Abstract
Background:
Previous observational studies suggested an association between sepsis and neurodegenerative diseases, but causality remains unclear.
Objective:
Determining the causal association between sepsis and four neurodegenerative diseases (Alzheimer’s disease, Parkinson’s disease, amyotrophic lateral sclerosis, and Lewy body dementia) through bidirectional two-sample Mendelian randomization (MR) analysis.
Methods:
Genome-wide association study summary statistics for all traits were obtained from publicly available databases. Inverse variance weighted (IVW) was the primary method for evaluating causal associations. In addition, three additional MR methods (MR-Egger, weighted median, and maximum likelihood method) were employed to supplement IVW. Furthermore, various sensitivity tests were conducted to assess the reliability: 1) Cochrane’s Q test for assessing heterogeneity; 2) MR-Egger intercept test and MR-PRESSO global test for evaluating horizontal pleiotropy; 3) leave-one-out sensitivity test for determining the stability.
Results:
The results of IVW indicated that sepsis significantly increased the risk of Alzheimer’s disease (OR = 1.11, 95% CI: 1.01–1.21, p = 0.025). In addition, three additional MR methods suggested parallel results. However, no causal effect of sepsis on the three other neurodegenerative diseases was identified. Subsequently, reverse MR analysis indicated that the four neurodegenerative diseases do not causally affect sepsis. Furthermore, sensitivity tests demonstrated the reliability of the MR analyses, suggesting no heterogeneity or horizontal pleiotropy.
Conclusions:
The present study contributes to a deeper comprehension of the intricate interplay between sepsis and neurodegenerative disorders, thereby offering potential avenues for the development of therapeutic agents that can effectively mitigate the multifarious complications associated with sepsis.
Keywords
INTRODUCTION
Neurodegenerative disorders, such as Alzheimer’s disease (AD), Lewy body dementia (LBD), Parkinson’s disease (PD), and amyotrophic lateral sclerosis (ALS), are a diverse group of progressive and age-dependent neurological diseases, affecting an estimated 50 million individuals worldwide [1]. As the global population continues to age, the prevalence of neurodegenerative diseases is steadily rising, posing a significant public health concern across various nations [2]. Despite notable advancements in molecular biology, genetics, and pharmaceutical research, the development of efficacious drugs to impede or postpone neurodegeneration remains elusive [3–5]. Consequently, there is an urgent need to identify modifiable risk factors and pursue potential therapeutic approaches to decelerate the progression or even preempt the onset of these diseases. Since bacterial infections have been shown to be a potential risk factor for dementia [6, 7], our investigation aims to explore infection-related risk factors in neurodegenerative diseases, with a specific focuson sepsis.
Sepsis, a life-threatening condition characterized by organ dysfunction resulting from a dysregulated host response to infection, is presented as a refractory disease with significant morbidity and mortality [8–10]. Various complications such as malaise, renal impairment, respiratory compromise, and cognitive dysfunction are observed in at least 20% of sepsis survivors after discharge from the hospital, severely affecting their quality of life [11, 12]. The systemic inflammation caused by sepsis is believed to have long-term detrimental effects on the brain [13]. Studies have demonstrated that a substantial number of sepsis survivors, approximately 3 million annually, develop cognitive dysfunction, as evidenced by persistent slowing of electroencephalographic (EEG) activity, impaired memory, and reduced hippocampal volume [14]. Numerous observational studies have reported that sepsis is an independent risk factor for dementia and that sepsis survivors are more likely to suffer from dementia [15, 16]. However, this conclusion remains controversial. A comprehensive national study conducted in Sweden reported no association between sepsis and an elevated risk of dementia [17]. In addition, investigations have confirmed that hospital-treated infections were linked to an increased risk of AD and PD, but not ALS [18, 19]. Furthermore, previous research has indicated that patients with neurodegenerative diseases are more susceptible to developing infections and sepsis [20–22]. However, it is important to note that the majority of studies investigating the relationship between sepsis and neurodegenerative diseases have been observational, making it challenging to establish a causal relationship. Consequently, these findings may be prone to biased conclusions, highlighting the need for more robust evidence to support the proposed association.
Mendelian randomization (MR) represents an epidemiological approach that leverages genetic variation as an instrumental variable in order to evaluate the causal association between modifiable risk factors and disease outcomes [23]. By exploiting the random assignment of single-nucleotide polymorphisms (SNPs) at conception and their independence from confounding factors, MR effectively overcomes the limitations inherent in observational studies [24]. In addition, MR investigations are good alternative approaches for exposures that cannot be intervened in certain clinical practices due to ethical issues. Therefore, we conducted a bidirectional two-sample MR analysis based on publicly available genome-wide association study (GWAS) summary statistics. The present study aimed to establish a causal relationship between sepsis and neurodegenerative diseases.
MATERIALS AND METHODS
Overall study design
We conducted a bidirectional two-sample MR analysis. First, we obtained GWAS summary statistics for sepsis, AD, PD, ALS, and LBD, which were derived from separate cohorts. Subsequently, SNPs associated with sepsis were considered as instrumental variables (IVs) for forward MR analysis to estimate the causal effects of sepsis on four degenerative diseases. Finally, SNPs associated with four degenerative diseases were considered as IVs for reverse MR analysis to estimate the causal effects of four degenerative diseases on sepsis. Figure 1 shows the overall flowchart of this study, and detailed information will be illustrated in subsequent parts of this section. This MR study was primarily conducted using the “TwoSampleMR” R package and the “MRPRESSO” R package in R software (version 4.3.1).
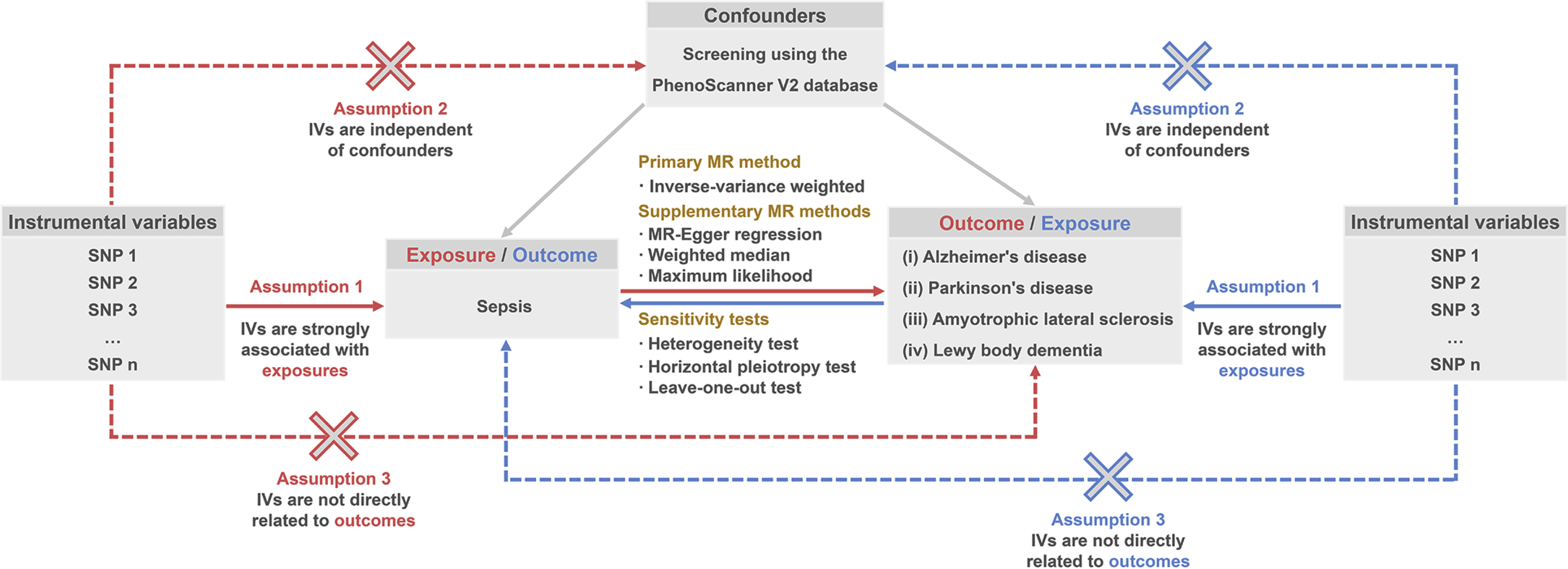
The overall flowchart of the present MR study.
Data sources for GWAS summary statistics
The GWAS summary statistics used for this MR study were downloaded from the IEU Open GWAS database and GWAS Catalog database, which are two open-source databases containing massive amounts of GWAS summary statistics [25, 26]. GWAS summary statistics for sepsis were derived from the UK Biobank cohort, containing 10,154 cases and 454,764 controls. In addition, the present MR study included four degenerative diseases: AD (21,982 cases and 41,944 controls) [27], PD (33,674 cases and 449,056 controls) [28], ALS (20,806 cases and 59,804 controls) [29], and LBD (2,591 cases and 4,027 controls) [30]. Details of the included GWAS summary statistics are shown in Supplementary Table 1.
Selection of IVs
The IVs were screened based on the three core assumptions recognized for MR studies: 1) Relevance: IVs exhibit a strong correlation with exposure; 2) Independence: IVs are not related to potential confounders; 3) Exclusion restriction: IVs only affect outcomes through exposure, with no direct association with outcomes [31].
To fulfill the first MR core assumption, for forward MR analysis, IVs associated with sepsis were screened using a threshold of p < 5e-6 due to the limited number of IVs obtained when using a threshold of p < 5e-8. For reverse MR analysis, IVs were screened for AD, PD, ALS, and LBD, respectively, based on a threshold of p < 5e-8. In addition, SNPs in linkage disequilibrium would be excluded from IVs (threshold of r2 < 0.001 within 10000 kb). Furthermore, the F-statistics of each IV were calculated. Only IVs with F-statistics >10 were retained for subsequent MR analysis to avoid bias caused by weak IVs [32]. To fulfill the second MR core assumption, all traits associated with IVs were obtained using the PhenoScanner V2 database [33]. SNPs associated with potential confounders would be removed from the IVs. To fulfill the third MR core assumption, we thoroughly examined the correlations between each IV and both exposure and outcome, ensuring that the IVs exhibit a stronger correlation with exposure than with the outcome. Furthermore, various sensitivity tests were administered to evaluate horizontal pleiotropy, namely, the degree to which IVs do not affect outcomes through exposure.
Statistical analysis
The primary MR method for assessing the causal effect of exposure on outcome was the inverse variance weighted (IVW), which first estimates the causal effect by the Wald ratio method using each individual IV, followed by a meta-analysis of these results [34]. In addition, three other MR methods were employed to supplement the IVW, namely MR-Egger regression, weighted median, and maximum likelihood. MR-Egger regression can provide relatively reliable estimates in the presence of horizontal pleiotropy [35]. Weighted median provides reliable estimates when only half of the IVs meet the criterion [36]. The maximum likelihood method, a conventional approach, was employed to estimate probability distribution parameters by maximizing the likelihood function and minimizing standard errors [37]. Since the outcome variable is binary, odds ratios (OR) and 95% confidence intervals (CI) were used to present the MR results, and p < 0.05 was considered statistically significant. Causal estimates could be considered reliable only if the IVW method identified statistically significant MR results. In addition, the ORs identified by three additional supplementary MR methods were required to be consistent with IVW.
Subsequently, a series of sensitivity tests were conducted to assess the overall reliability of the present MR study: 1) Cochrane’s Q test was performed to assess heterogeneity, namely variability in causal effects assessed by separate IVs; 2) The MR-Egger intercept test and MR-PRESSO global test were utilized to assess horizontal pleiotropy, which refers to the extent to which IVs affect the outcome independent of the exposure; 3) The leave-one-out test was used to assess stability, whereby replicate MR analyses were performed after eliminating each individual IV, thus assessing whether the MR results were interfered with by outlier IVs.
RESULTS
Results of forward MR analysis
This section will present the causal effects of sepsis on AD, PD, ALS, and LBD evaluated by MR analysis. For IVs proxying for sepsis, rs3130320 was excluded since it was associated with multiple sclerosis identified by PhenoScanner V2. In addition, rs12203592 was excluded as its F-statistics were less than 10. Subsequently, after excluding SNPs that were missing from the GWAS summary statistics of outcomes, 24, 26, 26, and 25 SNPs were ultimately utilized as IVs for evaluating the causal effect of sepsis on AD, PD, ALS, and LBD, respectively. In addition, all IVs showed stronger associations with exposure than with outcome. Furthermore, the F-statistics of all IVs used for MR analysis ranged from 12.33–31.28. Details of the IVs utilized for forward MR analysis are presented in Supplementary Table 2.
Figure 2 shows the results of the primary MR method, IVW, evaluating the causal effect of genetically determined sepsis on four neurodegenerative diseases. Interestingly, sepsis significantly increased the risk of AD (OR = 1.11, 95% CI: 1.01–1.21, p = 0.025). In addition, the MR results of three additional methods were parallel to IVW (OR > 1), supporting the reliability of the causal relationship (Table 1). However, no causal effect of sepsis on PD, ALS, and LBD was examined (p > 0.05) (Fig. 2).
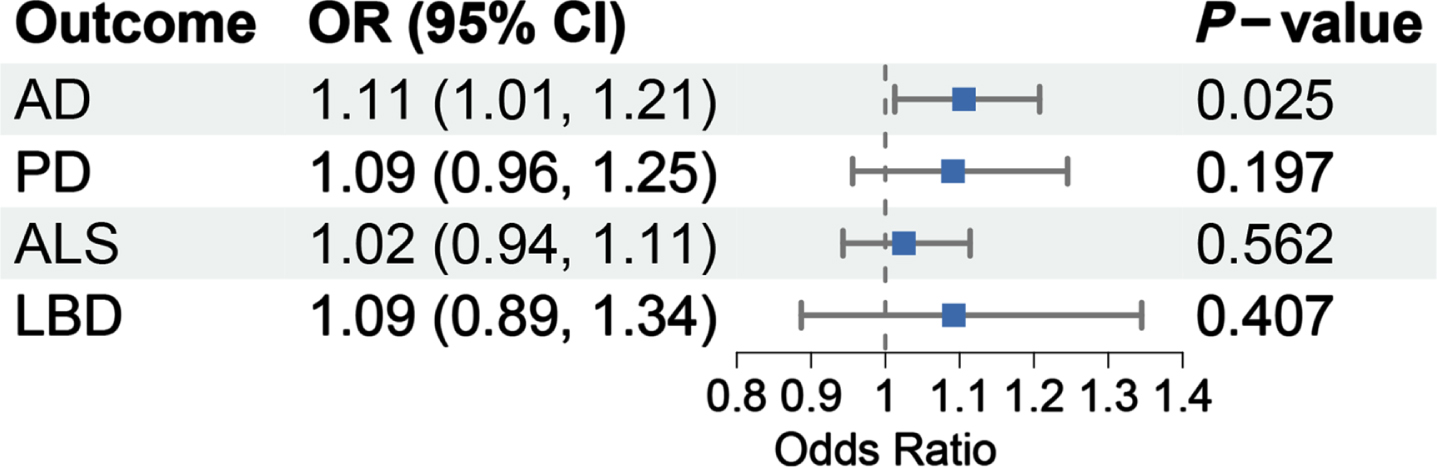
Primary MR results regarding the causal effect of sepsis on four neurodegenerative diseases evaluated by IVW. OR, odds ratio; CI, confidence interval; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; LBD, Lewy body dementia.
Causal effects of sepsis on four neurodegenerative diseases evaluated by various MR methods
OR, odds ratio; CI, confidence interval; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; LBD, Lewy body dementia; IVW, inverse variance weighted.
Various sensitivity tests demonstrate the reliability of forward MR results. Firstly, Cochran’s Q test showed no significant heterogeneity among the IVs used for forward MR analysis (p > 0.05) (Table 2). Secondly, the MR-Egger intercept test and the MR-PRESSO global test showed that forward MR analysis was not significantly influenced by horizontal pleiotropy (p > 0.05) (Table 3). Finally, the leave-one-out sensitivity test indicated the stability of the forward MR analysis, since excluding any IV did not significantly alter the results (Supplementary Figure 1).
Results of Cochran’s Q heterogeneity test for forward MR analysis
AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; LBD, Lewy body dementia; IVW, inverse variance weighted.
Results of the horizontal pleiotropy test for forward MR analysis
SE, standard error; RSS, residual sum of squares; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis; LBD, Lewy body dementia.
Results of reverse MR analysis
This section will present the causal effects of AD, PD, ALS, and LBD on sepsis evaluated by MR analysis. For IVs proxying for the four neurodegenerative diseases, SNPs associated with white cell counts identified by PhenoScanner V2 were excluded from IVs since they could be confounders affecting sepsis. Details of the IVs that were ultimately utilized for the reverse MR analysis are displayed in Supplementary Table 3. All IVs exhibited stronger correlations with exposure than with outcome, and F-statistics were all > 10 (29.82–962.36).
As shown in Supplementary Table 4, all MR methods did not support a causal effect of AD, PD, ALS, and LBD on sepsis (p > 0.05). Cochran’s Q test showed that the reverse MR analysis was not affected by remarkable heterogeneity (p > 0.05) (Supplementary Table 5). Both the MR-Egger intercept test and MR-PRESSO global test showed that the reverse MR analysis was not disturbed by significant horizontal pleiotropy (p > 0.05) (Supplementary Table 6). The leave-one-out sensitivity test demonstrated the stability of the MR results, showing no causal effects of the four neurodegenerative diseases on sepsis, regardless of which IV was excluded (Supplementary Figure 2).
DISCUSSION
In the present study, we employed a bidirectional MR approach to explore the causal association between sepsis and the risk of neurodegenerative disorders, including AD, LBD, PD, and ALS. Our results indicated that sepsis increased the risk of AD but had no causal effect on the risk of LBD, PD, and ALS. Reverse MR analysis demonstrated that the four neurodegenerative diseases did not have any causal association with susceptibility to sepsis. This is the first large-scale MR investigation using genetic instruments from publicly available GWAS to provide evidence for the causal relationship between sepsis and neurodegenerative disorders, thereby avoiding the confounding effect of traditional observational studies.
Previous clinical studies have reported sepsis as a risk factor for dementia. A retrospective observational study in Germany showed that sepsis survivors had a significantly higher risk of developing mid-stage dementia than non-sepsis patients [38]. Another study of elderly Medicare beneficiaries found that infection or severe sepsis, neurologic impairment, and acute dialysis during intensive care hospitalization exhibited independent associations with a subsequent diagnosis of dementia [16]. Similarly, a cohort study from U.S. veterans found that peripheral bacterial infections were associated with a significantly increased risk of developing dementia [7]. In addition, a systematic review and meta-analysis also found that sepsis survivors, especially those with severe sepsis, had a significantly increased risk of all-cause dementia [15]. Furthermore, a recent 40-year study suggests that the risk of all-cause dementia rises after hospital-diagnosed infections [39]. AD, a common type of dementia, has gained substantial attention in numerous studies due to its robust association with bacterial infections, including Chlamydia pneumoniae, Borrelia burgdorferi, and Porphyromonas gingivalis. Consistent with the above evidence, our findings established a causal link between sepsis and an elevated risk of AD development, while revealing a positive albeit nonsignificant genetic correlation between sepsis and other types of dementia such as LBD. However, contrasting outcomes emerged from a population-based study conducted in Taiwan, where sepsis survivors exhibited a heightened risk of non-Alzheimer’s dementia among dementia subtypes (HR: 2.26, 95% CI: 2.07–2.47, p < 0.001), without any discernible association observed with AD risk (HR: 1.17, 95% CI: 0.84–1.62, p > 0.05) [44]. This discrepancy could potentially be attributed to the limited sample size, regional bias, and the inclusion of individuals with prior sepsis and stroke in the study cohort.
The precise mechanism underlying the association between sepsis and AD remains incompletely understood [45]. Nonetheless, several mechanisms have been speculated to be involved. First, when sepsis-induced systemic inflammation occurs, pro-inflammatory mediators can disrupt the BBB, facilitating their passage into the brain [46]. Upon entry, these mediators activate cytotoxic microglia in the brain, leading to neuronal damage and apoptosis, cognitive deterioration, and an increased risk of dementia [47]. A recent investigation has shown that acute systemic inflammation has the potential to induce a phenotypic alteration in neuroinflammation, characterized by an interleukin-1α (IL-1α)-driven amplified response of microglia, astrocytes, and neuronal network dysfunction, thereby exacerbating neuroinflammation in AD [48]. Additionally, sepsis induces enduring innate immune memory within the murine brain, rendering it more susceptible to Aα oligomers, central neurotoxins implicated in AD [49]. Moreover, in mouse models of AD, sepsis has demonstrated the ability to worsen cognitive decline and AD pathophysiology by modulating the gut microbiome, augmenting hippocampal Aα plaque accumulation, and promoting neuroinflammation [50]. It is worth noting that investigations elucidating the relationship and mechanisms between sepsis and AD are predominantly based on rodent models, with limited human-based studies. Consequently, further exploration is imperative to unravel the potential pathologicalmechanism.
In contrast, a causal association between sepsis and other neurodegenerative disorders such as LBD, PD, and ALS was not established in our study. This discrepancy may arise from the distinct pathogenesis underlying AD when compared to other diseases. However, we acknowledge that our findings cannot definitively exclude the possibility of the association, as the current sample size may be inadequate for detection. With future GWAS summary statistics having larger sample sizes, the causal correlation between sepsis and other neurodegenerative diseases may become significant.
There is limited evidence from current studies on whether neurodegenerative diseases increase susceptibility to sepsis. Advanced dementia and PD have been associated with an elevated susceptibility to infections, especially pneumonia, and subsequently sepsis, due to the patients’ decline in body resistance and their difficulty swallowing food and choking which can result in the inhalation of food into the lungs [51–54]. A recent study showed that individuals with PD exhibit a heightened vulnerability to CNS infections and sepsis when compared to controls [21]. ALS patients may also face an increased risk of infections due to compromised immune systems and weakness in respiratory muscles [55]. Lee et al. reported that ALS patients are more prone to hospitalization for sepsis following diagnosis compared to those without ALS [22]. However, our MR analyses suggested a trend toward a positive but not significant association between these four neurodegenerative diseases and susceptibility to sepsis. This association awaits further analysis in the future when GWAS data with larger sample sizes become available.
The present MR study has several advantages: 1) large-scale GWAS summary statistics were employed for the study, and the large sample size provided relatively reliable results; 2) comprehensive bidirectional MR analyses were conducted using multiple MR methods, thus clarifying the directionality of the causal associations; and 3) a variety of sensitivity tests were conducted to enhance the reliability of the conclusions.
However, some limitations of this study need to be acknowledged: 1) first, the GWAS summary statistics included in this study were derived from European populations, and therefore it is unclear whether the findings are generalizable to other populations. Future studies utilizing large-scale GWAS summary statistics from diverse populations are essential to assess the generalizability of these findings across different ethnic groups; 2) Due to the use of GWAS summary statistics for MR analyses and the unavailability of GWAS data specific to diverse demographic groups, stratified analyses based on age or sex were not feasible; and 3) despite the multiple sensitivity tests, completely eliminating the effect of horizontal pleiotropy is challenging.
Conclusion
The present study contributes to a deeper comprehension of the intricate interplay between sepsis and neurodegenerative disorders, thereby offering potential avenues for the development of therapeutic agents that can effectively mitigate the multifarious complications associated with sepsis.
Footnotes
ACKNOWLEDGMENTS
We want to acknowledge the IEU OpenGWAS project database (https://gwas.mrcieu.ac.uk/) and GWAS Catalog database (![]() ) for providing GWAS summary data.
) for providing GWAS summary data.
FUNDING
This research was funded by the National Natural Science Foundation for Distinguished Young Scholars of China (No. 81900634 to Dr. Tang), the Natural Sciences Foundation of Hunan Province for Distinguished Young Scholars (No. 2021JJ40947 to Dr. Tang), the Natural Science Foundation of Changsha City (No. kq2208356 to Dr. Lin).
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
