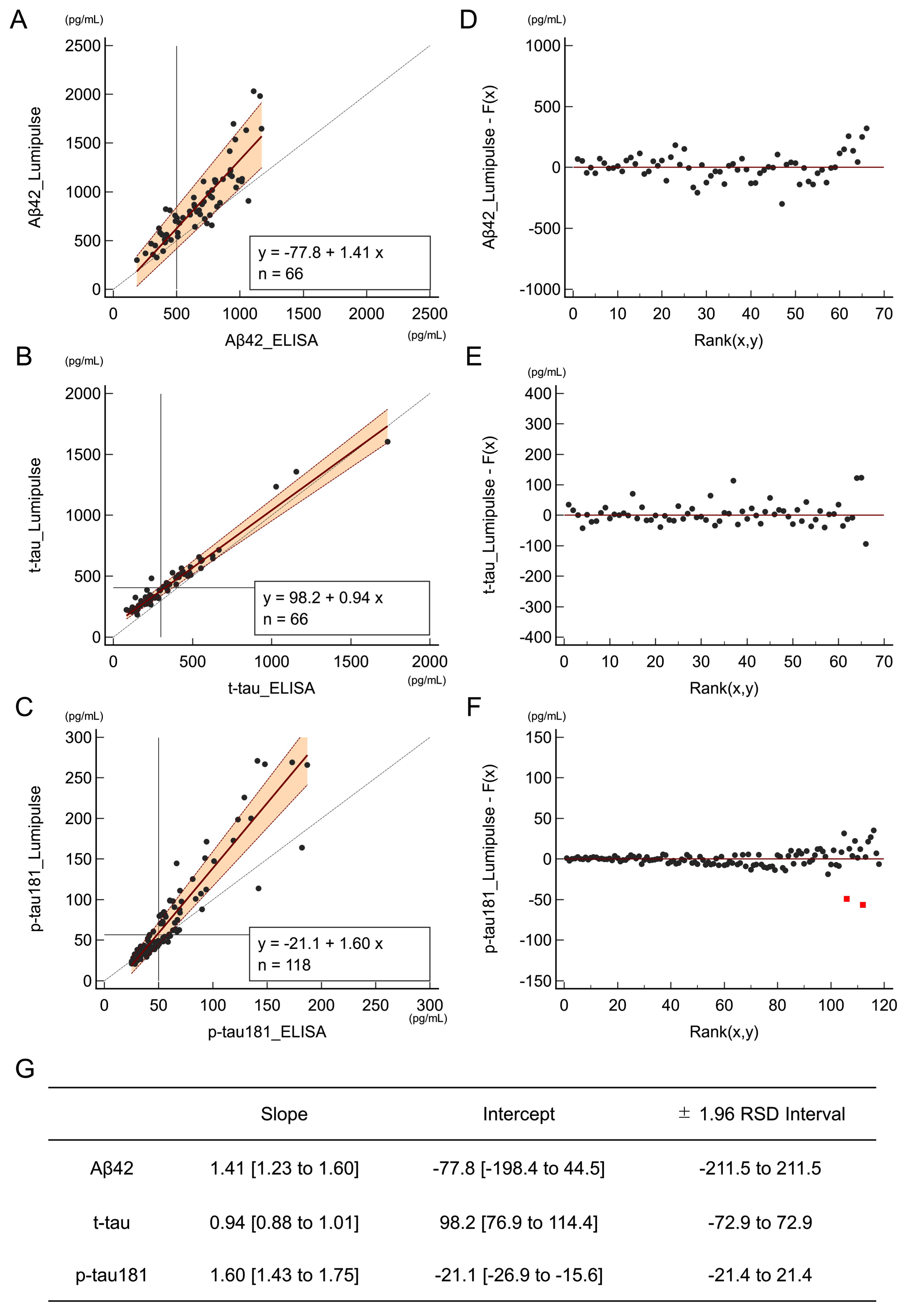
Passing-Bablok regression lines and residual plots for Aβ42, t-tau, and p-tau181 using Lumipulse and ELISA. Each point represents the measurement results for each participant. Passing-Bablok regression lines with 95% confidence intervals (A–C) and residuals from the regression line plotted by rank order (D–F) are shown for Aβ42, t-tau, and p-tau181. The diagonal dotted lines represent lines or equalities (x = y). The horizontal and vertical lines represent the predetermined cutoffs. In the residual plots, outliers, defined as residuals outside the 4 SD limit are shown as red squares (F). (G) Median [95% CI] of slopes and intercepts and±1.96 residual standard deviation (RSD) intervals are summarized. Aβ42, amyloid-β 1-42; t-tau, total tau; p-tau 181, tau phosphorylated at threonine 181; ELISA, enzyme-linked immunosorbent assay.
A flowchart of the screening process is provided in Fig. 2. First, 745 publications were identified via PubMed search. A total of 634 studies were excluded because they did not meet the inclusion criteria and thirty-six were excluded because of duplication. We carefully read the complete texts and supplementary materials of 75 publications and identified 13 that met the predetermined criteria. The reference lists of these publications were also evaluated, and we did not identify any additional publications that met the predetermined criteria. Therefore, 13 publications were included in this systematic review [15–25, 38].
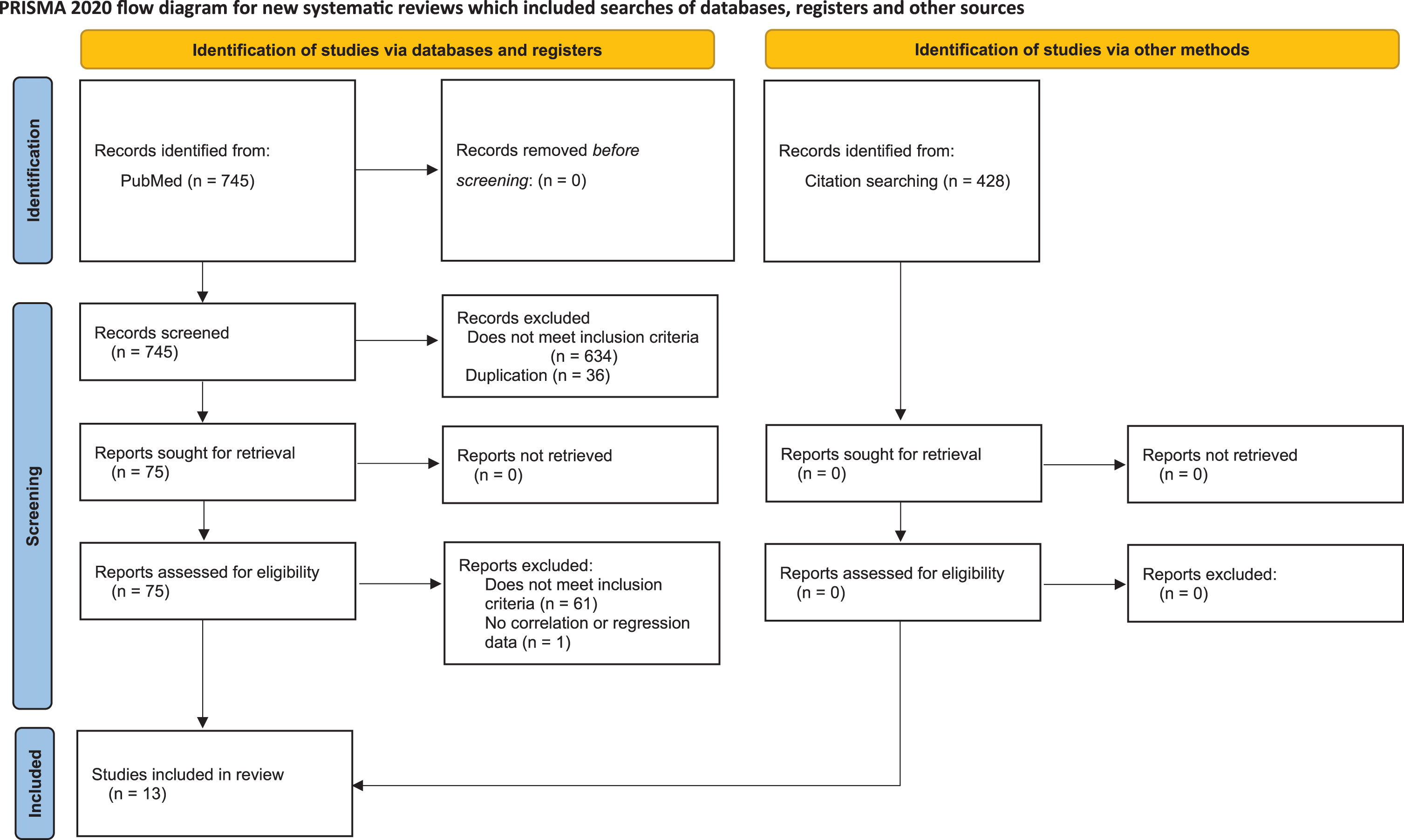
Study screening process following the Preferred Reporting Items for Systematic Reviews and Meta-analysis (PRISMA) 2020.
A total of 2,117 patients were included in 13 studies (Table 2). Eleven studies were conducted in European countries, one was from the United States, and one was from Korea. The disease types of the populations included in the studies were AD dementia, mild cognitive impairment (MCI), cognitive normal, subjective cognitive decline, and other disease controls. Seven studies used Passing-Bablok analysis of the actual values to determine the regression line, whereas some studies used Pearson’s correlation line or log transformation before analyses or did not report the method to analyze the regression line. To measure the correlation coefficients, five studies used Pearson’s method, and five studies used Spearman’s method with actual values.
Baseline characteristic information of studies included in the systematic review
USA, United States of America; UK, United Kingdom; AD, Alzheimer’s disease (dementia); MCI, mild cognitive impairment; CN, cognitive normal; DLB, dementia with Lewy bodies; FTD, frontotemporal dementia; SCD, subjective cognitive decline; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; CIDP, chronic inflammatory demyelinating polyneuropathy; FTLD, frontotemporal lobar degeneration; MSA, multiple system atrophy; iNPH, idiopathic normal pressure hydrocephalus; ALS, amyotrophic lateral sclerosis; NIID, neuronal intranuclear inclusion disease; n/a, not available.
The results of the quality assessment as a biomarker measurement procedure comparison study using the QUABICS assessment tool are presented in Table 3. The agreement between the two raters and distribution of the total scores are summarized in Supplementary Figures 3 and 4. Four and six studies were evaluated to be of high and medium quality, respectively (Table 3).
Quality assessment of studies based on the biomarker measurement procedure comparison studies (QUABICS) assessment tool
QC, quality control. Based on the distribution of total scores, total scores 16–20 and 12–15 were classified as high and medium quality respectively in this study (Supplementary Material).
The primary outcome measures (slopes and intercepts of the regression line) and additional outcome measures (correlation coefficients) for each study are summarized in Table 4. As only one study for Aβ42 and one study for Aβ40 used other ELISA, and the remaining studies used INNOTEST or identical ELISA (Innotest), we focused on the relationship between Lumipulse and Innotest ELISA. Forrest plots and results of the meta-analysis of the primary outcome measures for each biomarker are shown in Fig. 3. Overall, although regression lines slopes of Aβ42 varied between studies and a study reported in 2019 showed values higher than 1 [20], the remaining results other than our study distributed close to 1 and the pooled slope was 1.17 (95% CI: 1.01 to 1.34). Intercepts tended to be lower than 0 and pooled intercept was –95.8 (95% CI: –125.0 to –66.5) (Fig. 3A). Slopes and intercepts of t-tau were distributed around 1 and 0 and pooled slopes and intercepts were 1.00 (95% CI: 0.91 to 1.08) and 34.0 (95% CI: 2.4 to 65.6), respectively (Fig. 3B). A single study reported slope lower than 1 [18] and the remaining five studies reported slopes of p-tau181 higher than 1 distributing from 1.5 to 2.0 [19, 38] and the pooled slope was 1.49 (95% CI: 1.18 to 1.80) (Fig. 3C). The intercepts of p-tau181 were lower than 0 in all studies and pooled intercept was –24.9 (95% CI: –31.1 to –18.7) (Fig. 3C). Only two studies were found assessing for Aβ40 [19, 25] with different results (Fig. 3D). Pooled slope and intercept were 0.92 (95% CI: 0.74 to 1.10) and –504 (95% CI: –4420 to 3412), respectively. The correlation coefficient was 0.83–0.94, 0.79–0.98, 0.91–0.97, and 0.76–0.89 for Aβ42, t-tau, p-tau181, and Aβ40, respectively. The pooled correlation coefficients were 0.91 (95% CI: 0.89 to 0.93) for Aβ42, 0.96 (95% CI: 0.93 to 0.98) for t-tau, 0.95 (95% CI: 0.93 to 0.96) for p-tau, and 0.87 (95% CI: 0.83 to 0.90) for Aβ40 (SupplementaryFigure 5A–D). Publication bias was within the acceptable range, as assessed using funnel plots (Supplementary Figure 5E–H and SupplementaryFigure 6).
Summary characteristics of studies included in the systematic review
Mean/median (95% confidence interval) are listed. n: number of samples, n/a: not available. *the confidence interval of Aβ42 intercept by Leitão et al. did not include the median value [19]. **the Pearson’s correlation coefficient by Keshavan et al. was based on log-transformed values [22]. ***the slope of p-tau181 was provided as 0.88 in the manuscript by Bayart et al., although in the figure the slope seems to be more than 1 [18]. Aβ42, amyloid-β 1-42; t-tau, total tau; p-tau 181, tau phosphorylated at threonine 181.
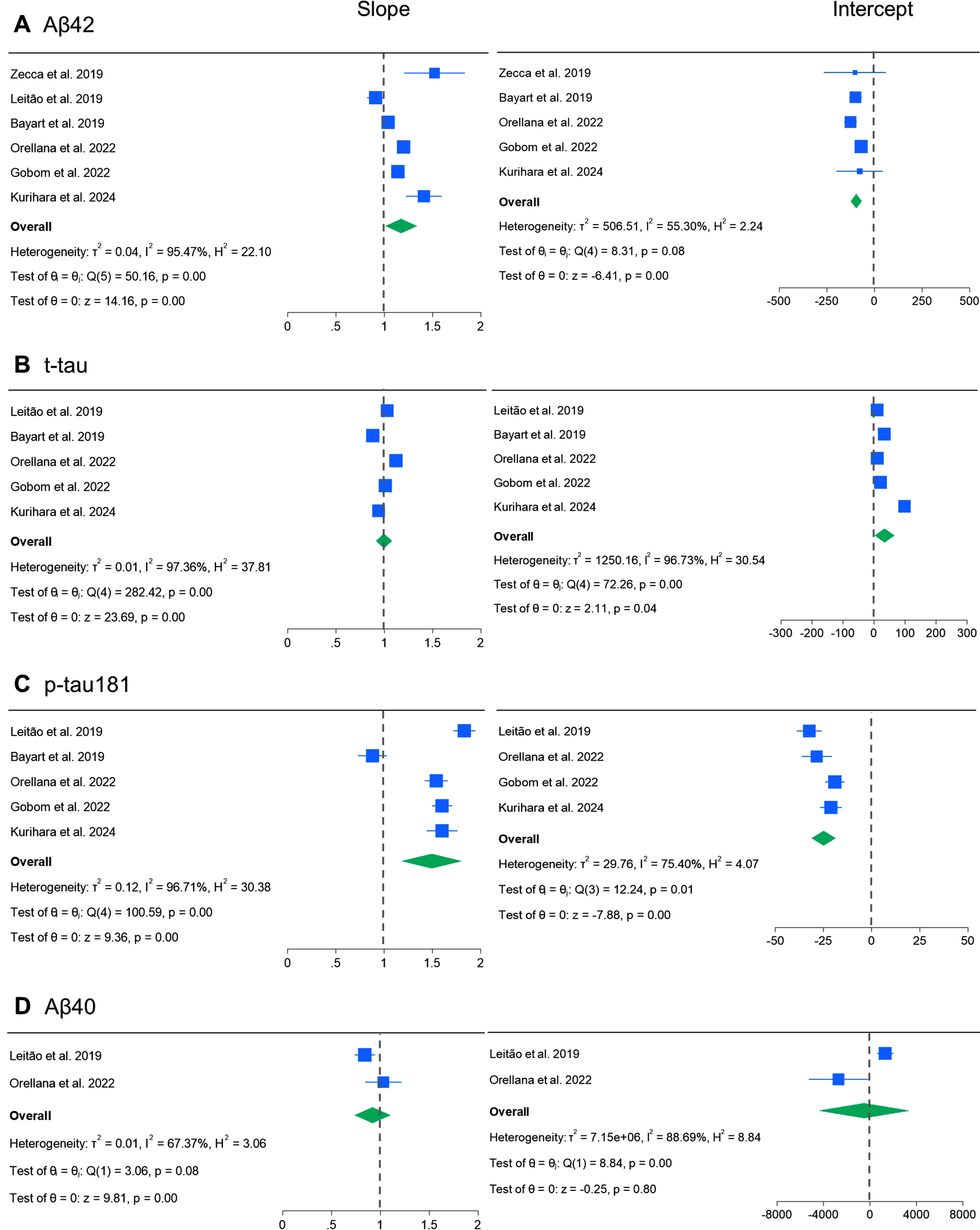
Forrest plot and meta-analysis of slopes and intercepts of the regression lines. Forrest plot and meta-analysis using random effect model for slopes and intercepts of Aβ42 (A), t-tau (B), p-tau181 (C), and Aβ40 (D). The square size for each study represents the study weight. Aβ42, amyloid-β 1-42; t-tau, total tau; p-tau 181, tau phosphorylated at threonine 181.
We reported the results of a single-center analysis and systematic review of the relationship between cerebrospinal fluid AD biomarker values measured using Lumipulse assays and conventional Innotest ELISA. Although we confirmed a good correlation between all biomarkers, we identified different characteristics between the measured values for each biomarker.
The slope of the regression lines of Aβ42 varied between studies and two studies reported in 2018–2019 showed values close to 1.5 [15, 20]. This may be explained by the Aβ42 adjustment measured via Lumipulse assay based on certified reference materials since December 2018, which divides previous values by 1.46 [24]. Excluding these two reports, although some variances were observed between the studies, the slopes seemed to be distributed around 1 except for our study. The intercepts were lower than 0 in most studies, suggesting systematic differences. Although our results reflect the above adjustment, the regression line slope was higher than previous studies. Previous studies reported higher inter-lab CV in manual ELISA compared to automated Lumipulse assays [14, 39]; this difference is likely because of lower Aβ42 values in our ELISA measurement compared to these studies. Our study has a limitation that while CSF sample handling procedures were in line with the current recommendations [26] for all p-tau181 ELISA measurements and majority of the Lumipulse measurements, Aβ42 and t-tau ELISA were conducted using biobank samples using different sample handling protocols involving aliquoting, which may have decreased the concentration of Aβ42 through protein binding to tubes [26, 41]. Although the sample size was extremely small to draw conclusions, subgroup analysis has suggested this hypothesis (Supplementary Figure 2). We did not compare the measurement values for Aβ40, since our institution have not measured Aβ40 in ELISA in the past, and we could only find two studies reporting slope or intercept of the regression line of Aβ40, with somewhat different results.
While some differences were observed between studies, the slopes and intercepts of t-tau were distributed close to 1 and 0, respectively, suggesting minimal proportional and systematic bias. Based on the small overall bias and variability between studies, the measurement results of t-tau between Lumipulse and Innotest ELISA may be interconvertible using a conversion formula, although the relationship should be confirmed in each laboratory and the formula must be locally adjusted.
In contrast, significant proportional and systematic differences were observed between the p-tau181 values. The regression line slope was significantly higher than 1 in five of six previous studies and in our study, distributed between 1.5–2.0. Although one study reported an outlying median slope value of 0.88 [18], the median regression line slope seemed to be greater than 1 in the figure. The regression line intercept was significantly lower than 0 in all studies distributing around –25 pg/mL. Pooled slope and intercepts were 1.49 (95% CI: 1.18 to 1.80) and –24.9 (95% CI: –31.1 to –18.7), respectively. The results of this systematic review suggest that the proportional and systematic differences in p-tau181 values reported in previous studies [19, 25] are not specific to certain races or laboratories, but are fundamental in the Lumipulse and Innotest assays. The reason for this relationship, despite the use of similar antibody combinations, remains unclear and warrants further evaluation for future standardization.
One of the main advantages of fluid biomarkers is that we can obtain not only binary results (positive versus negative), but also continuous measurement values. Although the current research framework uses binary classification with single cutoffs [2], different cutoffs may be optimal for different situations. Tertiary classification using two cutoffs has been employed in the United States for CSF Aβ42/40 ratio [42]. Recently, using neuropathological data at autopsy as gold standard, we have shown that CSF p-tau181 levels are associated with both amyloid and tau pathology and suggested that using two different cut-offs may be useful to differentiate between patients into three groups; amyloid-negative, amyloid-positive with tau pathology limited in the transentorhinal region, and amyloid-positive with expanding tau pathology [8]. Moreover, fluid biomarkers may change dynamically after the administration of disease-modifying therapies in line with clinical benefits [43, 44] and could be important as disease-monitoring biomarkers. Therefore, understanding the direct relationship between biomarker measurement values across different platforms is important.
Although validating predetermined cutoff values was not the aim of this study, our scatter plots and regression lines indicated that predetermined Lumipulse cutoff values for t-tau and p-tau181 highly corresponded to our ELISA cutoff values based on previous autopsy data [28, 29] and the binary results matched well between the two assays. These results may support the use of predetermined Lumipulse cutoff values based on data obtained from other racial groups in Japanese patients.
While we conducted this systematic review, we experienced several issues that could be addressed in future biomarker measurement procedure comparison studies. First, the patient populations differed widely between the studies. Combining the results, as we did in this systematic review, is important to obtain a complete overview of the characteristics of the different platforms. Second, although CSF sample handling and analysis methods seem to be relatively standardized by virtue of global standardization efforts [12, 26], statistical methods varied widely between studies. Based on previous recommendations [32, 33], we believe that the concentration value itself (not the logarithmic transform) should be evaluated using difference plots and a robust nonparametric method that assumes error in both the x- and y-axes in a situation where a normal distribution is difficult to assume, for example, the Passing-Bablok regression analysis. Third, data reporting also varied between studies. As recommended, the adjusted median and CI of the slope and interval should be reported [32, 33] for future global harmonization of the data.
There are several limitations to this study. First, as mentioned above, we used aliquoted biobank samples for Aβ42 and t-tau ELISA, which likely led to decreased Aβ42 value in ELISA because of protein binding to tubes. Second, we have not participated in the quality control program, and the measurement value of ELISA was not adjusted based on certified reference materials. Third, we searched only English articles in the systematic review and there may be several other studies reported in different languages.
In conclusion, while biomarkers measured by the Lumipulse assay and Innotest ELISA showed good correlation, we identified different characteristics between the measured values for each biomarker, which could be useful for a seamless understanding of previous and future AD biomarker studies. Our results should aid in understanding the differences in CSF biomarker results measured on different platforms and in future global harmonization.
AUTHOR CONTRIBUTIONS
Masanori Kurihara (Conceptualization; Data curation; Formal analysis; Funding acquisition; Investigation; Project administration; Visualization; Writing – original draft); Soichiro Kondo (Investigation; Validation; Writing – review & editing); Kensuke Ohse (Investigation; Writing – review & editing); Hisashi Nojima (Investigation; Writing – review & editing); Emiko Kikkawa-Saito (Investigation; Writing – review & editing); Atsushi Iwata (Funding acquisition; Supervision; Writing – review & editing).
Footnotes
ACKNOWLEDGMENTS
We thank all participants of Tokyo Medical Biobank for donating their clinical information and biosamples. We also thank all members of the Department of Neurology, Research Team for Neuroimaging, Healthy Aging Innovation Center (HAIC), and the Integrated Research Initiative for Living Well with Dementia (IRIDE) of the Tokyo Metropolitan Institute for Geriatrics and Gerontology (TMIG) for their assistance.
FUNDING
This study was supported by the Integrated Research Initiative for Living Well with Dementia (IRIDE) of the Tokyo Metropolitan Institute for Geriatrics and Gerontology, KAKENHI from the Japan Society for the Promotion of Science (JSPS) to M. Kurihara (JP23K14789), and by the Japan Agency for Medical Research and Development (AMED) to A. Iwata (JP23dk0207057h0002).
CONFLICT OF INTEREST
Hisashi Nojima and Emiko Kikkawa-Saito are employees of FUJIREBIO INC. Atsushi Iwata received a research grant from FUJIREBIO INC. The remaining authors declare no conflicts of interest relevant to this manuscript.
DATA AVAILABILITY
The datasets and full protocol for the present study are available from the corresponding author upon request.
