Abstract
Background:
Alzheimer’s disease (AD) brains accumulate DNA double-strand breaks (DSBs), which could contribute to neurodegeneration and dysfunction. The genomic distribution of AD brain DSBs is unclear.
Objective:
To map genome-wide DSB distributions in AD and age-matched control brains.
Methods:
We obtained autopsy brain tissue from 3 AD and 3 age-matched control individuals. The donors were men between the ages of 78 to 91. Nuclei extracted from frontal cortex tissue were subjected to Cleavage Under Targets & Release Using Nuclease (CUT&RUN) assay with an antibody against
Results:
The AD brains contained 18 times more DSBs than the control brains and the pattern of AD DSBs differed from the control brain pattern. In conjunction with published genome, epigenome, and transcriptome analyses, our data revealed aberrant DSB formation correlates with AD-associated single-nucleotide polymorphisms, increased chromatin accessibility, and upregulated gene expression.
Conclusion:
Our data suggest in AD, an accumulation of DSBs at ectopic genomic loci could contribute to an aberrant upregulation of gene expression.
Keywords
INTRODUCTION
Alzheimer’s disease (AD), a progressive neurodegenerative disorder that accounts for most dementia cases, has become a global challenge with the aging of the population [1]. At the molecular level, AD is hallmarked by extracellular amyloid plaques and intracellular neurofibrillary tangles [2]. Extracellular amyloid plaques are formed from the abnormal accumulation of amyloid-β protein (Aβ), which potentially disrupts communication between nerve cells. Intracellular neurofibrillary tangles occur when tau protein becomes abnormally phosphorylated, leading to the formation of clumps within nerve cells [2]. These pathological changes result in the death and functional impairment of brain cells and a gradual decline in cognitive function featuring memory loss and difficulty with language, problem solving, and other complex tasks [3]. Despite recent advances in AD research, the exact cause of AD remains unclear. Currently, there is no cure for AD, and all treatment options aim to lessen AD symptoms and improve quality of life. Thus, there is an urgent unmet need to understand the molecular mechanism underlying AD in order to identify therapeutic targets for AD prevention and treatment.
Mounting evidence suggests that genomic integrity is essential for the survival and function of neuronal cells and that loss of genomic integrity and accumulation of DNA damage are involved in neuronal aging and degeneration [4–10]. Since neurons are post-mitotic and long-lived, DNA damage could be especially damaging. Among all forms of DNA damage, DNA double-strand breaks (DSBs) are perhaps the most deleterious. DSBs can occur as a result of various factors, such as ionizing radiation, exposure to chemicals or other genotoxic agents, errors in DNA replication and repair mechanisms. The numbers of DSBs are increased in AD brain cells [10–12]. When DNA breaks occur, cells respond by activating a complex signaling pathway known as the DNA damage response (DDR) to repair the damage and maintain genomic stability. Mutations in the genes that encode proteins in the DDR pathway have been linked to neurodegenerative diseases, including AD [13–16]. In addition, multiple mouse models of neurodegeneration exhibit increased DSB formation, including tauopathy models, the CK-p25 model, 5XFAD, and the hAPP-J20 model [11, 17–19]. Taken together, these results implicate DSBs in AD pathology.
DSBs are formed in brain cells as a physiological response when mice are exposed to novel environments and repaired within 24 h [17]. Further studies have shown that these DSBs are not randomly distributed in the genome but positioned in certain loci to facilitate the expression of early response genes [20, 21]. Thus, it is not known whether the landscape of DSBs is altered in AD pathology. Moreover, dysregulation of gene expression and chromosomal accessibility has been reported in human AD brains, but it is not known whether ectopic DSBs correlate with this dysregulation.
One of the earliest events following DSB formation is the phosphorylation of the histone variant H2AX at serine 139, resulting in the rapid formation of
MATERIALS AND METHODS
Human samples
Autopsy brain tissue samples (frontal cortex) were obtained from the University of Kansas Alzheimer’s Disease Research Center (KU ADRC) Neuropathology Core. The samples were homogenized on dry ice and aliquoted for CUT&RUN. For information on the human samples used in this study (AD,
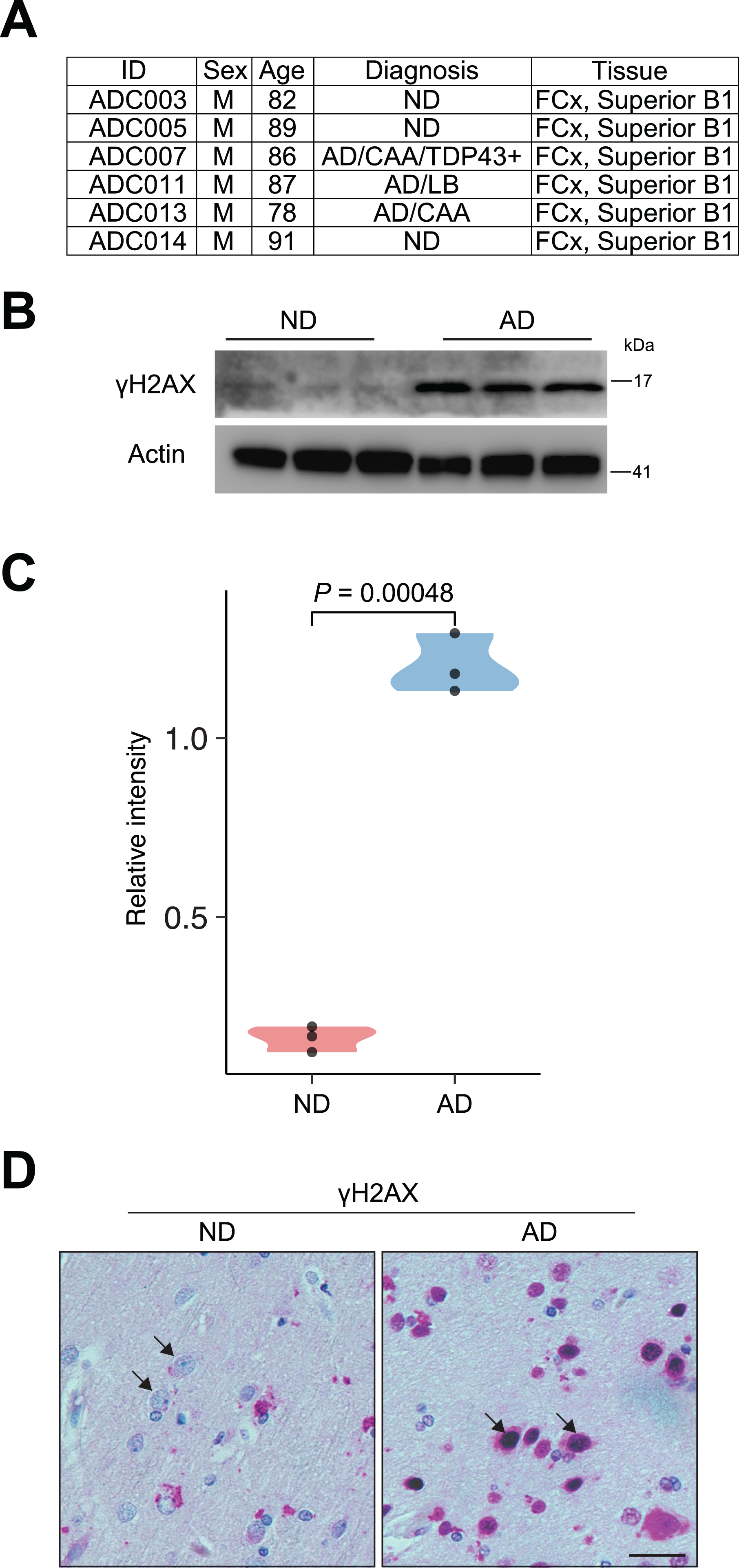
Western blot
Total protein was extracted in RIPA buffer containing 1 mM PMSF (Sigma) and protease inhibitor cocktail (Sigma P8340). The lysates were removed after centrifugation at 12,000×
Immunohistochemistry
For immunohistochemistry, human FFPE histological sections were dewaxed, rehydrated in an ethanol series, and then subjected to microwave antigen retrieval in 0.01 M citrate (pH 6.0) and methanol/H2O2 treatment. After blocking with 5% goat serum, the slides were sequentially incubated with
Preparation of nuclei from human brain specimens
Frozen human brain tissue was thawed on ice and minced sharply into <1 mm pieces. Next, 500μl of chilled Nuclei EZ Lysis Buffer (Millipore Sigma, NUC-101, no. N3408) was added, and the tissue was homogenized 10–20 times in a Dounce homogenizer. The homogenate was transferred to a 1.5-ml Eppendorf tube, and 1 ml of chilled Nuclei EZ Lysis Buffer was added. The homogenate was mixed gently with a wide-bore pipette and incubated for 5 min on ice. The homogenate was then filtered through a 70-μm mesh strainer and centrifuged at 500×
CUT&RUN
For CUT&RUN, 500,000 nuclei were washed 1× with CUT&RUN wash buffer (20 mM HEPES, pH 7.5, 150 mM NaCl, 0.5 mM spermidine), bound to activated ConA beads, permeabilized in wash buffer (wash buffer + 0.002% digitonin), incubated with
CUT&RUN Library Prep was performed using the Illumina NovaSeq 6000 Sequencing System at the University of Kansas Medical Center – Genomics Core (Kansas City, KS). Fragmented input and immunoprecipitated chromatin (5 ng) were used to initiate the TruSeq ChIP Sample Prep Kit library preparation protocol with modifications for CUT&RUN sample input (Illumina Cat# IP-202-1012). The fragmented chromatin underwent end repair and 3’ adenylation prior to Illumina indexed adapter ligation. No gel size selection of the ligation product was performed. Ten cycles of PCR amplification with a modified extension time of 10 seconds were performed using Illumina adaptor-specific priming with final library purification using KAPA Pure magnetic bead purification (KAPA Cat# KK8002).
Library validation was performed using a DNA 1000 Assay Kit (Agilent Technologies 5067-1504) on an Agilent TapeStation 4200. The concentration of each library was determined by qPCR using a Roche LightCycler 96 using FastStart Essential DNA Green Master Mix (Roche 06402712001) and KAPA Library Quant (Illumina) DNA Standards 1–6 (KAPA Biosystems KK4903). The libraries were pooled based on equal molar amounts to 1.85 nM for multiplexed sequencing.
The pooled libraries were denatured with 0.2 N NaOH (0.04 N final concentration) and neutralized with 400 mM Tris-HCl pH 8.0. Dilution of the pooled libraries to 370 pM was performed in the sample tube on the instrument, after which onboard clonal clustering of the patterned flow cell was performed using a NovaSeq 6000 S1 Reagent Kit v1.5 (200 cycle) (Illumina 20028318). A 2×101 cycle sequencing profile with dual index reads was completed using the following sequence profile: Read 1–101 cycles×Index Read 1–6 cycles×Index Read 2– 0 cycles×Read 2–101 cycles. Following collection, the sequence data were converted from the.bcl file format to the.fastq file format using bcl2fastq software and demultiplexed into individual sequences for data distribution using a secure FTP site or Illumina BaseSpace for further downstream analysis.
CUT&RUN data processing and analysis
TrimGalore was used to trim the raw.fastq files to remove adaptors. The trimmed.fastq files were then mapped to the hg19 genome utilizing Bowtie2. The same procedure was run to align the.fastq files to a masked Saccharomyces cerevisiae v3 (sacCer3) genome for spike-in control DNA, which was also downloaded from the University of California Santa Cruz (UCSC) (https://genome.ucsc.edu/). Sambamba was then used to remove duplicates. For IGV visualization, deepTools was used with the “bamCoverage” function to generate normalized CPM.bw files. For peak calling, the recently developed SEACR was utilized and run in “relaxed’ mode to produce peak files, as the BED files used were already normalized to the number of yeast spike-in reads. DeepTools was further applied for heatmap visualization with the functions “computeMatrix” and “plotHeatmap”. The “dba.peakset” function of the Diffbind R package was further applied to identify overlapping peaks on the basis of the bound peaks. The links of CUT&RUN peaks and their related genes were established with the “annotatePeaks.pl” function in Homer. Motif enrichment analyses were performed using the “findMotifsGenome.pl” function in Homer, leading to known enrichment results and de novo enrichment results, and the latter were chosen in this study.
RESULTS
Upregulated γ H2AX expression in AD brains
We obtained autopsy brain tissue (frontal cortex) from postmortem human AD and nondemented (ND) patients through the KU Alzheimer’s Disease Research Center Neuropathology Core. The age, sex, and diagnostic information of these patients is shown in Fig. 1A. First, we used
CUT&RUN to map DSB sites in ND and AD samples
DSB formation induces rapid phosphorylation of histone H2AX at Ser139 (
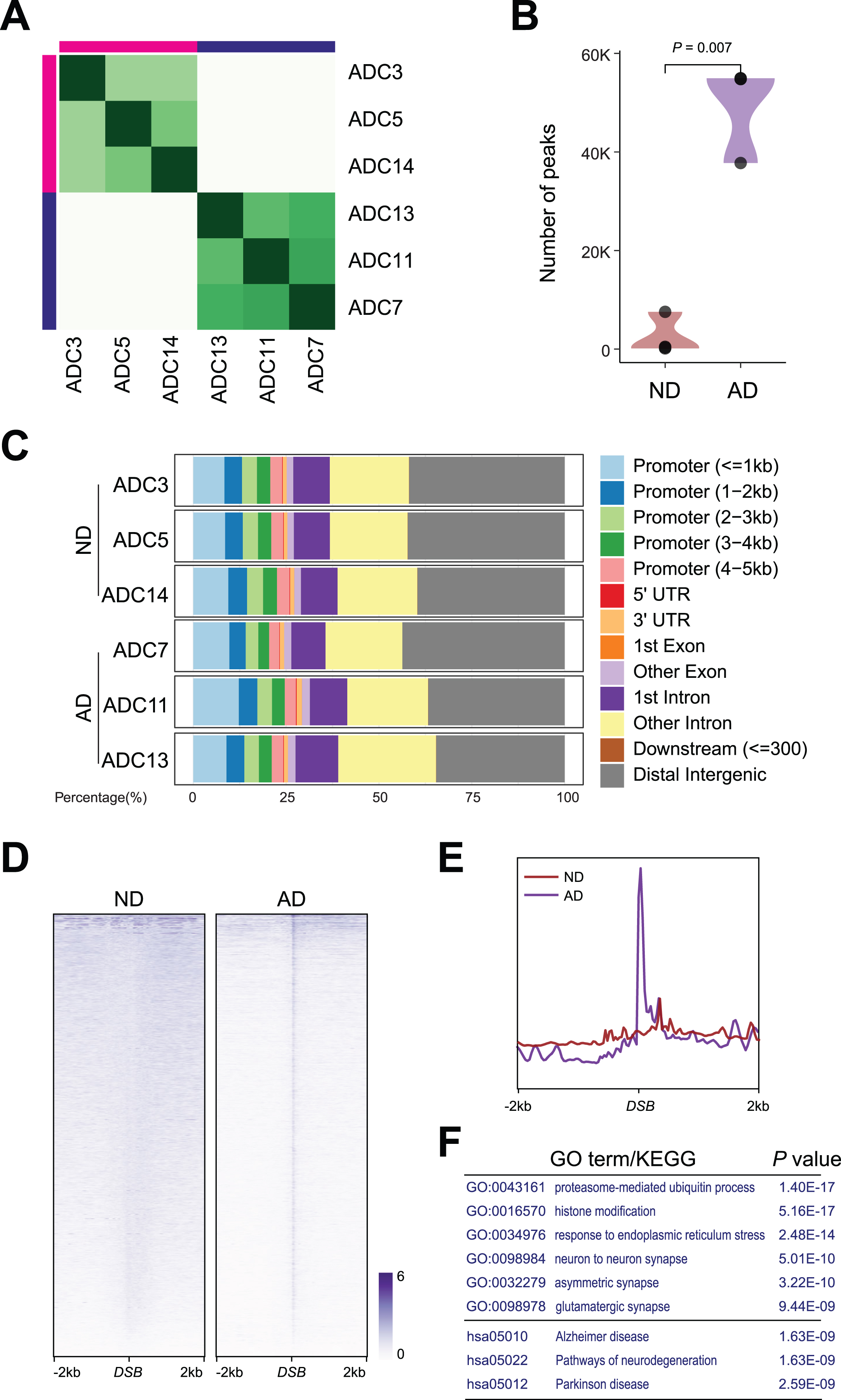
DSB sites in ND and AD samples. A) Heatmap representation of Diffbind clustering of the indicated CUT&RUN experiments. B) Violin plot showing number of peaks between AD and ND. C) Distribution of differential
The ratio of the genomic distribution of DSBs was comparable in both ND and AD samples.
Importantly,
Distribution of DSBs in ND and AD samples
We further examined the enrichment profiles of
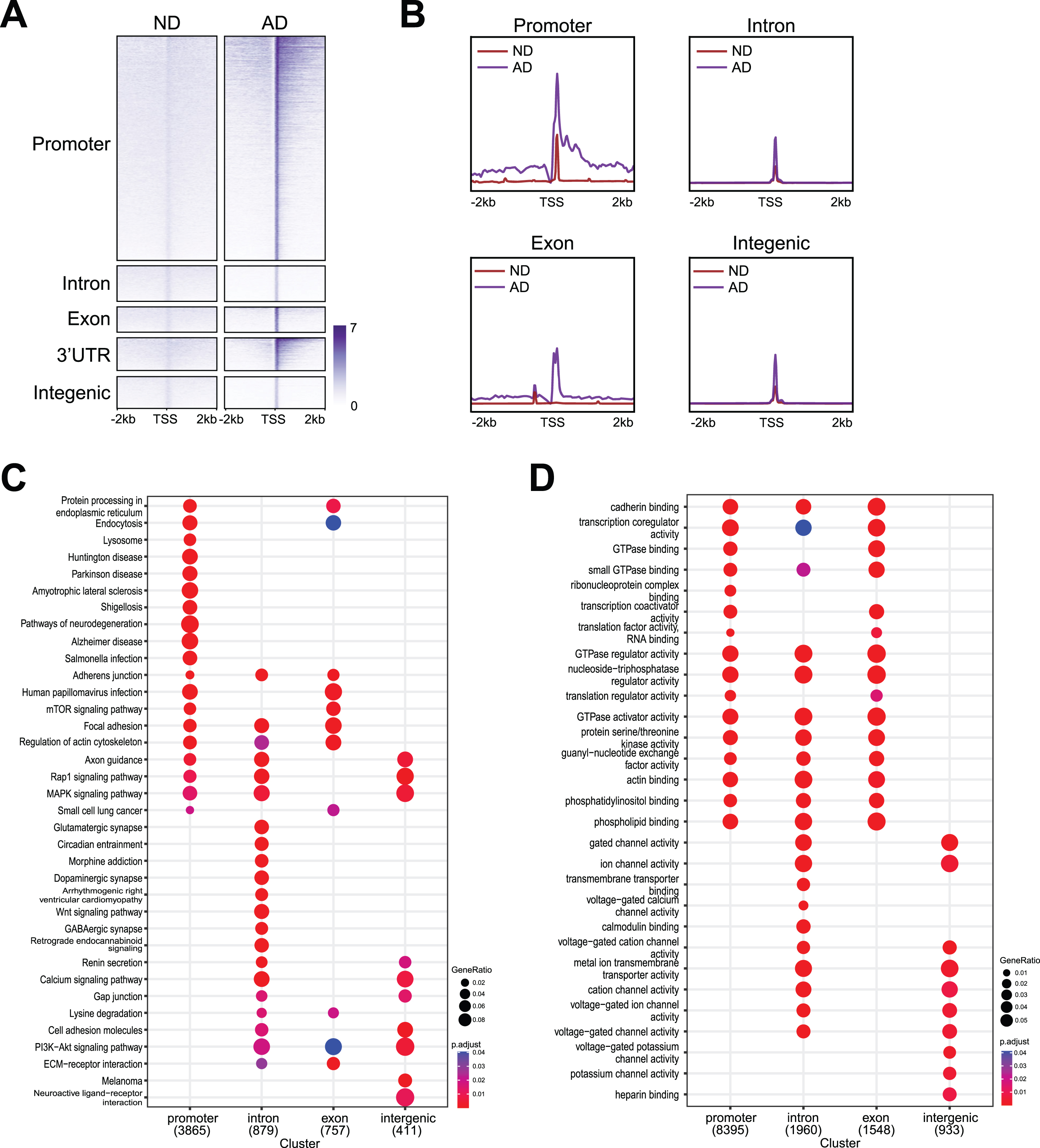
Genomic distribution of DSBs with functional analysis. A) Heatmaps showing distribution of DSBs binding at promoter, intron, exon, 3’UTR, and intergenic. B) Distribution of DSBs in a ±2 kb window of
Interestingly, our data showed that genes enriched for different functions were susceptible to DSBs at different genomic regions (Fig. 3C, D). For example, functional pathway identification analysis (Fig. 3C) demonstrated that genes that harbored DSBs at the promoters encode pathways related to neurodegeneration (e.g.,
Differential peak analysis
To identify differential DSBs in AD and ND samples, we performed differential peak analysis, and the result is shown by a volcano plot (Fig. 4A). We analyzed the DNA sequences around DSB sites near the TSSs of these genes for sequences related to transcription factor-binding sites by using Homer. Interestingly, our analysis showed that the binding site for PRDM9 was enriched in genes that had increased
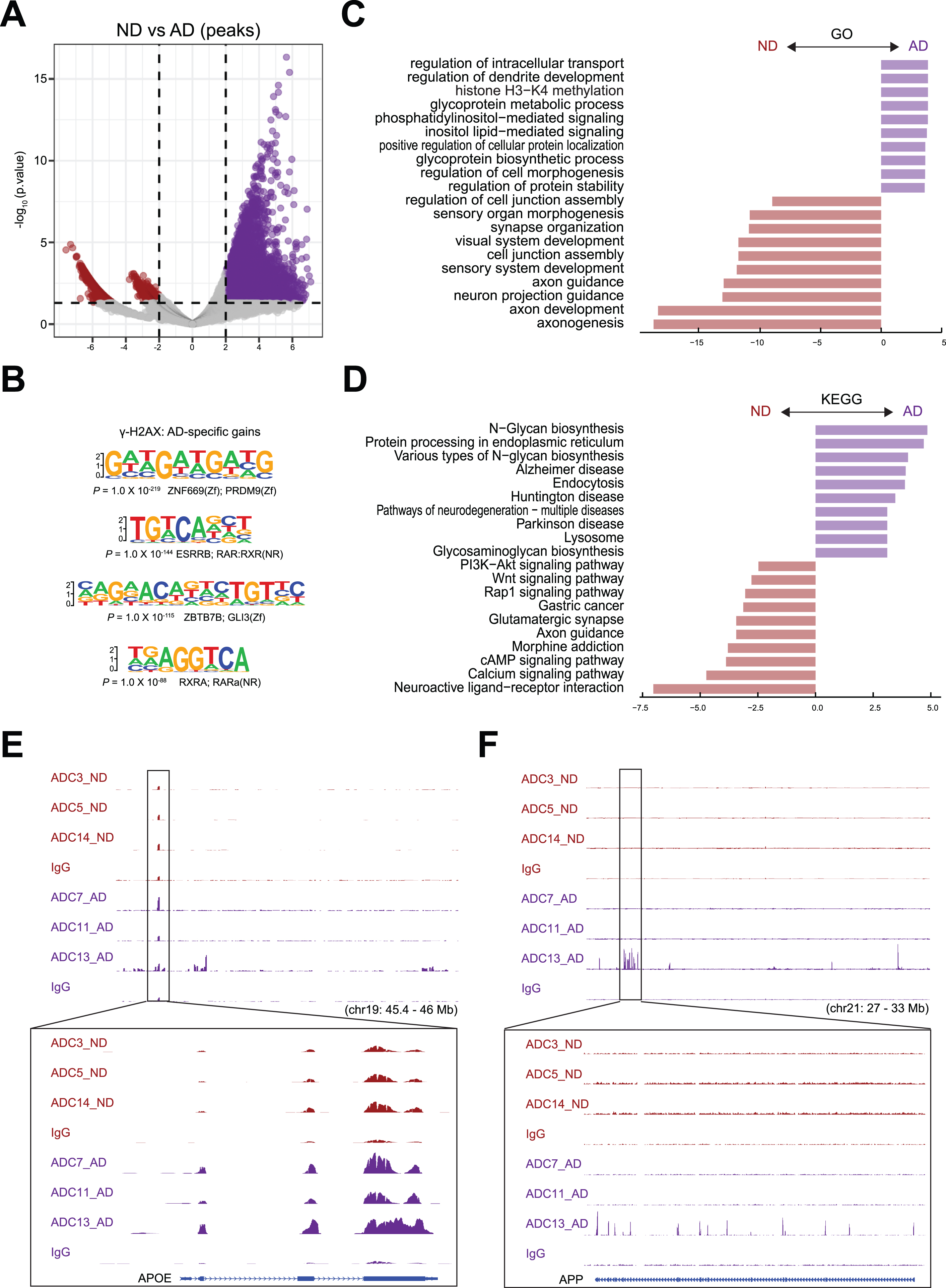
Differential peak analysis for DSBs. A) Differentially enriched
After annotating peaks to genes, we identified 12,345 genes with upregulated
Representative peaks at the known functional AD genes
Correlation of DSBs with upregulated gene expression in the AD brain
To test whether genes with increased numbers of DSBs exhibit dysregulated expression in AD, we analyzed their expression using a published RNA-seq dataset that contains data for 10 individual ND samples and 12 individual AD samples [34]. To identify the potential impact of DSBs on genes enriched for specific biological functions, we examined the expression of the genes according to the top 10 functional signatures with increased DSBs in AD samples (Fig. 4C). We found that the expression of genes in these functional signatures was upregulated in AD samples (Fig. 5A, upper panel). Conversely, genes in the top 10 functional signatures that lost
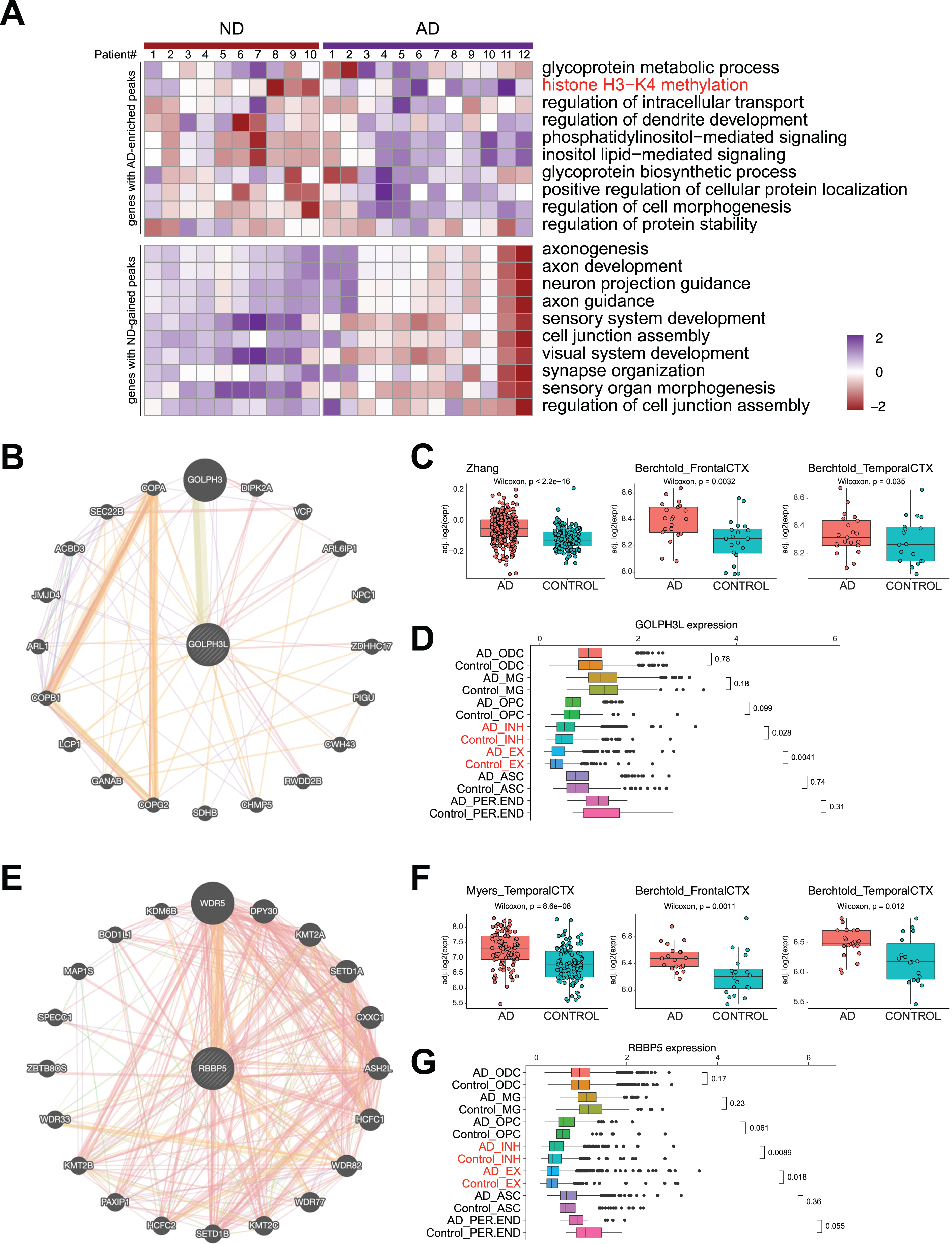
DSBs correlate with aberrant gene expression in AD. A) Heatmap showing AD and ND-associated signature level in AD and ND. B) STRING analysis of glycoprotein metabolic process-related genes revealing a protein interaction network with GOLPH3L. C) Boxplots showing GOLPH3L expression in three published RNA-seq datasets of AD and ND. D) Boxplots showing GOLPH3L expression in single cell RNA-seq datasets of AD and ND. E) STRING analysis of histone H3– K4 methylation-related genes revealing a protein interaction network with RBBP5. F) Boxplots showing RBBP5 expression in three published RNA-seq datasets of AD and ND. G) Boxplots showing RBBP5 expression in single cell RNA-seq datasets of AD and ND.
Next, we used STRING analysis [35] to identify the “hub” protein for the top GO functions, “glycoprotein metabolic process” and “histone H3-K4 methylation”. Analysis of the proteins encoded by genes in the “glycoprotein metabolic process” function revealed an interaction network of 20 proteins, of which GOLPH3L was located at the center (Fig. 5B). GOLPH3L is localized at the Golgi stack and may have a regulatory role in Golgi trafficking [36]. We examined
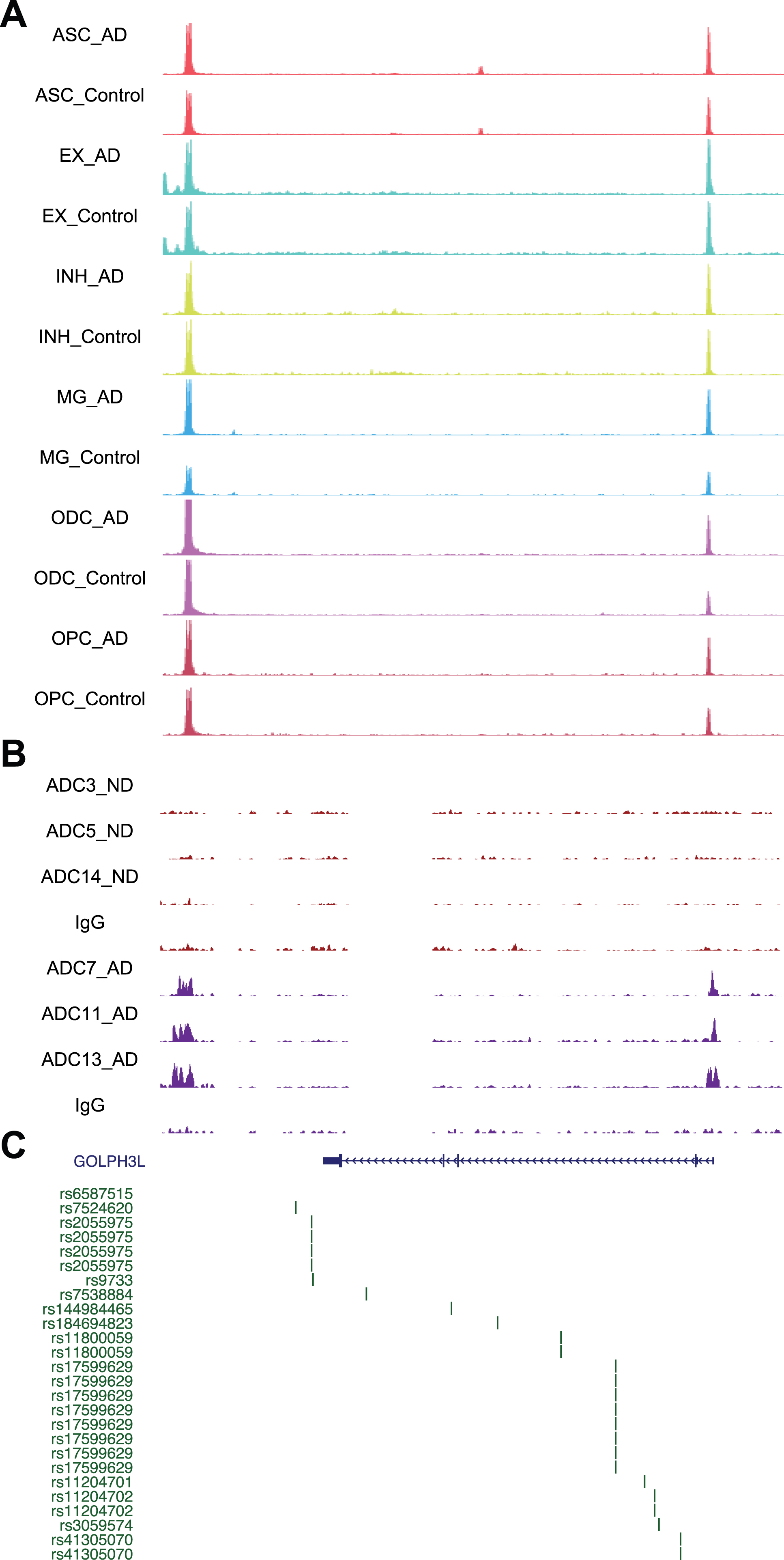
Chromatin accessibility, DSB sites, and GWAS for
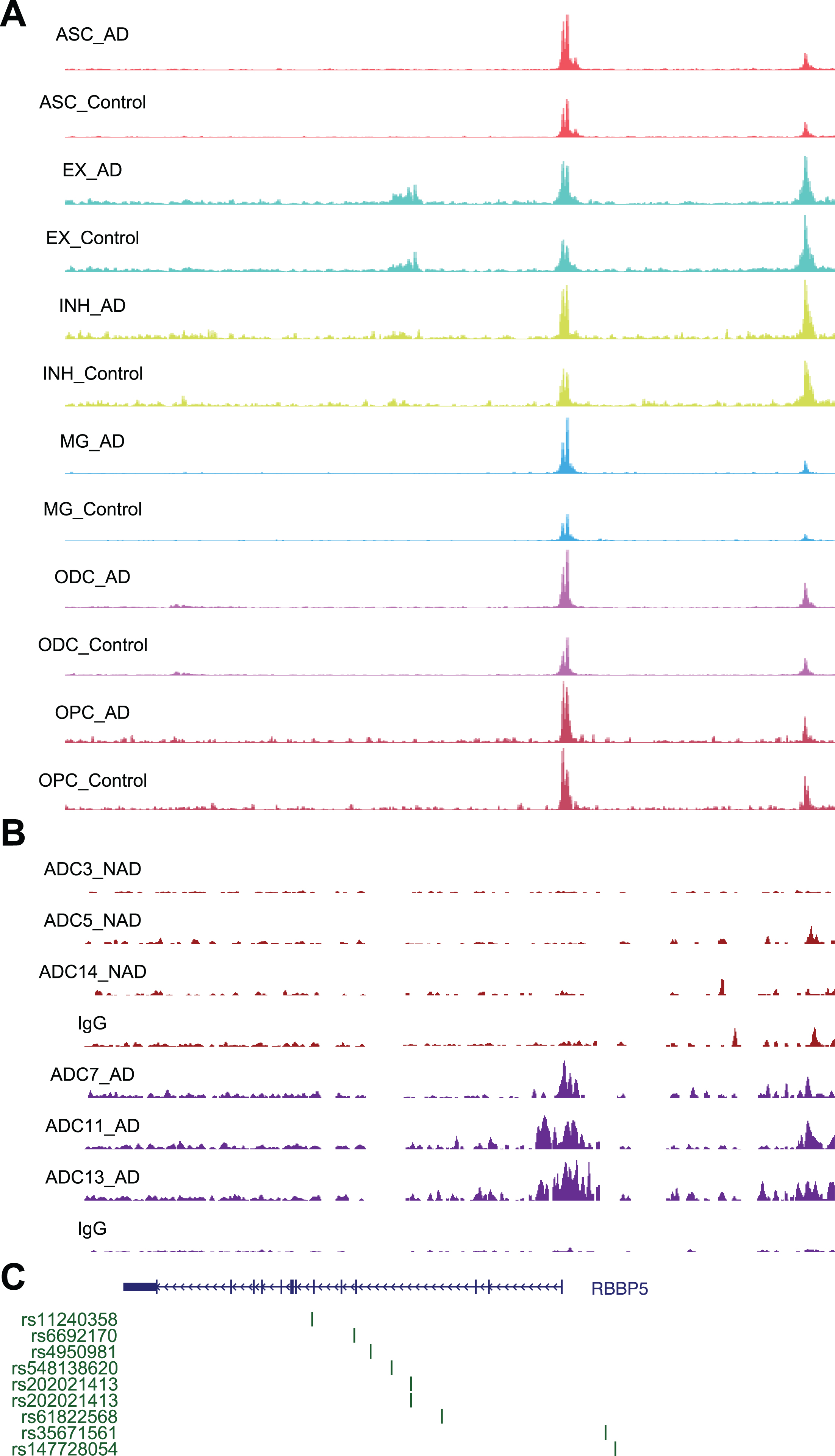
Chromatin accessibility, DSB sites, and GWAS for
Using a similar approach, we performed STRING analysis of proteins encoded by genes with the “histone H3-K4 methylation” function and revealed an interaction network of 20 proteins, with RBBP5 located at the center of the network (Fig. 5E). RBBP5 encodes a retinoblastoma protein-binding protein that regulates cell proliferation and is related to genes involved in transcription cis-regulatory region binding and histone H3K4 methyltransferase activity [43]. Comparison of
DISCUSSION
To our knowledge, this is the first report of an AD brain DSB landscape. Our data show that AD brains contain 18 times more DSBs than control brains and that the AD brain pattern of DSBs differs from the control brain pattern. In conjunction with published genome, epigenome, and transcriptome analyses, our data reveal that aberrant DSB formation correlates with AD-associated single-nucleotide polymorphisms (SNPs), increased chromatin accessibility, and upregulated gene expression. Thus, our data suggest that in AD, accumulation of DSBs at ectopic genomic loci contributes to aberrant upregulation of gene expression. The limitations of this study include a small sample size (3 ND and 3 AD) and a lack of female brain tissues. Thus, sex-specific differences in aberrant AD DSB formation were not addressed. In addition, gH2AX often spread across the region flanking DSBs for several hundred kilobases, particularly in euchromatin (spread less efficiently in heterochromatin), which prevent the precise and unbiased mapping of DSBsby gH2AX.
It is still not clear whether AD pathology itself causes DNA damage or whether accumulation of DNA damage contributes to the development of AD pathology. Studies have shown that several factors associated with AD pathology, such as Aβ and tau protein, can induce DNA damage in neurons [12, 45]. Aβ has been shown to damage DNA by generating free radicals and causing oxidative stress [46], while tau protein can indirectly contribute to DNA damage by disrupting the transport of molecules within neurons and by impairing DSB repair machinery [12, 47]. Additionally, inflammation, which is a hallmark of AD pathology [48], and mitochondrial dysfunction can also contribute to DNA damage by generating reactive oxygen species (ROS) [5, 49]. On the other hand, it is plausible that the accumulation of DNA damage occurs before the onset of AD. In this case, aging itself is a factor for the accumulation of DNA breaks [4, 50], and aging is known to be the most important risk factor for AD. Thus, the relationship between AD pathology and DNA damage is likely to be complex andreciprocal.
Our data show that genes bearing DSBs in AD exhibit increased chromatin accessibility. While DSBs are typically thought of as a form of DNA damage that jeopardizes genome integrity, our study shows that a consequence of ectopic DSBs in the AD brain is upregulated gene expression. This observation adds to the body of evidence that DSBs activate gene expression [51]. When a DSB occurs in the vicinity of a gene, it can trigger a cascade of molecular events that leads to the recruitment of repair proteins in the DDR pathway to the site of the break. These proteins can also recruit other proteins, such as histone modifiers and chromatin-remodeling factors, that can modify the structure of chromatin, resulting in opening of the chromatin structure and making the gene more accessible to transcription factors and RNA polymerase. This can ultimately lead to activation of the gene and to an increase in the expression of its encoded protein. In particular, in brain cells, this process is mediated by TOP2B to promote early response gene expression during physiological processes, including learning and memory, via triggering of the release of CTCF from cohesins and tethering of enhancer and promoter regions, which thereby activates RNA polymerase II activity [20]. Whether a similar mechanism underlies DSB-correlated upregulation of gene expression is not known and is worth furtherinvestigation.
The accumulation of DSBs likely results from both increased
Our data showed that the binding site for PRDM9 was enriched in genes that had increased
Footnotes
ACKNOWLEDGMENTS
We thank Cynthia Shaddy-Gouvion at KUMC Histopathology Core for preparing frozen autopsied tissues and FFPE sections.
FUNDING
This study was supported by R01-HD103888, 3R01-HD103888-03S1, KU ADRC P30 AG072973, the KU School of Medicine, and the Landon Center on Aging (N.W.). This project was supported by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number P20 GM103418 (X.Z.). The content is solely the responsibility of the authors and does not necessarily represent the official views of these funders. Kansas Intellectual and Developmental Disabilities Research Center (NIH U54 HD 090216), the Molecular Regulation of Cell Development and Differentiation – COBRE (P30 GM122731-03) - the NIH S10 High-End Instrumentation Grant (NIH S10OD021743) and the Frontiers CTSA grant (UL1TR002366) at the University of Kansas Medical Center, Kansas City, KS, USA.
CONFLICT OF INTEREST
Russell Swerdlow is an Editorial Board Member of this journal but was not involved in the peer-review process nor had access to any information regarding its peer-review.
The authors have no conflict of interest to report.
DATA AVAILABILITY
The dataset used in the current study has been uploaded and will be made publicly available at NCBI (GSE231568).
