Abstract
Background:
DNA methylation (DNAm), an epigenetic mark reflecting both inherited and environmental influences, has shown promise for Alzheimer’s disease (AD) prediction.
Objective:
Testing long-term predictive ability (>15 years) of existing DNAm-based epigenetic age acceleration (EAA) measures and identifying novel early blood-based DNAm AD-prediction biomarkers.
Methods:
EAA measures calculated from Illumina EPIC data from blood were tested with linear mixed-effects models (LMMs) in a longitudinal case-control sample (50 late-onset AD cases; 51 matched controls) with prospective data up to 16 years before clinical onset, and post-onset follow-up. Novel DNAm biomarkers were generated with epigenome-wide LMMs, and Sparse Partial Least Squares Discriminant Analysis applied at pre- (10–16 years), and post-AD-onset time-points.
Results:
EAA did not differentiate cases from controls during the follow-up time (
INTRODUCTION
Alzheimer’s disease (AD) pathology in the form of tau and amyloid-β (Aβ) starts accumulating years or decades before clinical symptoms [1], making early prediction critical. Recently, novel blood-based markers, for instance those reflecting different species of phosphorylated tau in plasma have shown promise for AD prediction [2, 3]. In addition, the study of polygenic risk-scores (PRS) for AD has come to explain an increasing proportion of AD heritability [4]. However, it is essential to study further non-invasive biomarkers, besides the neuropathology and genetic biomarkers, due to the highly heterogeneous and multifactorial nature of AD, which is influenced by multiple lifestyle and environmental factors across life [5–7].
DNA methylation (DNAm), an epigenetic process measurable in blood, is of increasing interest for AD prediction due to its potential to capture both inherited and acquired disease risk through the life course [8–10]. DNAm is a mechanism that can regulate gene expression, by binding of methyl groups to DNA nucleotides, most commonly at CpG sites (cytosine-guanine nucleotide pairs). DNAm patterns are partially heritable, influenced by lifestyle and environmental factors, and change in aging [9, 12]. Epigenetic biomarkers based on DNAm are also of interest for prediction of biological aging, aging-related diseases, and mortality [13, 14]. Of interest is that DNAm can bring potential mechanistic insights into disease etiology thanks to its potential to affect gene expression [15]. In AD, DNAm alterations located in gene regions with well-established roles in AD have been identified in both brain tissue and blood, including apolipoprotein E (
The relative consistency of DNAm changes with aging enables accurate estimation of chronological age based on DNAm patterns using machine learning methods, resulting in epigenetic age-estimators, or “clocks” [13, 27]. A higher epigenetic age than chronological age as estimated by epigenetic clocks, so-called epigenetic age acceleration (EAA), has been found to associate with cellular and physical aging, mortality [14, 28], several age-related diseases including Parkinson’s disease, some cancer types, coronary heart disease [13, 14] as well as AD [29]. Since advanced age is the largest risk factor for late-onset sporadic AD [30], it is of relevance to investigate whether measures capturing accelerated biological aging are predictive of AD. In studies of AD, EAA in brain tissue correlates with neuropathological biomarkers of AD, including brain tau and amyloid load, as well as with decline in cognition [29, 32]. The fact that AD-related DNAm alterations in brain tissue are enriched in sites displaying aging-related changes, with a concordance of effect direction, also suggests that EAA may be of relevance for AD prediction or disease progression [33]. However, previous studies have found mixed results with regards to the utility of EAA in blood in the context of dementia [8, 34–36], and there is a scarcity of longitudinal studies tracking EAA over longer time-periods preceding disease onset. Thus, although EAA in blood can successfully predict a number of aging-related outcomes, its predictive value for AD remains to be established.
The current study aimed at testing the hypothesis of accelerated epigenetic aging in blood in clinical late-onset AD using a longitudinal case-control design, with up to 20 years of longitudinal follow-up data from the same carefully characterized individuals from a Swedish population-based study [37, 38]. We tested three validated epigenetic clocks thought to capture different aspects of cellular and physiological aging [14, 40], as well as a DNAm biomarker designed to capture the rate of biological aging across multiple organ systems [41, 42]. In addition, we investigated whether we could identify sets of CpG sites predictive of AD up to 16 years before clinical AD onset, using both univariate and multivariate statistical methods. First, using CpG-wise univariate linear mixed effects models (LMMs), we leveraged our longitudinal data to identify CpG sites that stably differentiated AD cases from controls across the entire follow-up period, with the rationale that such sites may be particularly robust biomarkers. Secondly, we used a multivariate supervised machine-learning method for variable selection by sparse partial least squares discriminant analysis (sPLS-DA), which accounts for the covariance among CpG sites, and permits selecting sets of CpGs able to conjunctly discriminate AD cases from controls. By applying the sPLS-DA in specific time subsamples, long before (16-10 years) and after AD onset, we tested whether different sets of CpGs were predictive at different disease stages, substantially expanding the time frame of previous studies [43, 44]. Finally, we tested whether the findings from our primary study cohort replicated in a cross-sectional Australian sample [45], or corresponded to previously reported DNAm differences in AD based on a literature search. An overview of the study design is provided in Supplementary Figure 1.
MATERIALS AND METHODS
Study populations
Our main study sample is from the Betula study, a longitudinal, prospective population-based study, aimed at investigating health, aging, cognition and dementia. It comprises 4,425 participants followed for up to 25 years, with cognitive, health-related, social, and biological assessments [37, 46]. As fully described previously [38, 46], the recruited participants were native Swedish speakers with no dementia, congenital or acquired intellectual disabilities at study entry, nor severe impairments in hearing or vision. The study was initiated in 1988 with consecutive follow-ups at five years intervals (T1–T6 test waves), evaluating cognitive status and dementia at each time point. Blood sampling was carried out at test waves T2–T6 (Supplementary Table 1).
The Australian Imaging, Biomarker & Lifestyle Study (AIBL) was used as an independent external cohort to validate the analyses. Data was collected by the AIBL study group. AIBL study methodology has been reported previously [45, 47]. The AIBL study aimed at recruiting and characterizing 1000 participants, including at least 200 AD cases, 100 mild cognitive impairment (MCI) cases, and 700 healthy controls (aimed at including both carriers and non-carriers of the
This research was conducted in accord with the Declaration of Helsinki. The Betula study has been approved by the Regional Ethical Review Board in Umeå and the Swedish Ethical Review Authority, and written consent for study participation was obtained from each participant. Informed written consent was also given by all AIBL volunteers, and ethics approvals for the study were obtained from ethics committees of Austin Health, St. Vincent’s Health, Hollywood Private Hospital and Edith Cowan University.
Clinical characterization and dementia diagnosis assessments
The dementia diagnosis assessment in the Betula study has previously been described [37]. In brief, the diagnostic characterization was based on multiple sources of information and included each participant’s healthcare history as expressed in medical records, supplemented by relevant data from the repeated health and cognitive assessments that were part of the Betula study protocol. The dementia diagnoses were defined according to the Diagnostic and Statistical Manual of Mental Disorders 4th edition (DSM-IV) dementia classification. Participants diagnosed with AD showed representative symptoms of clinical AD, including an insidious onset and progressive cognitive decline.
For the AIBL sample, AD diagnoses were based on NINCDS-ADRDA Alzheimer’s Criteria (probable or possible), evaluating the impairment of memory, language, perceptual skills, attention, constructive abilities, orientation, problem solving, and functional abilities [48]. MCI diagnosis was based on Winblad et al., in which subjects previously diagnosed with MCI by a clinician additionally showed a score at least 1.5 standard deviation from the age-adjusted mean on one or more neuropsychological tasks [49].
Inclusion and exclusion criteria
The Betula sample inclusion criteria were having at least one blood sample ≥3 years prior to clinical AD onset, and at least one sample at or after onset, as well as a ≥5-year duration between samples. Only cases with an onset age of ≥65 years were considered. In total, 49 AD cases fulfilled these criteria. An additional two cases without post-onset samples, but with pre-onset samples >10 years prior to onset were added to enrich the sample with measurements collected long before onset. Thus, the final sample comprised 51 AD participants and 51 healthy age- and sex-matched controls subjects, born between 1909 and 1944. This selected sample comprised 20 AD cases and 20 matched controls with three longitudinal time-points, 29 AD cases and 29 matched controls with two longitudinal time-points, and 2 AD cases and 2 matched controls with one time-point long before AD onset. Controls were considered healthy when not diagnosed with AD or any other dementia subtypes, and not showing memory decline according to a previous classification model applied to the sample [50, 51]. The majority of the included controls (
The AIBL DNAm sample comprised 471 healthy controls, 94 MCI cases, and 161 AD cases (see sample demographic characteristics in Supplementary Table 2). We excluded subjects younger than 65 years of age (
DNAm analyses
DNA bisulfite conversion and methylation array analysis
DNA was extracted from peripheral blood (whole blood or buffy coat) collected at multiple blood sampling time points, by previously described methods in Betula [52] and AIBL [53]. In Betula, the DNA were sodium bisulfite converted using the EZ DNA methylation kit (Zymo Research, CA, USA) according to the manufacturer’s protocol. AIBL DNA methylation was obtained from the NCBI Gene Expression Omnibus GSE153712 [54]. Infinium Methylation EPIC BeadChip arrays (Illumina inc., San Diego, CA) were used in Betula and AIBL for methylation profiling of the bisulfite converted DNA. These arrays interrogate over 850 000 CpG sites across the genome at single-nucleotide resolution. For the Betula cohort the quality of the methylation data was assessed using the bead arrays controls reporter (Illumina) and the multiple samples from the same individual were confirmed using single-nucleotide polymorphisms (SNPs) included on the array. In both Betula and AIBL arrays, raw methylation data from the arrays (β-values) was extracted using the the
Epigenetic clocks estimation
Hannum’s DNAm age (71 CpGs) was originally calculated composing a weighted average (formed by regression coefficients) of CpGs, which then is transformed to DNAm age using a calibration function [39]. Horvath’s epigenetic clock (353 CpGs) is based on a similar regression model approach [40]. Hannum’s and Horvath’s clocks were constructed with the Illumina 450k methylation array, with 6 and 17 included CpGs, respectively, missing on the currently used Methylation EPIC array [13]. The PhenoAge clock (513 CpGs) was obtained by a penalized regression model that accounted for several disease risk biomarkers [14]. All CpGs used by the PhenoAge clock are available on the EPIC array. We also estimated the DNAm biomarker Dunedin Pace of Aging Calculated from the Epigenome (DunedinPACE) [41], an updated version of the DunedinPoAm clock designed to predict the longitudinal rate of change in 18 biomarkers from multiple organ systems across 12 years [42]. For the epigenetic clocks estimation, missing β-values were imputed by a K-nearest neighbor model.
To estimate epigenetic age acceleration/deceleration, delta epigenetic ages were used instead of age-acceleration residuals from linear regression-based estimation, since the longitudinal measures violate the assumption of independence of observations (see also [58]). Thus, after the estimation of Horvath, Hannum, and PhenoAge epigenetic age clocks, Δepigenetic ages were obtained by subtracting the chronological age from the epigenetic age. A positive Δepigenetic age indicates accelerated epigenetic aging (i.e., the individual is biologically older than their chronological age) and a negative Δepigenetic age indicates slower epigenetic aging (i.e., an individual is biologically younger than their chronological age). The raw estimated values were used for the DunedinPACE health clock, as it is not an age estimator. One healthy control subject presented values below 3 standard deviations from the mean Hannum, Horvath and PhenoAge clocks, and these outlier values were replaced by the second lowest value of the full sample to avoid exclusion and loss of data points, according to a previous study [59].
Covariates
For the Betula sample
Relative leukocyte telomere length (RTL) was compared to the novel DNAm biomarkers, as an established blood-based biomarker previously associated with increased AD incidence in non-
Statistical analyses
Generalized additive mixed models (GAMMs)
GAMMs were used to depict the longitudinal profile of the raw estimated DNAm clocks in cases and controls separately, in order to check for potential non-linear associations in the Betula sample. Subsequent analyses used linear models, as observed associations were highly linear. Unadjusted GAMM models were performed in R using the
Linear mixed-effects models
In the Betula sample, longitudinal changes in the DNAm biomarkers (Hannum, Horvath, PhenoAge, or DunedinPACE), and blood cell proportions were assessed employing LMMs. As we intended to access differential longitudinal changes between AD cases and controls, all LMMs include an interaction term between AD status as a binary indicator variable and time, calculated in years to clinical onset (year 0). The time-scale ranged from – 16 to 7, i.e., a 23-year follow-up duration was modelled. Chronological age was used as an alternative time-scale. The longitudinal measures from the same subject and the sex- and age-matched pairs were modelled as nested random effects to account for variability within these blocking variables, therefore there was no need to control the models for age and sex. In particular, subjects were nested within matched pairs such that each matched pair of subjects are unique to that pair. LMMs were performed in R using the
Univariate longitudinal analysis of differentially methylated sites
We used LMMs with individual CpG-sites as dependent variables to identify differentially methylated sites that stably discriminate AD cases from controls across the entire follow-up period (see the statistical analysis section above for a more detailed description of the LMMs). Models were adjusted for
As a sensitivity analysis, we additionally ran the LMMs on 78 filtered out meQTL sites that have previously been associated with AD [56, 57]. None of the sites fulfilled the
Multivariate analysis of AD-predicting sets of CpGs
Machine learning-based sparse partial least squares discriminant analysis (sPLS-DA) [68] with 685,514 CpGs was used to identify CpGs that together may differentiate AD from controls. Two different cross-sectional subsamples were used for the sPLS-DA analyses: 1) the ‘long before’ AD subsample, that comprised samples from 16 to 10 years before AD onset and their respective matched controls (21 AD cases and 19 controls); and 2) the ‘after AD’ subsample, comprising samples from the years of AD onset to 7 years after AD onset (47 AD cases and 49 controls). sPLS-DA combines variable selection (identifying the most predictive or discriminative CpGs using lasso penalization) and classification in a one-step procedure. The algorithm uses a linear transformation that converts the data into a reduced dimensional space, in which the principal components (PCs) are the estimated features that represent the reduced dimensions that best separate the labeled groups with the smallest error rate. The number of PCs and CpGs within the PCs was selected by the lowest obtained balanced error rate (BER) after within-sample cross-validation (3-fold repeated 50 times). The sPLS-DA analyses were performed by the
Logistic regression models
The ability of each DNAm clock and novel DNAm biomarker to differentiate AD cases from controls was evaluated in the Betula sample at baseline time-point of each participant on average 8 years before AD onset, by logistic regressions. Models were adjusted for the covariates
Cox proportional hazard regression models
The ability of each DNAm clock and novel DNAm biomarkers to predict the risk of AD was evaluated at baseline time-point of each participant, on average 8 years before AD onset, by Cox regression. Models were adjusted for the confounders
Internal validation analyses
C-statistics of the logistic regression models were used to compare the discriminatory accuracy of the novel DNAm biomarkers estimated in Betula with the established biomarker
External validation analyses
Logistic regressions and their respective area under the receiver operating characteristic (ROC) curves AUC evaluating models’ discriminatory accuracy were used to validate the novel DNAm biomarkers in the AIBL validation sample. As for Betula, logistic models were adjusted for
Enrichment analysis
Tests of enrichment of CpGs-associated genes in AD or AD-related pathways by ‘Pathway’, ‘Disease’, and ‘Human Phenotype’ was performed by protein function analysis using Toppgene (https://toppgene.cchmc.org/enrichment.jsp). Enrichment by ‘disease biomarker networks’, and ‘diseases (by biomarkers)’ was performed with GeneGo MetaCoreTM software (https://portal.genego.com/).
RESULTS
Sample characteristics
The Betula and AIBL samples selected for the DNAm analysis were compared to investigate potential sample differences (Table 1). The percentage of males was similar between cases and controls within the samples, but higher in AIBL (∼43%) than in Betula cases (∼18%) (Table 1). In Betula, the proportion of
Demographic characteristics of the Betula and AIBL study populations
Data are expressed as number (percentage) or median (min-max). AD, Alzheimer’s disease. APOE , apolipoprotein E. MCI, mild cognitive impairment. *Age at study entry for Betula, and age at blood sampling for AIBL.
Demographic characteristics of the Betula and AIBL study populations
Data are expressed as number (percentage) or median (min-max). AD, Alzheimer’s disease.
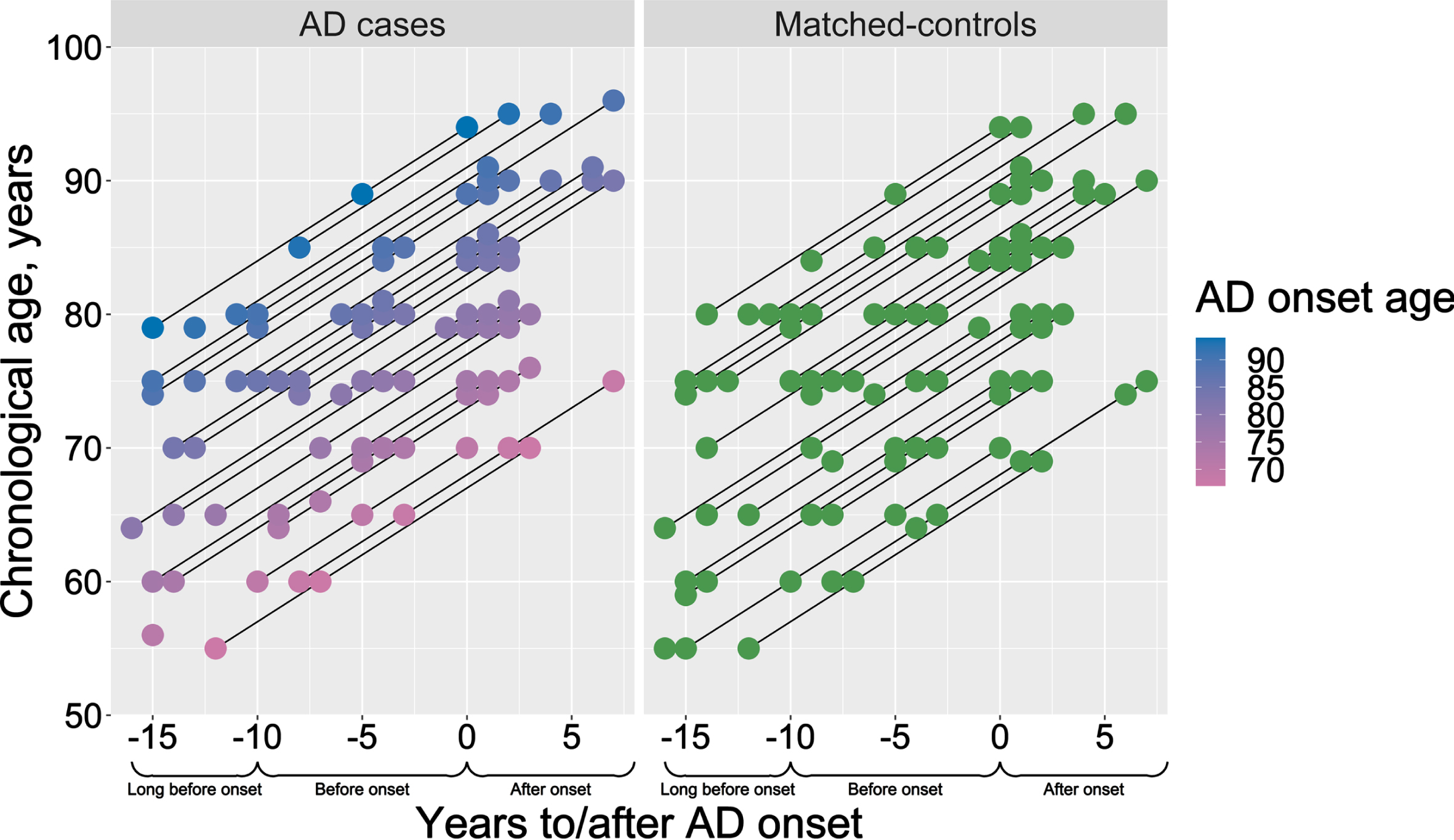
Description of the study design. Chronological age and AD onset age of the selected AD cases (pink-to-blue) and their respective sex- and age-matched controls (green). Y-axis presents the participants’ chronological age at blood sampling, and the color scale bar represents the age of AD onset. X-axis represents the time-scale, where blood samples for the DNAm analysis were selected aiming three time-points: long before (–16 to –10 years before AD onset), before (–9 to –3 years before AD onset) and after AD onset (0 to 7 years after AD onset).
In Betula, among health and lifestyle variables, BMI was significantly lower in the AD cases after clinical onset but the remaining of the considered covariates did not differ between the study groups at baseline (Supplementary Table 1).
Longitudinal changes in blood cell proportions
Prior to biomarker analyses, LMMs were used to evaluate whether aging or AD were associated with longitudinal changes in estimated blood cells proportions (Supplementary Table 5 and Supplementary Figure 3) in the Betula sample. The AD cases had a faster rate of increase in the NK cell proportion associated with increased chronological age (interaction beta coefficient = 0.0006,
In Betula, the estimated Hannum (Fig. 2a), Horvath (Fig. 2b), and PhenoAge (Fig. 2c) DNAm age clocks showed high and approximately linear associations with chronological age at blood sampling (
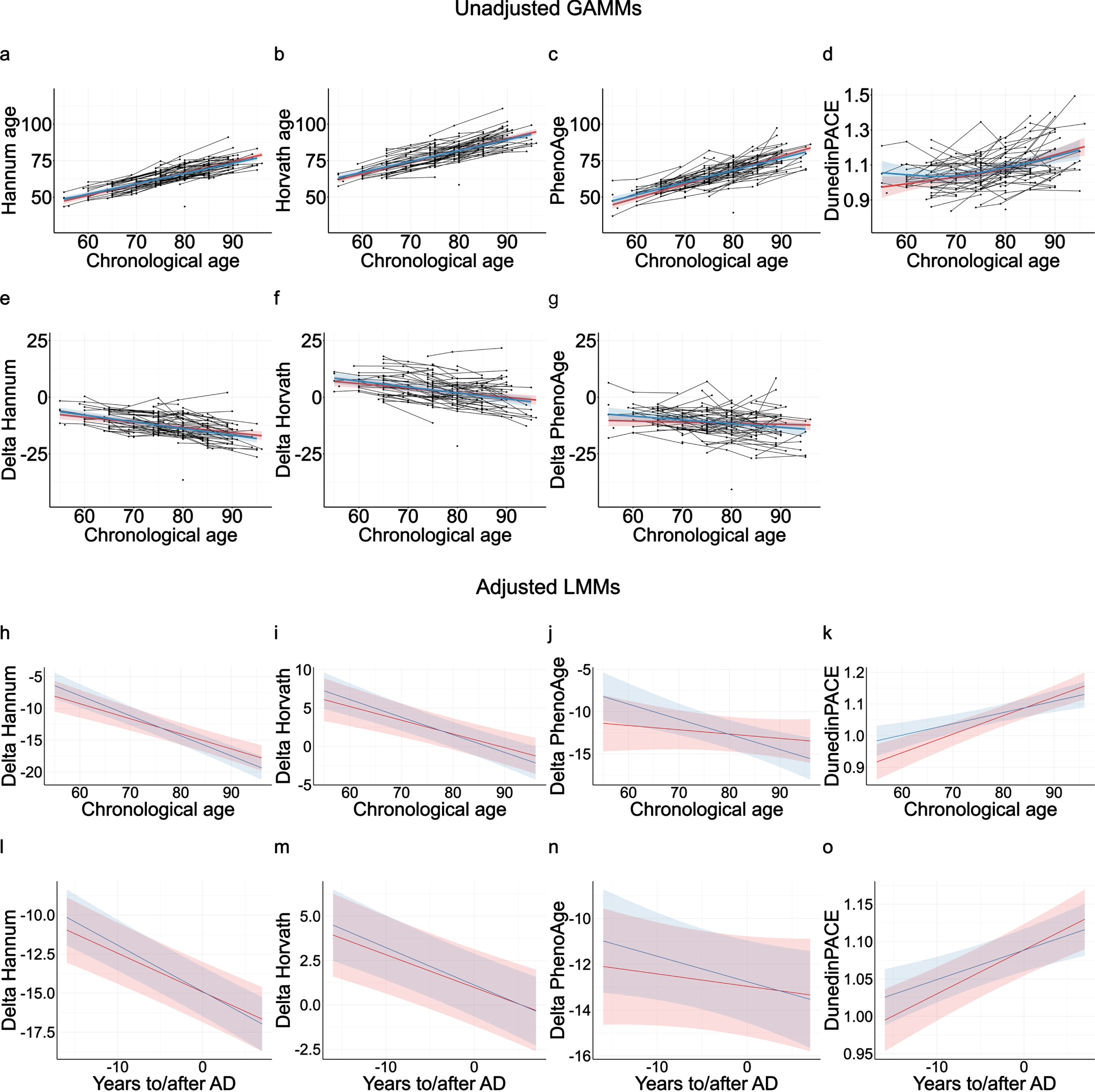
Longitudinal analyses of DNAm clocks in AD cases (red) and matched-controls (blue). Unadjusted generalized additive mixed models (GAMMs) of the raw estimated DNAm clocks (a–d) and Δages (e–g) with chronological age as the time-scale. Note negative association between Δage and age, indicating a deacceleration of epigenetic ages at higher chronological age for both cases and controls. Linear mixed-effects models (LMMs) of the Δages and the DunedinPACE clock with chronological age (h–k) and time to/after AD onset (l–o) as time-scales. LMMs were adjusted by
Delta ages were used to capture EAA, i.e., whether subjects were epigenetic older or younger than expected from their chronological ages. ΔHannum age (Fig. 2e), ΔHorvath age (Fig. 2f), and ΔPhenoAge (Fig. 2g) did not differ significantly between AD cases and controls in the unadjusted LMMs, and no evidence was obtained for differential longitudinal rates of epigenetic aging between cases and controls (
LMMs adjusted for additional covariates,
We additionally ran a supplementary set of analyses with age at AD onset instead of chronological age as a covariate to test for potential differences driven by age of onset of the cases. However, due to the sampling scheme in this study, which prioritized cases with a long follow-up time (i.e., available blood samples long before AD onset), chronological age at blood-sampling was highly correlated with age at onset (
When estimated in the AIBL cohort, the DNAm clocks were in accordance with the null findings in the Betula sample, none of the estimated delta ages nor the DunedinPACE clock were able to significantly discriminate MCI (
Longitudinal AD panel of differentially methylated sites is predictive of AD 8 years before clinical onset
In the Betula sample, univariate LMMs were used to identify differentially methylated sites that significantly discriminated AD cases from controls longitudinally, i.e., across the study duration of 20 years. There was no indication of inflation, as suggested by the obtained genomic inflation factor (lambda) of 1.050. No CpGs survived FDR-correction, but the models identified 73 CpG sites that fulfilled our exploratory criteria of
Further characterization of the 73 identified sites showed that the median difference in methylation β-values between cases and controls was 4.4% (min 2.6%, max 20.3%) at baseline, and 6.1% (3.6–17.6%) after AD onset (only sites with beta coefficient≥|0.05| at AD onset, see Methods). A majority of the identified sites (57 of 73) were hypomethylated in the AD cases when compared with the controls, and from the 27 CpGs that had significant AD-by-time interactions, all but one had a negative coefficient, i.e., almost all had decreased methylation over time in AD (Supplementary Table 3). Moreover, 22 CpGs had significant longitudinal aging effect (i.e., change over time across the whole sample), of which 19 (86% of 22 CpGs) showed an increase in methylation over time (i.e., positive beta coefficients, but with low effect sizes).
Complementary enrichment analyses of the annotated genes associated with the longitudinal AD panel’s CpGs were performed. These analyses did not show significant enrichment in AD using GeneGo (‘Alzheimer disease core network’ false discovery rate (FDR) >0.1, and ‘Alzheimer disease, late onset’ FDR = 0.084) or AD-, neurodegeneration-, or inflammation-related pathways using Toppgene (FDRs >0.1).
Pre- and post-AD scores from sPLS-DA are predictive of AD 8 years before clinical onset
Multivariate analysis by sPLS-DA was used to select CpGs that significantly discriminate AD cases from matched-controls cross-sectionally, at two different time-points. Two sets of PCs were identified in the
Similar to the longitudinal AD panel, the DNA sites of the pre- and post-AD scores were predominantly hypomethylated in the AD cases (Supplementary Table 3). The median |Δ
β| of the pre-AD score CpGs was 1.9% (min 1.3%, max 10.4%) long before AD onset. In CpGs of the post-AD scores, the median |Δ
β| was 1.4% (min 0.3%, max 8.4%) after AD onset. In addition, 3 CpGs overlapped between the longitudinal AD panel and the post-AD scores, the cg03688665 in the gene body/promoter region of the mitogen-activated protein kinase 4 (
Enrichment analyses of the annotated genes associated with pre- and post-AD scores CpGs were performed and did not show significant enrichment in AD-associated genes using GeneGo (‘Alzheimer disease core network’ FDR >0.1, and ‘Alzheimer disease, late onset’ FDRs >0.1) or AD-, neurodegeneration-, or inflammation-related pathways using Toppgene (FDRs >0.1).
Finally, we also tested whether the novel DNAm panels and scores, measured at study baseline, explained unique variance when considered simultaneously in a Cox regression, along with age, sex, granulocyte proportion,
Internal validation analyses of the novel DNAm biomarkers
The DNAm biomarkers estimated from the baseline time-point of each participant, on average 8 years before onset, were compared in an internal validation analysis to test their discriminatory accuracy using C-statistics. Including the well-established biomarker
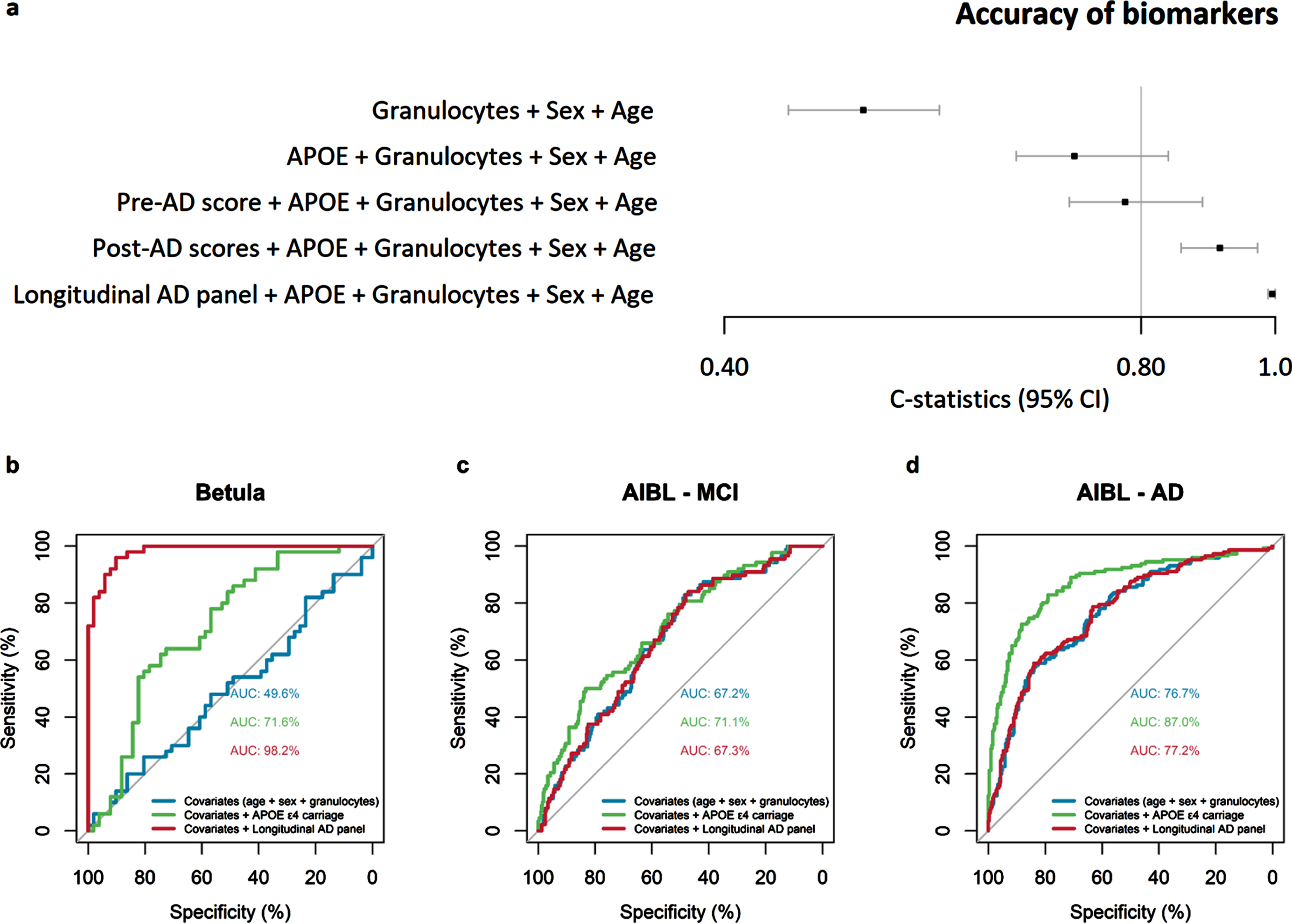
Accuracy of novel DNAm biomarkers in the Betula and The Australian Imaging, Biomarker & Lifestyle of Ageing (AIBL) samples. The forest plot shows the discriminatory accuracy (C-statistics) of the logistic regression models comparing AD cases versus matched-controls when including the biomarkers apolipoprotein E (
External validation analyses of the novel DNAm biomarkers
Next, we estimated the novel DNAm biomarkers in the AIBL sample among MCI cases (violin plots in Supplementary Figure 6) and AD cases (Supplementary Figure 7), and employed equivalent logistic regression models to compare their discriminatory ability between the Betula and the AIBL samples (Supplementary Tables 10 and 11). The novel DNAm biomarkers did not significantly discriminate MCI cases from controls (odds ratio [OR]: 0.98–1.16,
The discriminatory accuracy of the longitudinal AD panel was lower in the AIBL sample (see AUC Fig. 3c, d) compared to the Betula sample (Fig. 3b). In the Betula baseline subsample, on average 8 years before onset, including
We next explored whether the individual CpGs obtained in the longitudinal AD panel from the Betula univariate LMMs were associated with MCI and/or AD in the AIBL sample. A univariate logistic regression analysis was implemented, adjusted for
Previously identified differentially methylated sites overlap poorly across studies
We additionally conducted a literature search of studies published until August 2022 to investigate whether the CpGs of the longitudinal AD panel were reported in previous array-based epigenome-wide association study (EWAS) studies reporting a list of AD-associated CpGs in whole peripheral blood or white blood cells [10, 72–76]. However, no DNAm site selected by our analyses was previously reported (Supplementary Table 15). We also note that among these previous studies reporting several CpGs associated with AD exclusively in blood, only a few reported overlapping CpGs (Supplementary Table 15). From the 1000 [43], 477 [76], and 503 [73] CpGs previously reported, in total ten overlap in two different studies; however, six of these are reported with opposite directions of association with AD [43, 76]. Thus, only four CpGs had the same direction of association in AD between different studies [43, 76]. These were cg08787968 and cg01693350 (both in the gene body of the
DISCUSSION
The present study aimed at leveraging a unique longitudinal design with up to 16 years of prospective pre-diagnosis data from an age- and sex-matched case-control sample of clinical AD to test the hypothesis that EAA measures in blood are predictive of AD. A secondary aim was to identify new potentially predictive or diagnostic CpGs using novel longitudinal and machine learning methods. No evidence was obtained for EAA being predictive of AD in our longitudinal cohort, or a cross-sectional validation sample. Our longitudinal AD panel was the only novel biomarker identified in Betula that replicated in AIBL, although with negligible discriminatory value.
The absence of evidence for blood-based EAA measures as biomarkers of AD conversion is consistent with several recent studies on pre-symptomatic dementia cases [77, 78], MCI, or manifest dementia cases [79], as well as observed null associations with Aβ, p-tau, or t-tau status in cerebrospinal fluid [36]. Similarly, a previous AIBL study reported largely null findings between age-acceleration and cross-sectional and longitudinal measures of neuroimaging, cognition, and Aβ load; except for a robust cross-sectional association between the Hannum clock and hippocampal volume in cognitively unimpaired individuals with high brain Aβ load [35]. In contrast, a previous small study from our group suggested an association between DNAm age and dementia status in Betula (
In addition to our null findings for the epigenetic clocks, our attempts to identify novel panels/scores of DNAm biomarkers with predictive value for AD had limited success. Given the scarcity of replication evidenced in the literature, it is noteworthy that our longitudinal AD panel consisting of 73 CpG sites replicated nominally in the AIBL validation sample. This despite differences in sample characteristics, such as diagnostic criteria, sex, and
A limited replication of CpGs was further observed in our literature review of blood-based DNAm array studies of AD with published lists of CpGs. This literature review showed that only 4 out of 3,275 identified CpGs replicated with concordant direction of effects across the studies [43, 76]. These four CpGs were located in genes that participate in macrophage and immune responses, WNK4 and PTPN2 [88, 89], and the synaptic plasticity-related gene WT1 (2 CpGs) [90], respectively. The literature review did not identify previously replicated findings from candidate-gene or array-based DNAm studies, such as
Many factors, in addition to methodological and analytical ones [94, 95] may contribute to the limited replication of blood-based DNAm EWAS findings in AD. The heterogeneity of the disease itself may also be a main contributor, as AD comprises several subtypes concerning genetic factors, neuropathology, cognitive symptoms, and biological pathways [7, 97]. Different study cohorts may differ on important disease characteristics potentially influencing DNAm. For instance, the Betula and AIBL samples differed in
With regards to findings from DNAm studies, it is also relevant to consider the magnitude of the methylation differences observed. For instance, the mean methylation differences between Betula cases and controls after AD onset ranged from 4.7 to 8.2%, when considering the three validated CpGs of our longitudinal AD panel. This can be compared with a previous study in monozygotic and dizygotic AD-discordant twins with median methylation differences of 18.4% (min 15.9, max 29.7%) in blood examined by a different DNAm analysis method [99]. In previous studies in blood the median |Δ β| between AD cases and controls ranged from 1.0% to 4.6% (min 0, max 19.3%), indicating that methylation differences <5% are commonly reported [43, 92]. It is still unknown if methylation changes in this range lead to biologically significant changes in gene expression, but some evidence indicates biological relevance of subtle DNAm changes, for instance by resulting in protein isoform diversity [100]. Furthermore, in studies based on several cell types, small average changes in methylation levels may mask larger underlying changes in specific cell types. Regardless of functional consequences, small methylation differences could still provide valuable information as indicators rather than causes of dysregulated biological pathways, or serve a predictive role regardless of their functional relevance, as exemplified by the CpGs included in the epigenetic clocks that successfully predict other age-related disorders and mortality.
We note that within our longitudinal AD panel the majority of CpGs (78%), and all three validated CpGs, were cross-sectionally hypomethylated in AD cases compared to controls at the time of diagnosis. This is concordant with some recent studies on blood DNAm [10, 101] but not with others, that instead observed hypermethylation [43, 92]. Also longitudinally, almost all sites where DNAm rate of change significantly differed between cases and controls (i.e., time-by-AD interaction) in our longitudinal panel evidenced decreased methylation over time in AD. A decrease in global methylation levels with aging has been seen in several tissues [102], and thus it is possible that DNAm changes follow the same direction in aging as in AD, as previously proposed [10, 33].
Our current findings may have methodological and study design implications. Firstly, we applied a novel multivariate method, sPLS-DA, to try to identify CpGs in any parts of the genome that could jointly differentiate AD cases from controls. Although novel machine-learning or artificial intelligence methods are emerging for DNAm analyses [72], the majority of AD studies so far have relied on univariate methods (or differentially methylated regions across adjacent CpGs). In our data, the generated pre- and post-AD scores that were estimated based on sPLS-DA in the Betula sample were not replicated in AIBL. sPLS-DA is considered to be able to outperform other machine learning methods of feature selection due to its sparsity assumption [68], that aims at reducing the number of features that conjunctly discriminate the analyzed condition. This would help avoiding the selection of “noise” variables [103]. Even so, overfitting does happen [103], reinforcing the need for internal and external validation of the selected features. There was low concordance in the identified CpGs between the two different methods used in the current study, LMMs for the longitudinal AD panel and sPLS-DA for the pre- and post-AD scores. The fact that out-of-sample replication was seen only for the longitudinal panel, comprising stably differentially methylated sites between cases and controls across up to 26 years of follow-up, may speak to the superiority of longitudinal study designs for identifying DNAm-based disease biomarkers. This may be particularly true for small sample sizes such as in the current study, where the repeated measurements may act as intra-individual replication aiding identification of more reliable CpGs.
A strength of our study is that we considered estimated blood cell proportions in our analyses, which may otherwise confound DNAm estimates if differentially affected by the health status or age of the study participants [62, 104]. We also separately analyzed potential longitudinal differences in estimated blood cell counts in AD cases and controls. An increase in NK cells proportion over time was seen only in AD cases, again indicating a potential change in AD inflammation/immune response with aging. To the best of our knowledge, this is the first study of longitudinal changes in blood cell compositions in AD, and additional studies are needed to consolidate this finding. We also replicated some previous age-related changes in blood cell counts, like a decrease in subtypes of CD4+ T cells and B cells [105], and increase in monocytes [106].
Limitations
The limited AD sample size in the Betula study is an important limitation of our study, but uniqueness of the dataset with longitudinal retrospective blood samples up to 16 years prior to diagnosis nevertheless had potential to bring novel insights into the long-term predictive ability and temporal dynamics of blood-based DNAm biomarkers for AD. All available AD cases in the Betula study database who fulfilled the inclusion and exclusion criteria were included. Still, we acknowledge that this study was likely underpowered, particularly for the EWAS analyses, and lack of correction for multiple comparisons may have increased risk of false positive results. Another limitation was the lack neuropathological data or gold standard biomarkers (cerebrospinal fluid or positron emission tomography neuroimaging) to confirm AD diagnoses, which were not available in the Betula study, our primary cohort. Even so, the diagnostic evaluation integrated health-related, clinical, and cognitive assessments, resulting in a reliable clinical characterization [37, 45]. An inherent limitation for epigenetic clocks is the underestimation of epigenetic age in older subjects, which can lead to a loss of precision in older samples [70]. Finally, it is important to stress that the EPIC array covers only a small fraction, 850,000 of the ∼28 million CpG sites in the genome [107] and CpGs associated with AD could be outside the currently analyzed regions. Whole genome bisulfite sequencing has the potential to identify novel disease-associated CpGs.
Conclusions
The findings of this 16-year longitudinal study concur with the majority of recent observations in the literature that blood-based EAA measures developed so far are of limited value as AD biomarkers, particularly when other easily available indicators such as age, sex, and blood cell proportions are accounted for. Tentatively, inflammation and immune-system related processed may be reflected in DNAm patterns in AD, but overall our findings underscore the difficulty of identifying replicable epigenome-wide DNAm alterations that can reliably distinguish AD cases from controls beyond known markers such as
Footnotes
ACKNOWLEDGMENTS
We thank all the Betula study participants, and SciLifeLab Uppsala for the array analysis. The AIBL study (http://www.AIBL.csiro.au) is a consortium between Austin Health, CSIRO, Edith Cowan University, the Florey Institute (The University of Melbourne), and the National Ageing Research Institute. We thank all the investigators within the AIBL who contributed to the design and implementation of the resource and/or provided data but did not actively participate in the development, analysis, interpretation or writing of this current study. A complete listing of AIBL investigators can be found at ![]() .
.
FUNDING
This work was supported by grants from the Swedish Research Council (2018-01729) and the Kempe Foundation (JCK-1922.1) to SP. Financial support was also provided through a regional agreement between Umeå University and Västerbotten County Council, grants: RV-735451 (2018–2020); RV-453141 (2015–2017); RV-225461 (2012–2014) and year-wise RV-741571, RV-678571, RV-582111, RV-491371, RV- 400741, RV-322831, RV-243741(2012–2018) to RA; as well as year-wise RV- 932787, RV-865381 and RV-745571 to MH. This work was also supported by the Medical Faculty at Umeå University (SD, MH, SP), the Kempe Foundation (SD), and Uppsala-Umeå Comprehensive Cancer Consortium (SD, MH). The Betula project is supported by the Bank of Sweden Tercentenary Foundation [grant number 1988-0082:17; J2001-0682]; the Swedish Council for Planning and Coordination of Research [grant numbers D1988-0092, D1989-0115, D1990-0074, D1991-0258, D1992-0143, D1997- 0756, D1997-1841, D1999- 0739, B1999-474]; the Swedish Council for Research in the Humanities and Social Sciences [grant number F377/1988–2000]; the Swedish Council for Social Research [grant numbers 1988–1990:88-0082, 311/1991–2000]; and the Swedish Research Council [grant numbers 345-2003-3883, 315-2004- 6977]. AIBL DNAm data was supported through funding from the National Health and Medical Research Council (NHMRC) awarded to SML, specifically project grant GNT1161706 a Boosting Dementia Research Grant (GNT1151854) linked to the Joint Programming Neurodegenerative Disease (JPND) BRAIN-MEND grant.
CONFLICT OF INTEREST
The authors have no conflict of interest to report.
DATA AVAILABILITY
The Betula dataset used and/or analyzed in the current study is available from the corresponding author on reasonable request, as long as the data transfer is in agreement with the European Union legislation on the General Data Protection Regulation and Umeå University data protection policies. AIBL DNAm data are available from the GEO repository accession number GSE153712.
