Abstract
Background:
Recent studies had explored that gut microbiota was associated with neurodegenerative diseases (including Alzheimer’s disease (AD), Parkinson’s disease (PD), and amyotrophic lateral sclerosis (ALS)) through the gut-brain axis, among which metabolic pathways played an important role. However, the underlying causality remained unclear.
Objective:
Our study aimed to evaluate potential causal relationships between gut microbiota, metabolites, and neurodegenerative diseases through Mendelian randomization (MR) approach.
Methods:
We selected genetic variants associated with gut microbiota traits (
Results:
Greater abundance of
Conclusion:
Our study firstly applied a two-sample MR approach to detect causal relationships among gut microbiota, gut metabolites, and neurodegenerative diseases. Our findings may provide new targets for treatments and may offer valuable insights for further studies on the underlying mechanisms.
Keywords
INTRODUCTION
Neurodegenerative diseases are characterized by progressive loss of structure or function of neurons in the central or peripheral nervous system, which involves irreversible long-term motor or cognitive impairments [1]. The prevalence of neurodegenerative diseases, including Alzheimer’s disease (AD), Parkinson’s disease (PD), and amyotrophic lateral sclerosis (ALS), are rising worldwide with the increasing life expectancy. In recent years, emerging evidence has indicated that gut microbiota derived metabolites including short-chain fatty acids (SCFAs) [2, 3] and neurotransmitters such as glutamate [4], serotonin [5, 6], and γ-aminobutyric acid (GABA) [7] may play a central role in the gut-brain axis alterations and risk of neurodegenerative diseases [8]. However, few consistent links connecting gut microbiota and diseases or their associated metabolic pathways were found.
Increasing number of cross-sectional studies have implicated the association between gut microbiota and neurodegenerative diseases, including AD, PD, and ALS [9]; however, such associations differed across studies. For example, an observational study (
Mendelian randomization (MR) approach, which uses genetic variants as instrumental variables (IVs), has been widely accepted to determine the causal effect of exposures on diseases [16]. As single nucleotide polymorphisms (SNPs) are of random allocation and is independent of confounders, MR is similar to randomized controlled trial and circumvent the limitations of previous observational studies.
Therefore, our study firstly applied a two-sample MR approach to detect causal relationships among gut microbiota, metabolites, and neurodegenerative disorders including AD, PD, and ALS, using summary statistics from the largest genome-wide association studies (GWAS) so far.
MATERIALS AND METHODS
Data sources and instruments
Summary statistics applied for investigating traits had the largest sample sizes with similar populations, and exposure and outcome statistics were obtained from different consortia, sample overlap tended to be little [17]. Details of the contributing GWAS consortiums are listed in Supplementary Table 1.
Gut microbiota
We leveraged summary statistics from most comprehensive exploration of genetic influences on human gut microbiota so far. The MiBioGen consortium recruited 18,340 participants of multiple ancestries (including European, American Hispanic/Latin, East Asian, etc.) from 24 cohorts [18]. After extracting DNA from fecal samples, 16S rRNA gene sequencing was utilized to characterize the gut microbiome using SILVA as a reference database [19], with truncation of the taxonomic resolution to genus level.
Gut metabolites
Considering the important roles of gut metabolites in microbiota-host crosstalk, we also leveraged summary-level data from a GWAS of the human metabolome conducted among European-descent subjects (TwinsUK and KORA,
Neurodegenerative diseases
We utilized the GWAS summary statistics from the largest and most recent datasets for AD, PD, and ALS so far. We obtained the corresponding genetic variants from the International Genomics of Alzheimer’s Project (IGAP) including 17,008 cases and 37,154 controls [21], the International Parkinson’s Disease Genomics Consortium (IPDGC) including 37,688 cases and 141,779 million controls [22], and the International Amyotrophic Lateral Sclerosis Genomics Consortium (IALSC) including 20,806 cases with ALS and 59,804 controls [23]. Cases of those neurodegenerative diseases were all clinically confirmed using published criteria.
Ethical approval for each study had been obtained in all original articles [18, 21–24], and no ethical approval for the current analyses was needed as they were based on publicly available summary statistics.
Selection of instrumental variables
To ensure the validity of the conclusion, our study applied several quality control steps to select instrument variables. First, SNPs significantly related to gut microbiome were selected as instrumental variables. Two thresholds were used to select the instrumental variable. In order to obtain more comprehensive results and increase sensitivity to IVs, SNPs smaller than the locus-wide significance level (1×10–5) was selected as instrumental variables in primary analysis. We also selected exposure-related SNPs at genome-wide significance (
Statistical analyses
We applied two sample MR as our main statistical methods to estimate causal associations between each instrument-exposure (gut microbiota and metabolite) and instrument-outcome (AD, PD, and ALS). The MR approach was based on three key assumptions: 1) the genetic variant must be truly associated with the exposure; 2) the genetic variant should not be associated with confounders of the exposure-outcome relationship; and 3) the genetic variant should only be related to the outcome of interest through the exposure under study [25].
Primary analyses were performed using Inverse-variance weighted (IVW) method, which essentially assumed the intercept was zero, and our results were corrected for multiple hypothesis testing using the Benjamini and Hochberg false discovery rate (FDR), as significance threshold was set at FDR-corrected
To validate assumption 3 and improve the robustness of the findings, we also undertook a series of sensitivity analyses including MR-Egger regression, weighted mode, weighted median, simple median methods, and robust adjusted profile score (MR.RAPS) method, which provided different assumptions about horizontal pleiotropy [28, 29]. However, the MR-Egger method had the lowest power among the 6 methods and was based on the instrument strength independent of the direct effects (INSIDE) assumption, with no measurement error in the SNP exposure effects (NOME) assumption [30]. Therefore, MR Egger was performed when I2GX was > 0.9 [31].
Cochran Q statistic and leave-one-out sensitivity analysis were also adopted to the SNPs that may influence the outcome through an unaccounted causal pathway, and Steiger analysis was performed to explore direction of causal effects [32]. Furthermore, MR-Egger intercept and Mendelian Randomization Pleiotropy RESidual Sum and Outlier (MR-PRESSO) global test were used to detect the presence of pleiotropy [33].
At last, we conducted multivariable MR (MVMR) analyses using IVW method to estimate the direct and indirect effect of each exposure on an outcome, as we found a high degree of IV overlap across gut microbiota (Lentisphaerae at phylum level, Lentisphaeria at class level, and Victivallales at order level) in univariable MR analyses on PD [34]. Furthermore, we also conducted multivariable MR-Egger analyses to evaluate the horizontal pleiotropy for direct and indirect effects. The IVs used for MVMR analysis were listed in Supplementary Table 8.
A flowchart of our study was provided in Fig. 1. The MR analyses were performed in the R version 4.0.2 computing environment using the latest TwoSampleMR (https://github.com/MRCIEU/TwoSampleMR), MVMR (https://github.com/WSpiller/MVMR), and MRPRESSO (https://github.com/rondolab/MR-PRESSO) packages.
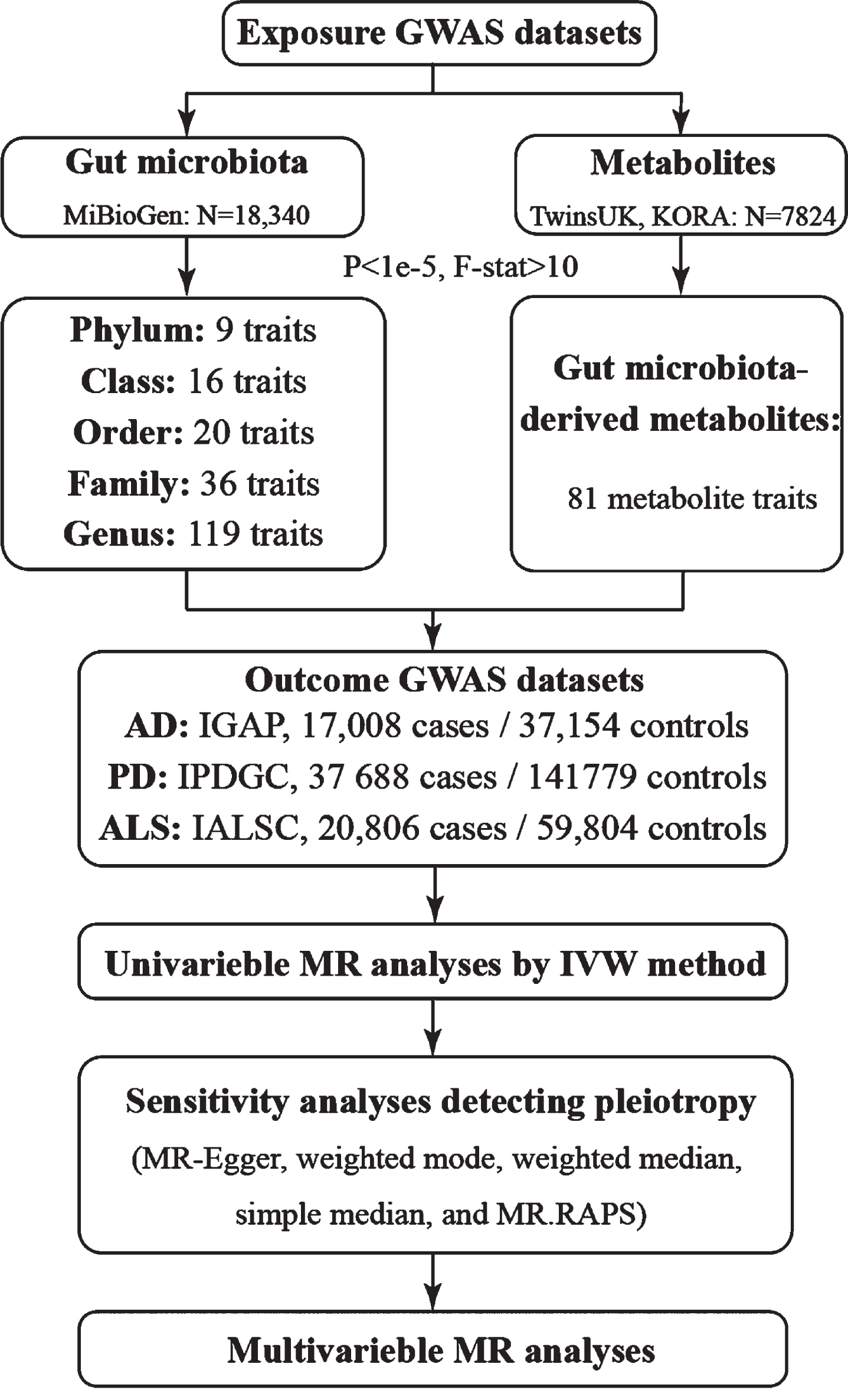
Flowchart of current study. AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis.
RESULTS
Associations between gut microbiota and neurodegenerative diseases
By the means of IVW method, results reaching a threshold of
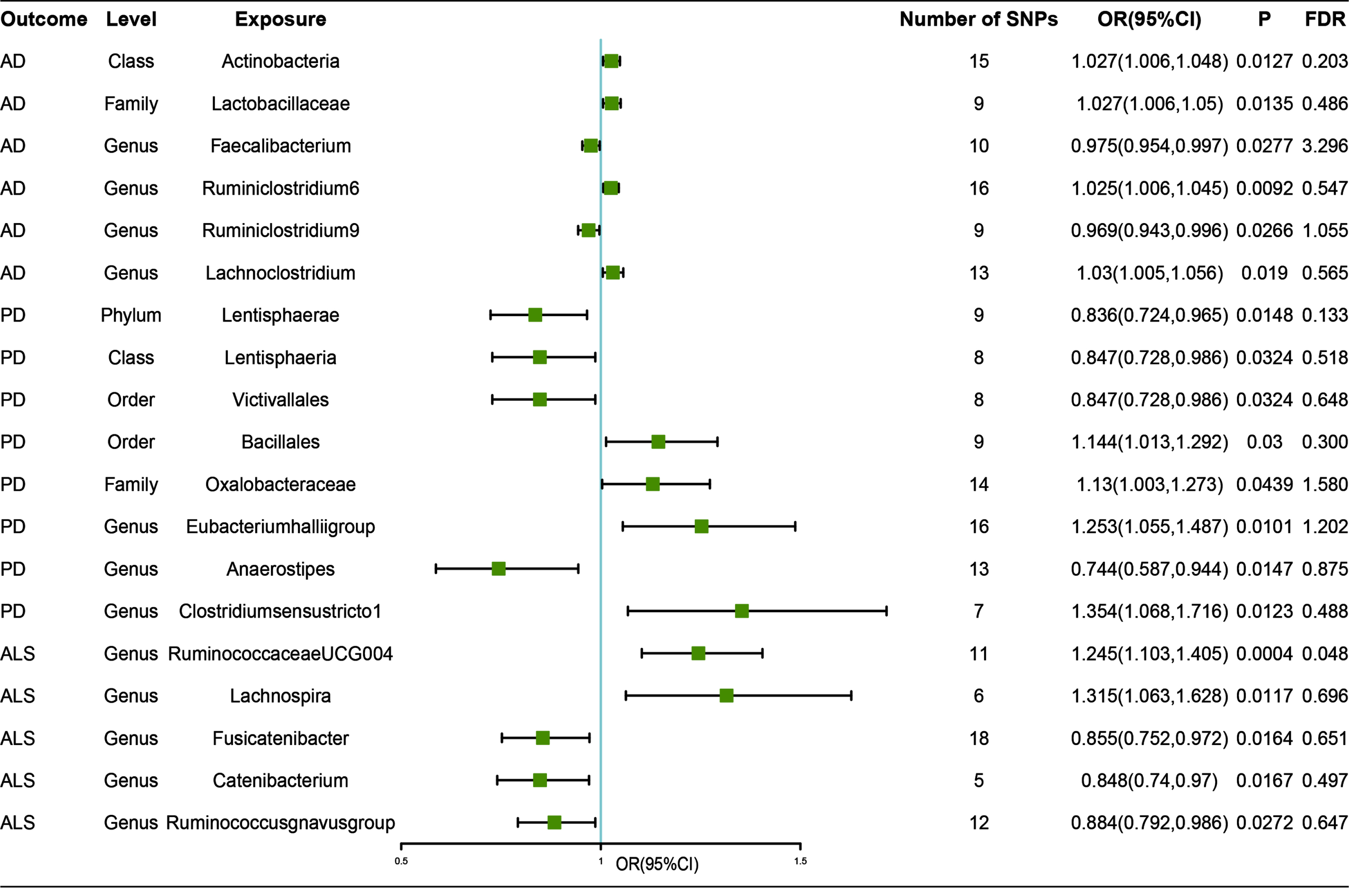
Associations of genetically predicted gut microbiota with risk of neurodegenerative diseases using IVW method. OR, odds ratio; CI, confidence interval; FDR, False discovery rate.
Sensitivity analyses of MR analyses of neurodegenerative diseases on gut microbiota by MR Egger, simple mode, weighted median, and weighted mode
OR, Odds ratios for associations of genetically predicted gut microbiota traits with neurodegenerative diseases; CI, confidence interval; MR, Mendelian randomization; RAPS, robust adjusted profile score; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis.
Causal relationship between gut microbiota and other neurodegenerative diseases were also analyzed by the same process. Our study revealed that genetically increased abundance of Lentisphaerae at phylum level (OR, 0.836; 95% CI, 0.724–0.965;
Besides, genetically increased
Those estimate effects mentioned above were considered robust (Table 1) with no directional pleiotropy or heterogeneity was significant (see Supplementary Table 7), and MR power calculation results were showed in Supplementary Table 6.
Associations between gut metabolites and neurodegenerative diseases
Among 81 gut microbiota-derived metabolites incorporated in our MR analyses, we found 11 suggestive estimate effects of gut metabolite on neurodegenerative diseases. Those metabolites were classified into two types: host-derived or dietary molecules [35].
With regard to host metabolites transformation, our study suggested that increased abundance of taurodeoxycholate, which was a product of primary bile acids (OR, 1.16 for risk ratio of ALS per SD unit of taurodeoxycholate; 95% CI, 1–1.345;
For the dietary molecules, amino acids, complex plant polysaccharides and polyphenols were considered to exert impact on brain function. In tryptophan metabolism, our study revealed that serotonin (OR, 0.535; 95% CI, 0.292–0.979;
Furthermore, those results were judged to be reliable without pleiotropy through sensitivity analyses (Table 2, Supplementary Table 7). However, no significant association was revealed (FDR-corrected
Sensitivity analyses of MR analyses of neurodegenerative diseases on gut microbiota dependent metabolites features by MR Egger, simple mode, weighted median, and weighted mode
OR, Odds ratios for associations of genetically predicted gut microbiota-derived metabolite traits with neurodegenerative diseases; CI, confidence interval; MR, Mendelian randomization; RAPS, robust adjusted profile score; AD, Alzheimer’s disease; PD, Parkinson’s disease; ALS, amyotrophic lateral sclerosis.
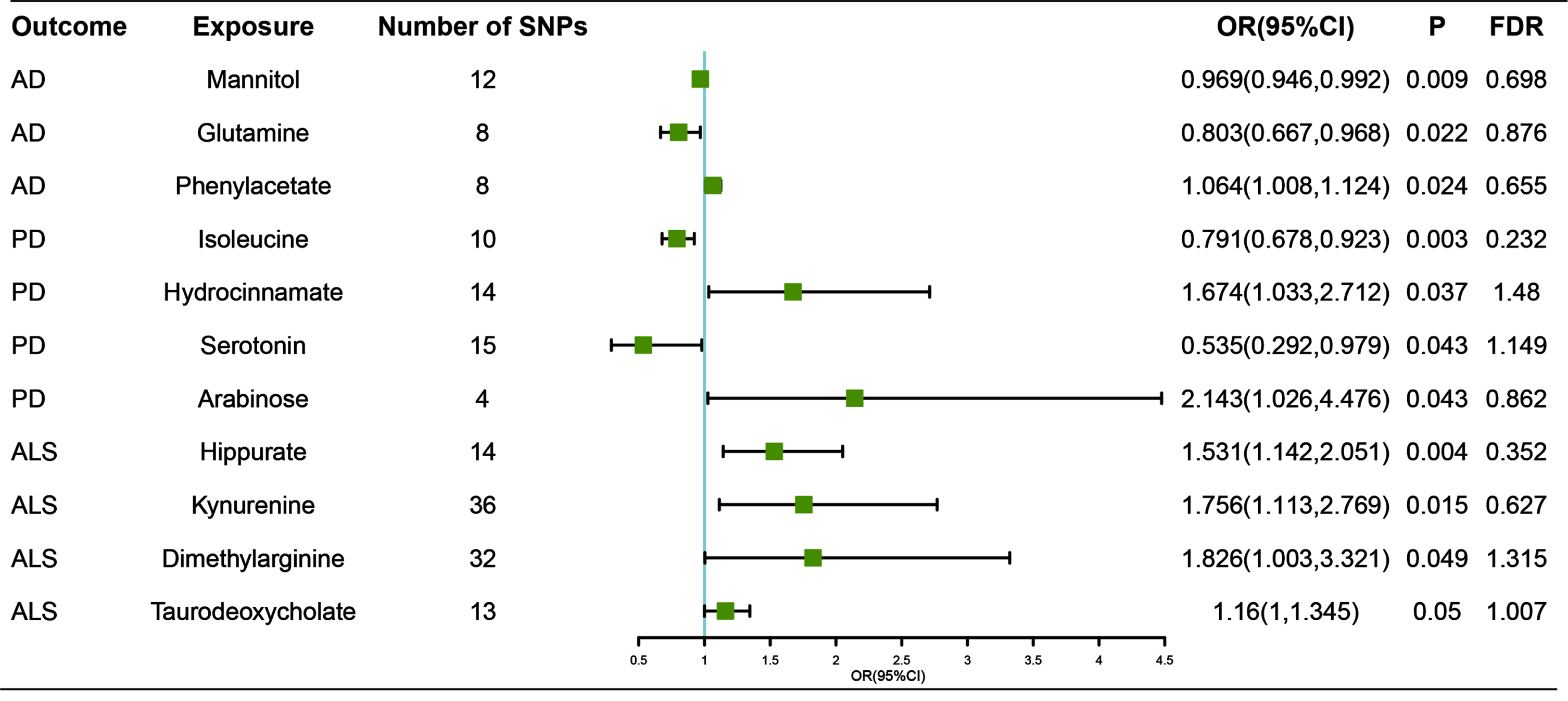
Associations of genetically predicted gut microbiota-dependent metabolites with risk of neurodegenerative diseases using IVW method. OR, odds ratio; CI, confidence interval; FDR, False discovery rate.
The results of MR analysis which applied a SNP selecting threshold of genome-wide significant level of
DISCUSSION
In the present MR study, we found significant association of increased abundance of genera
A previous MR study has suggested that increase in
Our study revealed suggestive causal effect of increased abundance of phylum Lentisphaerae, class Lentisphaeria, and order Victivallales on protective effects of PD; however, no direct effect revealed after multivariable MR analysis, while no relevant result was reported in previous studies either, therefore, such results should be treated with caution. Other associations of Family Oxalobacteraceae, Order Bacillales,
Tryptophan is broken down by the microbiota into indole derivatives and also tryptamine and kynurenine metabolites, and those metabolites were considered important in gut-brain axis [40, 41]. Previous studies have revealed that glutamate signals are destroyed by serotonergic overdrive, and serotonergic dysfunction is associated with the development of motor and non-motor symptoms and complications in PD [42]. Moreover, kynurenine pathway (KP) of tryptophan degradation is involved with several neuropathological features present in ALS including neuroinflammation, excitotoxicity, oxidative stress, immune system activation, and dysregulation of energy metabolism [43]. Previous clinical studies have revealed that serum kynurenine in controls were lower than that in ALS [44]. Our study proved that serotonin was protective factor of PD, while kynurenine was risk factor of ALS, and those molecules may become potential biomarkers to assess the progression of relative diseases. In addition, other amino acid such as glutamine and isoleucine were found causally associated with lower risk of AD and PD. Actually, up to 50% of all α-amino groups of glutamate and glutamine are derived from leucine. Leucine is a regulator of the mechanistic target of rapamycin (mTOR) complex 1 (mTORC1), which is critical on protein synthesis and degradation, autophagy as well as maintenance of glutamate homeostasis, and may have effects on the neuronal solute transport and the excitatory neurotransmitter function [45]. Moreover, in the glutamate-glutamine cycle, synaptically-released glutamate is rapidly transported into astrocytes, and glutamine is then released by astrocytes through SN-type glutamine transporters into the extracellular fluid. Aβ has been shown to reduce the surface expression of GLT-1and to impair astrocyte glutamate uptake [46, 47]. A recent study demonstrated that altered astrocyte glutamine synthesis directly impaired neuronal GABA synthesis in brain slices of the 5xFAD mouse model of AD [48], and our results provided clinical evidence to confirm that reduction of glutamine in peripheral blood was causally associated with occurrence of AD.
Bacterial metabolites produced from polyphenol precursors were also found at levels sufficient to exert biological effects enter circulation [49].
Among the strengths of the study are the most comprehensive MR study on association of gut microbiota and metabolite traits with neurodegenerative diseases, and the largest sample size so far. However, our study still suffers from several limitations. First, most of the results did not survive a strict FDR correction. However, MR was a hypothesis-driven approach, and it could be used to detect some causal relationships regardless of FDR adjusting when some biological evidence exists. Second, 16S rRNA gene sequencing describes gut microbiota from genus to phylum level only, and metagenomic and multiomic approaches may offer opportunities to target gut microbiota composition at a more specific level, avoiding bias if species of more specific level associated with neurodegenerative diseases. Third, our findings might have been affected by weak instrument bias as we included a loose cutoff of exposure-related SNPs at a threshold of
Footnotes
ACKNOWLEDGMENTS
All data used in the current study were based on summary-level data that have been made publicly available. Summary data from genome-wide association studies for the gut microbiota (Kurilshikov et al.) is available at https://www.mibiogen.org/. Metabolomic GWAS summary statistics (Shin et al.) are available for download from the metabolomics GWAS server (http://metabolomics.helmholtz-muenchen.de/gwas/index.php?task=download). Summary level data for AD (Jansen et al.) can be obtained from https://ctg.cncr.nl/, for PD (Nalls et al.) from https://pdgenetics.org/resources (participants from 23andMe Inc. were excluded), and for ALS (Nicolas et al.) from http://als.umassmed.edu. All data generated or analyzed during this study are included in this published article and its ![]() .
.
This study was supported by grants from the National Natural Science Foundation of China (82071201, 82071201), Shanghai Municipal Science and Technology Major Project (No.2018SHZDZX01) and ZHANGJIANG LAB, Tianqiao and Chrissy Chen Institute, and the State Key Laboratory of Neurobiology and Frontiers Center for Brain Science of Ministry of Education, Fudan University.
