Abstract
Background:
The complex and not yet fully understood etiology of Alzheimer’s disease (AD) shows important proteopathic signs which are unlikely to be linked to a single protein. However, protein subsets from deep proteomic datasets can be useful in stratifying patient risk, identifying stage dependent disease markers, and suggesting possible disease mechanisms.
Objective:
The objective was to identify protein subsets that best classify subjects into control, asymptomatic Alzheimer’s disease (AsymAD), and AD.
Methods:
Data comprised 6 cohorts; 620 subjects; 3,334 proteins. Brain tissue-derived predictive protein subsets for classifying AD, AsymAD, or control were identified and validated with label-free quantification and machine learning.
Results:
A 29-protein subset accurately classified AD (AUC = 0.94). However, an 88-protein subset best predicted AsymAD (AUC = 0.92) or Control (AUC = 0.92) from AD (AUC = 0.98). AD versus Control: APP, DHX15, NRXN1, PBXIP1, RABEP1, STOM, and VGF. AD versus AsymAD: ALDH1A1, BDH2, C4A, FABP7, GABBR2, GNAI3, PBXIP1, and PRKAR1B. AsymAD versus Control: APP, C4A, DMXL1, EXOC2, PITPNB, RABEP1, and VGF. Additional predictors: DNAJA3, PTBP2, SLC30A9, VAT1L, CROCC, PNP, SNCB, ENPP6, HAPLN2, PSMD4, and CMAS.
Conclusion:
Biomarkers were dynamically separable across disease stages. Predictive proteins were significantly enriched to sugar metabolism.
Keywords
INTRODUCTION
Early elucidation of Alzheimer’s disease (AD) is pivotal for constructing clinically impactful treatments. However, the pathophysiology of AD and the driving biochemical changes are not fully understood. Assessment of changes in protein expressions in the brain may assist in elucidation of multi-factorial biochemical changes that lead to AD [1]. Given the complexity and heterogeneity of AD, no single protein is likely to be predictive of all mechanisms or phenotypes which result in AD [2]. Nonetheless, predictive protein models may suggest novel disease mechanisms, improve assessment of patient risk, and signify disease stage-dependentbiomarkers [3].
This work identifies protein subsets that differentiate diagnostic labels for AD. AD diagnosis is often based on clinically measured functional cognitive decline. AD diagnosis is typically determined using a battery of neuropsychological tests in combination with suggestive imaging, genomic, or other clinical features. Common cognitive tests used in AD diagnosis include the Montreal Cognitive Assessment or Consortium to Establish a Registry for Alzheimer’s Disease (CERAD) neuropsychological battery [4]. There is no universal definition of asymptomatic AD (AsymAD). AsymAD is typically characterized by changes in age-adjusted biomarkers, such as increase in amyloid-
In particular, identification of subsets of proteins that better predict and stratify the asymptomatic AD stage is pivotal. Earlier identification of patients likely to transition to AD could enable earlier intervention. The ability to intervene early is likely key to improving outcomes, such as slowing progression or improving symptom-related quality of life. The amyloid-
The study goal was to determine which proteins in the brain (beyond amyloid-
METHODS
Methods consist of data collection and preprocessing; protein selection using a machine learning algorithm to identify the “best” subset of proteins to predict patient diagnostic classification; validation of the algorithm to accurately classify control, AsymAD, or AD patients using only the identified “best” subset of predictive proteins; and assessment of predictive protein functions. All data preprocessing, machine learning, and analysis was performed in Python 3.6.
Patient diagnostic class labels
Note that the patient diagnostic labels (Control, AsymAD, AD) were inherited from previously published work. Briefly, according to the definitions outlined by Johnson et al. [3], the neuropathological diagnostic classes were determined using CERAD criteria to quantify neuritic plaque distribution and Braak staging to quantify extent of neurofibrillary tangle pathology.
Data used for protein biomarker identification
Six public data sets were utilized [3]: Baltimore Longitudinal Study of Aging (BLSA) [12], Banner Sun Health Research Institute (Banner) [13], Mount Sinai School of Medicine Brain Bank (MSSB) [2], Adult Changes in Thought Study (ACT), Mayo Clinic Brain Bank and University of Pennsylvania School of Medicine Brain Bank. Four data sets (
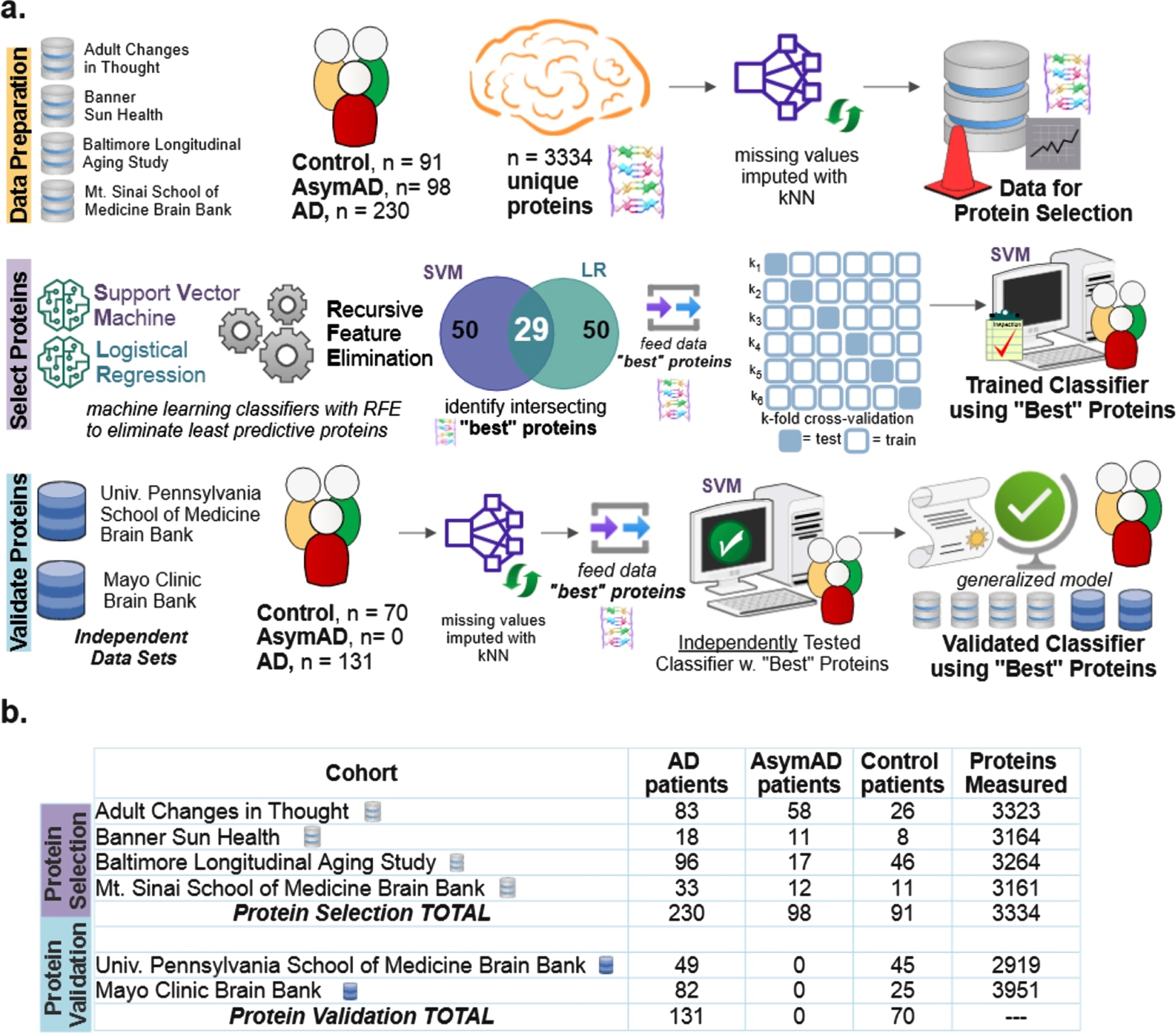
Diagram explaining the data and machine learning pipeline to identify a subset of “best” predictive protein biomarkers to accurately classify Alzheimer’s disease (AD), asymptomatic Alzheimer’s disease (AsymAD), or Control. a) Machine learning pipeline consisted of data preparation, protein selection, and model validation with selected proteins. Data was prepared by aggregating four cohorts (
Protein selection using machine learning
As shown in the protein selection row of Fig. 1a, proteins from the selection cohort (data from BLSA, Banner, ACT, and MSSB) were selected using a combination of classification algorithms with recursive feature elimination (RFE). RFE is a feature selection algorithm which recursively eliminates less important data features until a pre-defined number of features remain in the dataset. In this study, the “features” are the measured proteins. This iterative procedure is an instance of backward selection [14]. RFE [14] is used to determine the most predictive proteins for successful classification. The resultant predictive protein subset was then used to classify each subject as either control, AsymAD, or AD.
RFE is a wrapper-based feature selection algorithm where recursive rounds of elimination are used to determine the subset of proteins that best predict patient diagnostic classification. The final set of selected predictive proteins is, in part, sensitive to the classification method. Thus, two popular linear classification methods were independently used with RFE in the scikit-learn package of Python: support vector machine (SVM) and logistic regression (LR), both with linear kernels [15]. The two classifiers, SVM and LR, separately select a specified number of most predictive proteins equal to the RFE criterion. The RFE criterion is the number of proteins the algorithm is allowed to retain. Note that other classifiers were also tried in place or in combination with SVM and LR. However, the intersection of proteins selected by SVM and LR was most consistent and accurate; hence, all results shown utilized this method.
Proteins are selected based on their superior classification ability as quantitatively measured by the area under the precision-recall curve (AUPRC). The intersecting most predictive proteins become the “best proteins”. The RFE algorithm assessed RFE criterions ranging from 10 to 150 proteins. For example, the Venn diagram of Fig. 1a for protein selection illustrates that an RFE criterion of 50 for SVM and LR resulted in an intersecting set of 29 best proteins. Upon completion of protein selection using RFE, a new SVM classifier is constructed, trained, and validated to classify diagnosis (AD, control, AsymAD) using only the selected best proteins. With three classes (control, AsymAD, AD), a one versus rest approach was utilized (AD versus NonAD, AsymAD versus non-AsymAD, control versus non-control).
Note that alternative methods to RFE to identify the most predictive proteins were considered and tried on LFQ as well as held out data: penalized lasso (Supplementary Figure 5), random forest feature importance (Supplementary Figure 6), and statistical differential protein expression using the f-statistic (Supplementary Figure 7). Also, random forests were coupled with RFE to have a more stringent selection criterion - including a protein only when it is selected by three algorithms: SVM, logistic regression, and random forests (Supplementary Figure 8). Performance comparison to neural network, which played no role in feature selection, is also shown in Supplementary Figures 5–8. In some cases, the alternate methods shown in the supplementary figures performed marginally better on the UPenn dataset, which has only binary labels (Control/AD). In all cases, RFE chosen proteins performed substantially better on the LFQ dataset which has more samples (
Validation of “best” protein subsets to classify diagnosis
As shown in the Protein Validation row of Fig. 1a, the trained SVM classifier was independently tested using validation cohort data (Mayo, UPenn data sets). As part of independent validation, the best set(s) of proteins determined during protein selection with RFE was used to predict validation cohort diagnostic classes. However, there were a couple of exceptions due to required data harmonization. In the Mayo cohort, one of the “best” proteins was not quantified (CROCC|Q5TZA2) and a different protein isoform was quantified for APP; APP|A0A0A0MRG2 was included for Mayo, instead of APP|E9PG40). Similarly, for the UPenn cohort, two of the “best” proteins were not quantified (C4A|P0C0L4, DMXL1|Q9Y485), and a different isoform was quantified for APP (APP|A0A0A0MRG2 instead of APP|E9PG40).
Confusion matrices illustrate true positives (TP), false positives (FP), true negatives (TN), and false negatives (FN) used to calculate classification performance. Additionally, a precision-recall curve (PRC) is generated to assess aggregate final model performance. PRC assesses the model’s classification accuracy when using only the “best” protein subsets to classify diagnosis. PRC is a plot of precision versus recall. Recall is defined as [TP/ (TP+FN)], and precision is defined as [TP/(TP+FP)]. Area under the curve (AUC) provides an aggregate measure of performance across all classification thresholds.
A separate unsupervised learning technique, t-stochastic neighbor embedding (t-SNE), was used to assess separability of AD, AsymAD, and Control subjects using only the selected best proteins subsets determined during supervised learning with RFE.
Finally, principal component analysis (PCA), a dimensional reduction technique, was used to explore and validate RFE criteria. The scree plot and elbow method were used to separately verify how many proteins are necessary to explain the preponderance of variance. The elbow approximated the number of intersecting proteins selected during RFE for optimal diagnostic classification.
Analysis of protein function modules
Selected proteins were matched to their protein function using the color modules and algorithms published by Johnson et al. [3]. There are 14 possible functional modules comprising the entire protein data set (
RESULTS
A machine learning classification and recursive feature elimination process (Fig. 1a) determined which of 3,334 possible clinically measured proteins were most important for classifying control, AsymAD, or AD. RFE was used to identify the proteins that best predicted diagnostic class. Six public data sets were utilized (Fig. 1b). Four data sets (
Classification performance with 29 best proteins
Amyloid precursor protein (APP) is linked to the well-known amyloid-
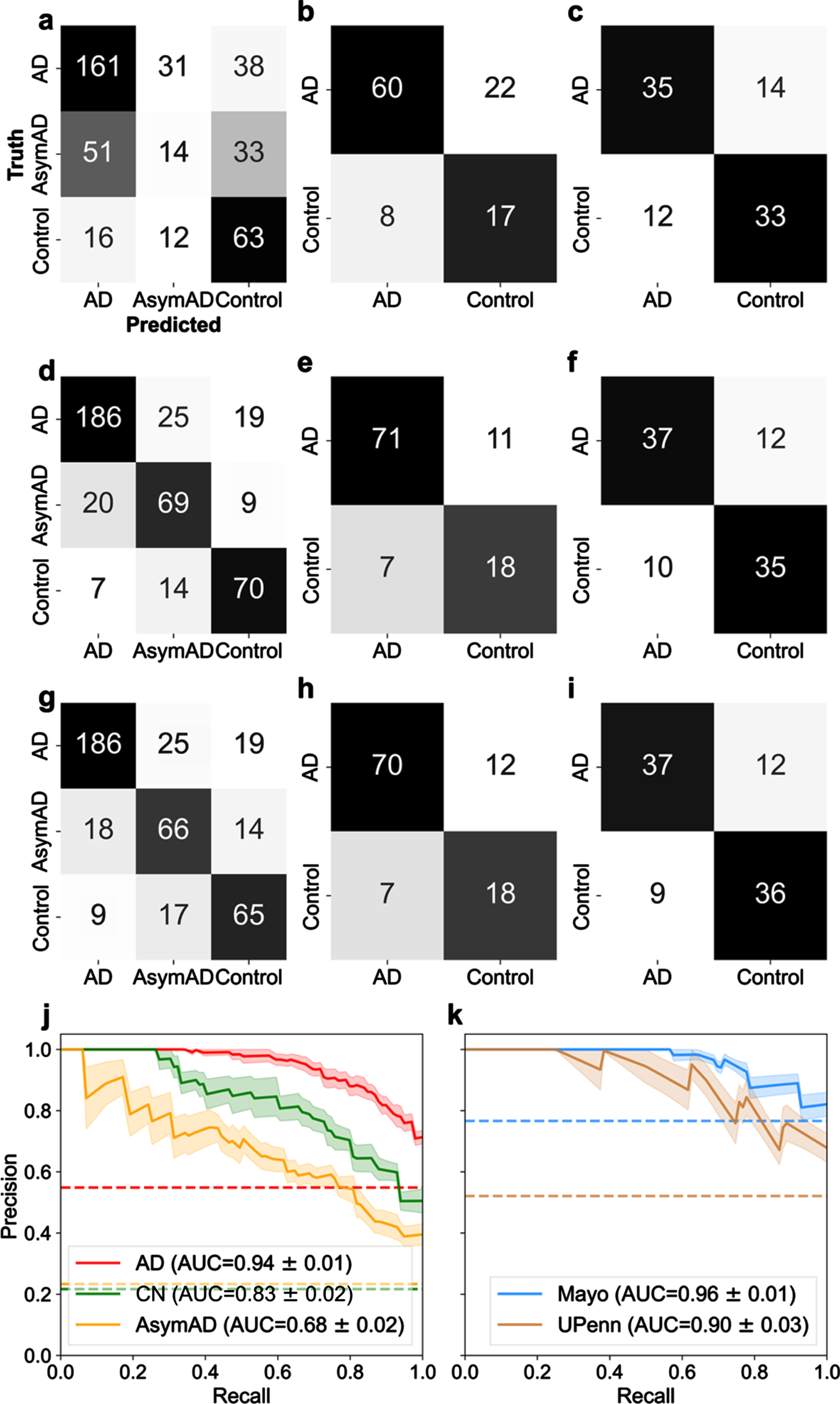
Examination of classification performance for Alzheimer’s disease (AD), asymptomatic Alzheimer’s Disease (AsymAD), and control using
A precision-recall (PR) curve curve is used to assess aggregate classifier performance. AUC is used to quantify the aggregate classification performance using the selected best protein subset(s). AUC = 1 is a perfect classifier; thus, an AUC closer to 1 is desirable. Figure 2j illustrates the PR curve and corresponding AUC for the protein selection cohort for AD, control, and AsymAD, respectively, using the selected 29 best proteins. The shaded area represents the standard error, ±
The best 29 proteins are listed in Fig. 3a with their corresponding model coefficient weight as determined from the SVM classification model. Since the one-versus-rest approach for multi-class classification is used, it results in three coefficients for each protein. The three coefficients for every protein correspond to the three diagnostic classes (AD, AsymAD, and Control). The corresponding heatmap illustrates how the selected proteins (
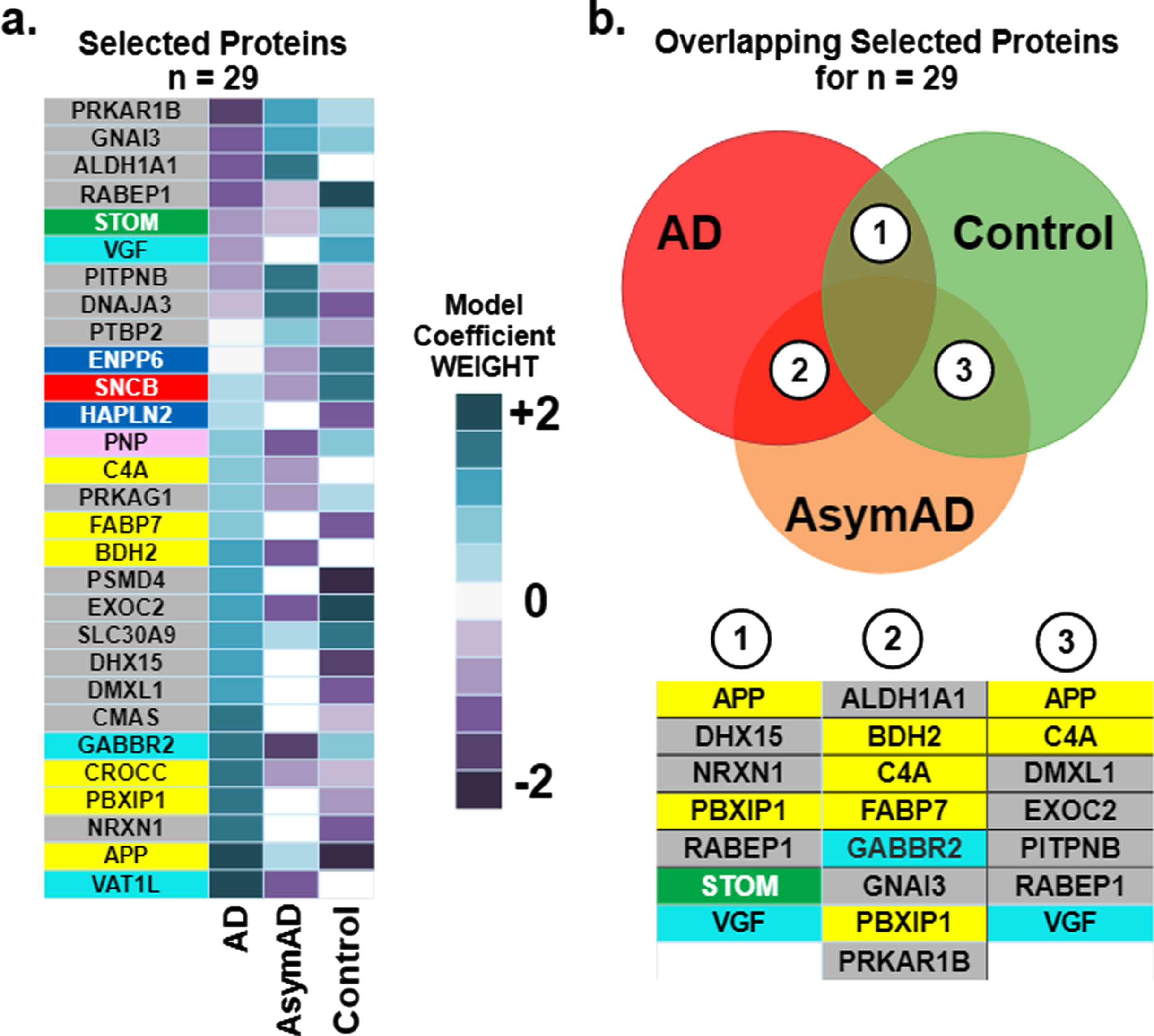
Driving effects of the selected 29 proteins on the prediction of diagnostic classes of Alzheimer’s (AD), asymptomatic Alzheimer’s (AsymAD), and Control. Note the color of the box around individual listed proteins corresponds to functional modules derived in Johnson et al. [3] and detailed later in Fig. 6. a) The heatmap illustrates the relative magnitude that each protein drives diagnostic class. Driving effects are determined from the coefficients of the SVM. Since the one-versus-rest approach for multi-class classification is used, results encompass three coefficients for each selected protein, one coefficient corresponding to each diagnostic class (AD, AsymAD, Control). b) Key proteins that drive correct prediction in multiple or overlapping classes. The Venn diagram illustrates the overlap in classes: area 1 denotes overlap between AD and Control, area 2 overlap between AD and AsymAD and area 3 between AsymAD and Control. The biomarker panels for each overlap area denote individual predictive proteins shared by overlapping classes.
Figure 3b examines the overlap of the 29 selected proteins in driving diagnostic class (AD, AsymAD, Control). AD and Control, labeled as area 1 on the Venn diagram, share 7 driving proteins: APP, DHX15, NRXN1, PBXIP1, RABEP1, STOM, and VGF. AD and AsymAD, labeled as area 2 on the Venn diagram, share eight driving proteins: ALDH1A1, BDH2, C4A, FABP7, GABBR2, GNAI3, PBXIP1, PKAR1B. AsymAD and Control, labeled as area 3 on the Venn diagram, share seven driving proteins: APP, C4A, DMXL1, EXOC2, PITPNB, RABEP1, and VGF. Note that the color coding of each protein, itself, in Fig. 3 corresponds to function as described in the
Classification performance with 88 proteins
An RFE of 50, resulting in 29 selected proteins, was sufficient for differentiating AD and Control classes. However, an RFE criterion of 150, resulting in 88 selected proteins, was optimal for differentiating AD and AsymAD classes. Figure 4a and 4b illustrate the PR curve and AUC for each class utilizing the 88 protein subset for predicting diagnostic classification. Utilizing the 88-protein subset increased AD and Control classification performance by approximately 4% and 9% respectively compared to the 29-protein subset (Fig. 4a). Utilizing the 88-protein subset increased AsymAD classification performance by approximately 24% (Fig. 4a). In the independent validation cohorts (Fig. 4b), which did not contain any AsymAD patients, the 29-protein subset marginally outperformed the 88-protein subset. In short, AsymAD requires substantially more proteins for accurate predictive classification.
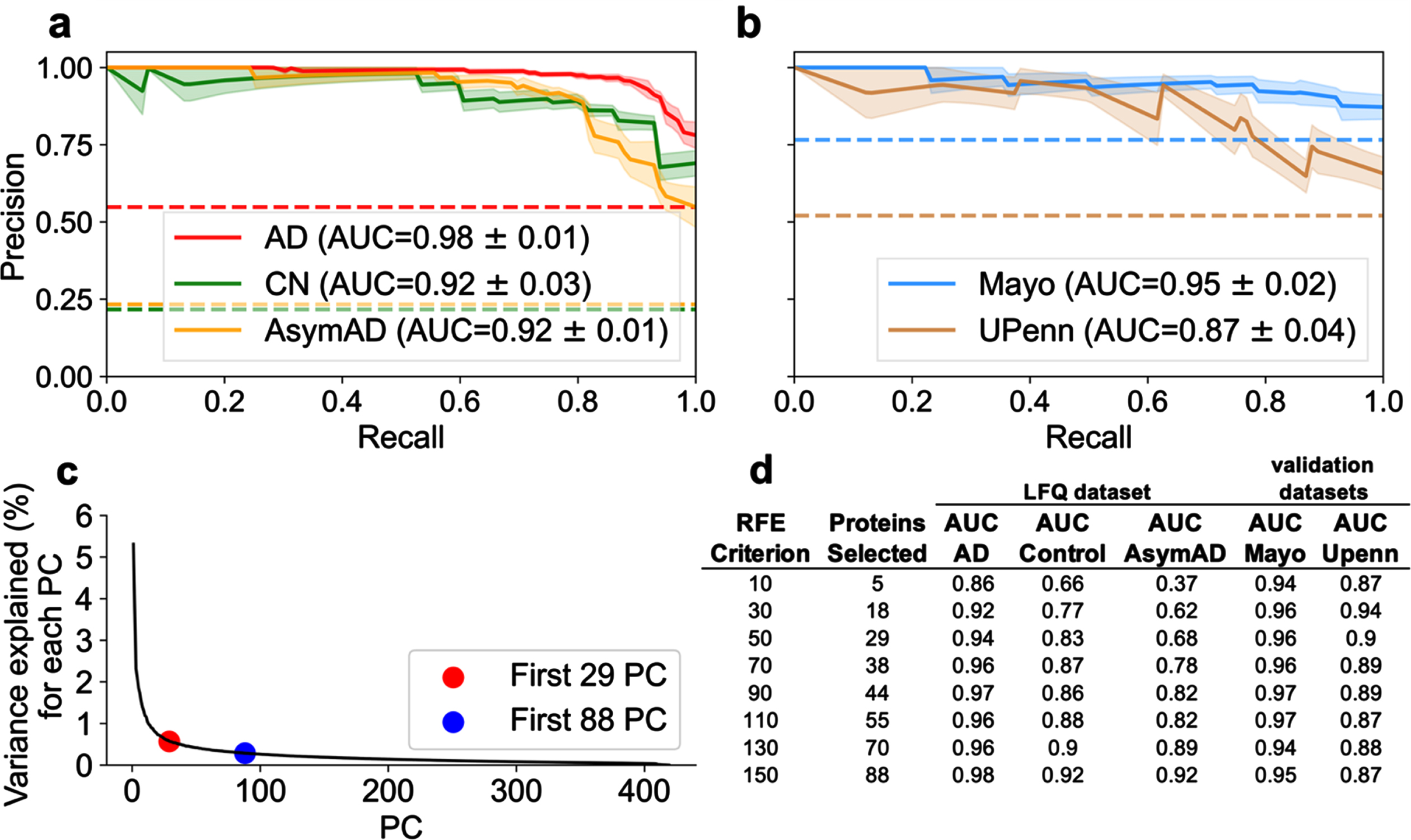
Examination of optimal number of proteins necessary for superior classification between AD, AsymAD, and Control. A best predictive protein set of 88 (
Further exploration of classification performance as a function of RFE criterion
The RFE criterion for protein selection was varied to determine the optimal protein subsets. Again, the RFE criterion determines the number of proteins each classifier (SVM and LR) can select. For a given RFE criterion, the intersection of proteins selected by both SVM and LR become the resultant number of “best” predictive proteins. As described above, RFE = 50, resulting in 29 proteins, was sufficient to classify AD versus control. The number of intersecting best predictive proteins for diagnostic classification is not random. Rather, these thresholds are explained by examining dimensional reduction with PCA. Figure 4c illustrates variance explained as a function of number of components. The scree plot approximates minimum components needed to explain the preponderance of variance. The “elbow” of the scree plot denotes the optimal range of components needed to account for the preponderance of variance. Figure 4c shows 29 components (red dot) corresponds to the start of the elbow and 88 components (black dot) corresponds to the end of the elbow. Variance per component beyond the elbow asymptotically approaches zero. Hence, those additional components should not substantively improve model performance. Figure 4d examines the impact of RFE criterion and the resultant number of selected best predictive proteins on diagnostic classification performance. Selecting a RFE criterion greater than 150 (not shown) did not result in increased classification performance.
Functional themes in the selected proteins
The biomarker proteins were mapped to their corresponding modules, which were identified by using the weighted gene correlation network analysis (WGCNA) algorithm. Colors and module number are used to define the function of the biomarkers [3] and are recapitulated in the first 3 columns of Fig. 5. Functional module frequency (expressed as a percentage) of selected best proteins (for
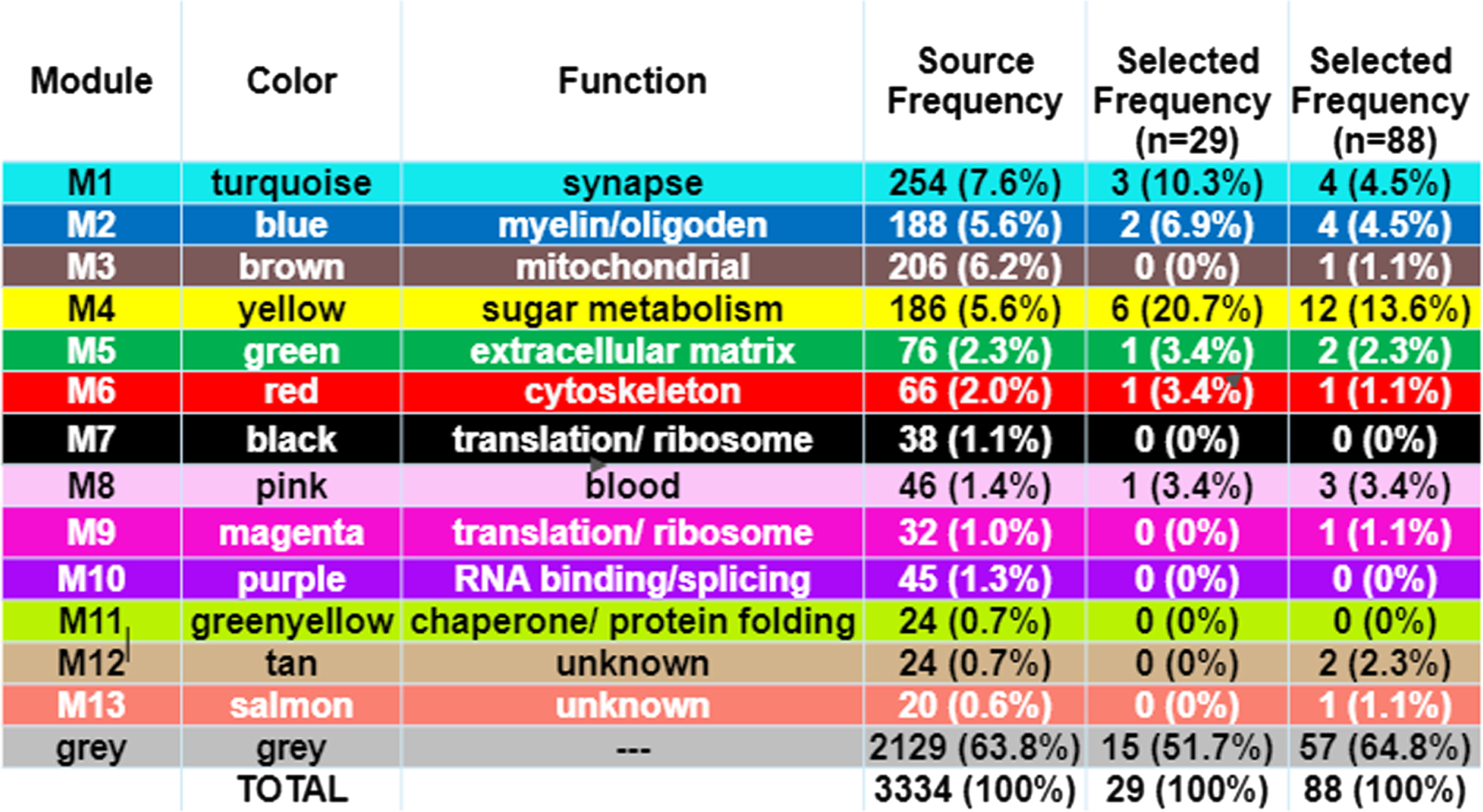
Functional Protein Module of Selected “Best” Predictive Proteins. The functional protein modules are as defined by Johnson et al. [3]. Source frequency is the frequency of the module in the source protein set (
DISCUSSION
Of the 3,334 proteins, machine learning determined a minimum 29-protein subset necessary to accurately classify AD and Control, but an 88-protein subset was necessary to accurately classify AsymAD. The additional proteins needed for AsymAD classification is likely due to greater complexity and heterogeneity of the AsymAD disease state. The “best” predictive protein subsets (
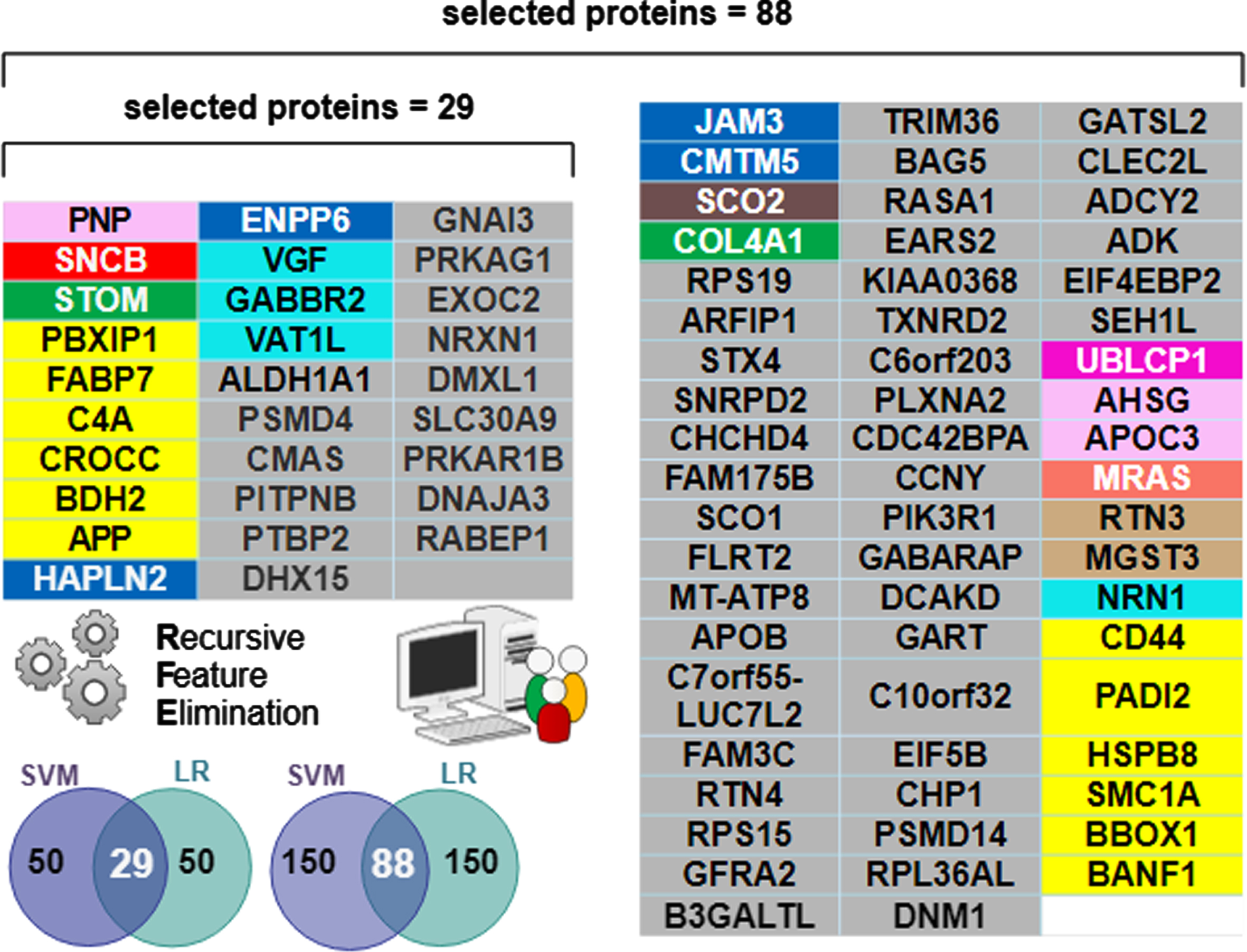

Individual proteins comprising the selected “best” predictive protein subsets color-coded by functional module. The
Homeostatic regulatory dynamics key to disease progression
There was relatively little overlap between the predictive proteins that drive control-AsymAD changes and predictive proteins that drive AsymAD-AD changes (see Fig. 3b). This finding indicates an associative relationship to multifactorial dynamic disease progression etiology. In short, the most predictive proteins dynamically change with disease stage (see Fig. 3a). Whether familial or sporadic AD, different underlying proteomic perturbations may result in multi-scalar system destabilization (e.g., failed homeostasis) with corresponding functional disease phenotypes. Homeostasis is critical for maintaining health, and thus, instabilities often appear in disease [17]. Multifactorial homeostatic instability has been suggested as an underlying propagating mechanism in other neurological pathology, including amyotrophic lateral sclerosis [18], absence epileptic seizures [19], Parkinson’s disease [20], and secondary spinal cord injury [21].
Overlapping proteins for class discrimination
Supplementary Table 1 presents literature details on functions and cited associations with each member of the 29-protein subset. Supplementary Table 1 includes the protein unique ID, brief description of its function and role in AD (if known), and a corresponding reference. Five proteins of the 29-protein subset overlapped in class discrimination (Fig. 3b): APP, VGF, RABEP1, C4A, PBXIP1. APP (upregulated in AD, AsymAD) was expected given its role in the amyloid cascade [16]. VGF (downregulated in AD) protects against amyloid-
Sugar metabolism biomarkers enriched in 29-protein and 88-protein set
Sugar metabolism proteins in both the 29-protein and 88-protein sets (Fig. 6) included: APP, BDH2, C4A, CROCC, FABP7, PBXIP1. Sugar metabolism proteins in the expanded 88-protein subset included CD44, an immune marker associated with AD [27]; PADI2, an age and AD-related marker [28]; BANF1, implicated in aging and progeria [29]; HSPB8, inhibitor of amyloid-
The significantly enriched sugar metabolism module (Fig. 5) supports the recent perspective that asymptomatic and symptomatic AD is characterized by dysregulation of energy metabolism [33, 34]. In short, the presented work supports the hypothesis that sugar metabolism becomes more impacted with disease progression. Insulin resistance in the brain modulates AD inflammatory markers and decreases amyloid clearance [35]. The exact link between AD and type 1 or 2 diabetes is under debate. Nonetheless, poorly controlled blood sugar appears to increase risk of AD [35]. Some researchers have referred to the dysregulation of blood sugar in the brain in AD “type 3 diabetes” [36]. Interconnections between inflammation, metabolism, and protein clearance are further evidence of a multifactorial homeostatic instability contributing to AD progression [3, 37].
The good and the bad of APP
APP is another example where homeostatic instability may play a role in disease progression. For quite some time, APP has been known to be involved in the formation of amyloid-
Blood-based biomarkers
Biomarkers detected in the blood are preferable for AD risk assessment and early diagnosis [41]. Only one blood-based module protein was selected in the 29-protein subset: PNP, a purine-related metabolite altered early in AD [42]. Two additional blood-based proteins were in the 88-protein subset: AHSG and APOC3. A higher apoE level in high density lipoprotein that lacks apoC3 was associated with better cognitive function [43]. AHSG, a highly glycosylated protein appears downregulated in AD [44].
Assessment of alternatives and limitations
The presented RFE method in Fig. 1, and its corresponding presented results above, were thoroughly vetted and compared to several other statistics-based and machine learning-based model alternatives. The presented method consistently outperformed all other alternative methods and models (see Supplementary Figures 5–8), especially in the mega-LFQ data set with three classes (Control, AsymAD, and AD). In summary, the presented 29-protein and 88-protein lists for diagnostic classification were quite stable. Relaxing the RFE criterion to include more proteins beyond the selected 88-proteins did not improve classification results (Fig. 4). Nonetheless, no model or method is perfect. While the model is stable, it is fair to expect that a small number of proteins included on the final presented list(s) could be substituted for non-included proteins (e.g., such as similarly co-expressed proteins or proteins from the same functional module). As such, regardless of method, a few proteins that relayed similar, correlated, or mutual information as the selected proteins may not have made the presented final selected proteins list(s). In full transparency, the performance of the protein list generated by each alternative method is shown in Supplementary Figures 5–8. Notably, many of the RFE selected final proteins presented in the main article were recurringly selected by the alternative methods. Finally, any proteins not included in the presented final lists (or even in the input study data) could have their relative importance deduced based on co-expression or their functional modules.
Future directions
The LFQ data covered 3,334 proteins from which the identified “best” biomarker subset is derived. However, the presented method could be extended to future more comprehensive data sets, such as tandem mass tag, to further optimize results. Future addition of larger validation cohorts, especially AsymAD, will ensure model generalizability. Additionally, future inclusion of traits such as gender and race (when available) are important to determine if there are specific feature biases that impact the predictive ability or discriminative expression of proteins. Finally, this work utilized the common 3-class AD staging system: control, AsymAD, or AD. However, it is possible there is a more optimal temporal disease staging system. For example, integrative data machine learning analysis suggested with Alzheimer’s Disease Neuroimaging Initiative data suggested at least four clusters of symptomatic AD patients [45].
Conclusions
Machine learning successfully identified proteins subsets most predictive for classifying AD, AsymAD, and Control subjects. The most predictive proteins subsets comprised < 3% of the 3,334 proteins assessed. A 29-protein subset accurately classified AD versus Control, but an 88-protein subset was needed to accurately classify AsymAD. The protein subsets resulted in a robust classifier model. The presented model generalized to accurately predict diagnostic labels on unseen data in independent validation cohorts regardless of brain region or minor data set differences. The predictive protein subsets included known important proteins like APP. However, diagnostic classification performance did not hinge upon APP or any single protein or pathway. Finally, the most predictive subsets were significantly enriched in proteins linked to sugarmetabolism.
Footnotes
ACKNOWLEDGMENTS
The authors have no acknowledgments to report.
FUNDING
This research was funded by National Science Foundation CAREER grant 1944247 to C.M, Alzheimer’s Association Research Grant Award 2018-AARGD-591014 to C.M., Goizueta Alzheimer’s Disease Research Center at Emory University grant awards P50 AG025688 and P30 AG066511 to C.M., National Institute of Health grant U19-AG056169 sub-awards to C.M, J.L., A.L.
CONFLICT OF INTEREST
C.M. is an Editorial Board Member of this journal but was not involved in the peer-review process nor had access to any information regarding its peer-review.
DATA AVAILABILITY
All data used in the analysis has been previously published in [3] and is publicly available at ![]() .
.
