Abstract
Background:
The previous studies have identified several genes in relation to Alzheimer’s disease (AD), such as
Objective:
To illustrate the association between the common variants within or adjacent to the coding regions of AD susceptible genes and CSF biomarkers in AD patients.
Methods:
75 sporadic probable AD patients were extracted from the dementia cohort of Peking Union Medical College Hospital. They all had history inquiry, physical examination, blood test, cognitive assessment, brain MRI, CSF testing of Aβ42, 181p-tau, and t-tau, and next-generation DNA sequencing. Sixty-nine common single nucleotide polymorphisms (SNPs) (minor allele frequency > 0.01) within or near the coding region of 13 AD susceptible genes were included in the analysis.
Results:
The rs7412-CC (
Conclusion:
Some common variants within or near the coding regions of
Keywords
INTRODUCTION
Alzheimer’s disease (AD) is the most prevalent neurodegenerative dementia. In 2010, there were about 3.71 million people in China who lived with AD [1]. However, the genetic basis of AD remains unclear. Pathogenic mutations in the amyloid precursor protein (
AD is pathologically characterized by extracellular deposition of amyloid-β (Aβ) plaques and intracellular aggregation of highly phosphorylated tau (p-tau)-containing neurofibrillary tangles [4]. Based on the 2018 AD research framework from the National Institute on Aging-Alzheimer’s Association workgroups, it is Aβ plaques and neurofibrillary tau deposits that define AD as a unique neurodegenerative disease. And low Aβ42 and elevated p-tau in cerebrospinal fluid (CSF) are biomarkers of Aβ plaques and fibrillar tau, respectively [5]. In this case, we aim to assist in the exploration of AD risk loci by identifying the risk SNPs which correlate with CSF biomarkers.
Genome-wide association studies and meta-analyses have identified several genes associated with AD risk, such as the alpha 2 Macroglobulin (
A few studies have been devoted to exploring the association between susceptibility loci and CSF biomarkers. With the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database, previous studies have investigated the correlation between 15
We hypothesized that the common SNPs within or near the coding regions of AD risk genes were potential functional loci for AD. The mutations close to the splicing junction might affect mRNA splicing. Missense mutations in the coding region could alter amino acid products. And synonymous mutations might influence promoter activity, pre-mRNAs conformation, stability, protein folding, or function [14–16]. The altered gene products might further affect cholesterol metabolism, immune response, Aβ processing, tau phosphorylation, etc., and ultimately contribute to AD pathogenesis.
In present study, we focused on the common SNPs within or near the coding regions of 13 known AD susceptible genes (
METHODS
Participants
The participants were from the dementia cohort of Peking Union Medical College Hospital. All subjects had basic tests, including history inquiry, physical examination, blood biochemical test, cognitive assessment, and brain CT/MRI. Cognitive assessment included the Mini-Mental State Exam (MMSE), activities of daily living (ADL), etc. 315 cases received whole exon sequencing, and 1,144 cases had targeted exon sequencing of 278 dementia-related genes. CSF testing of Aβ42, 181p-tau, and t-tau was performed in 362 subjects. Pathological evidence was not obtained.
75 Chinese Han patients were included in the study. The inclusion criteria were as follows: 1) met the diagnostic criteria for probable AD based on 2011 diagnostic guidelines for AD from National Institute on Aging-Alzheimer’s Association workgroups [17]; 2) no family history of dementia, and no potential pathogenic variant implicated in dementia; (3) no missing data on basic tests, gene sequencing, CSF testing of Aβ42, 181p-tau, and t-tau; and 4) informed consent was obtained. This study was approved by ethics committee of PUMCH (No. JS-1836).
Gene sequencing
IDT xGen Lockdown Probes were used in whole exon sequencing with 50 bp flanking intron length. The targeted exon sequencing covered 3927 exons with 20 bp flanking intron regions. The DNA libraries were sequenced on Illumina HiSeq X Ten Analyzers (Illumina, San Diego, USA). “Clean reads” were generated with AfterQC [18]. All reads were aligned to human reference genome 19 with Burrows-Wheeler Aligner (v.0.5.9) [19]. Local realignment and base quality recalibration were finished with GATK IndelRealigner and BaseRecalibrator (v3.5). SNVs and small indels were called with GATK UnifiedGenotyper (v 3.5).
This report focused on 13 known AD susceptible genes, including
CSF testing
CSF samples was collected using low protein binding tube (Eppendorf Protein LoBind Tube; 1.5 ml; Hamburg, Germany). The tubes were filled more than 75%. Within one day, the samples were centrifuged at 1800 rpm, 4°C for 10 min. After that, the supernatant was moved to new tubes and stored in freezers at –80°C. Within two weeks, CSF Aβ42, 181p-tau, and t-tau were measured using ELISA method (INNOTEST hTAU Ag, PHOSPHO-TAU, β-AMYLOID (1–42); Fujirebio, Ghent, Belgium).
The CSF biomarker cut-off values for clinical AD in our laboratory were submitted to Alzheimer’s Association International Conference in 2019 (https://doi.org/10.1016/j.jalz.2019.06.790). With the increasing sample size, the current CSF biomarker cut-offs for clinical AD have been set as follows: Aβ42 <570.9 pg/ml, p-tau > 56.49 pg/ml, t-tau > 241.6 pg/ml, p-tau/Aβ42 > 0.08465, and t-tau/Aβ42 > 0.5290. The sensitivities of these cut-offs are 56.6%, 88.7%, 81.1%, 77.4%, and 84.9%, respectively. The specificities are 84.6%, 84.6%, 90.4%, 90.4%, and 90.4%, respectively. These cut-offs are unpublished data.
Statistical analysis
The CSF biomarker data (Aβ42, 181p-tau, t-tau, 181p-tau/Aβ42, t-tau/Aβ42) were compared by analysis of covariate (ANCOVA). Sex, age, disease course, and
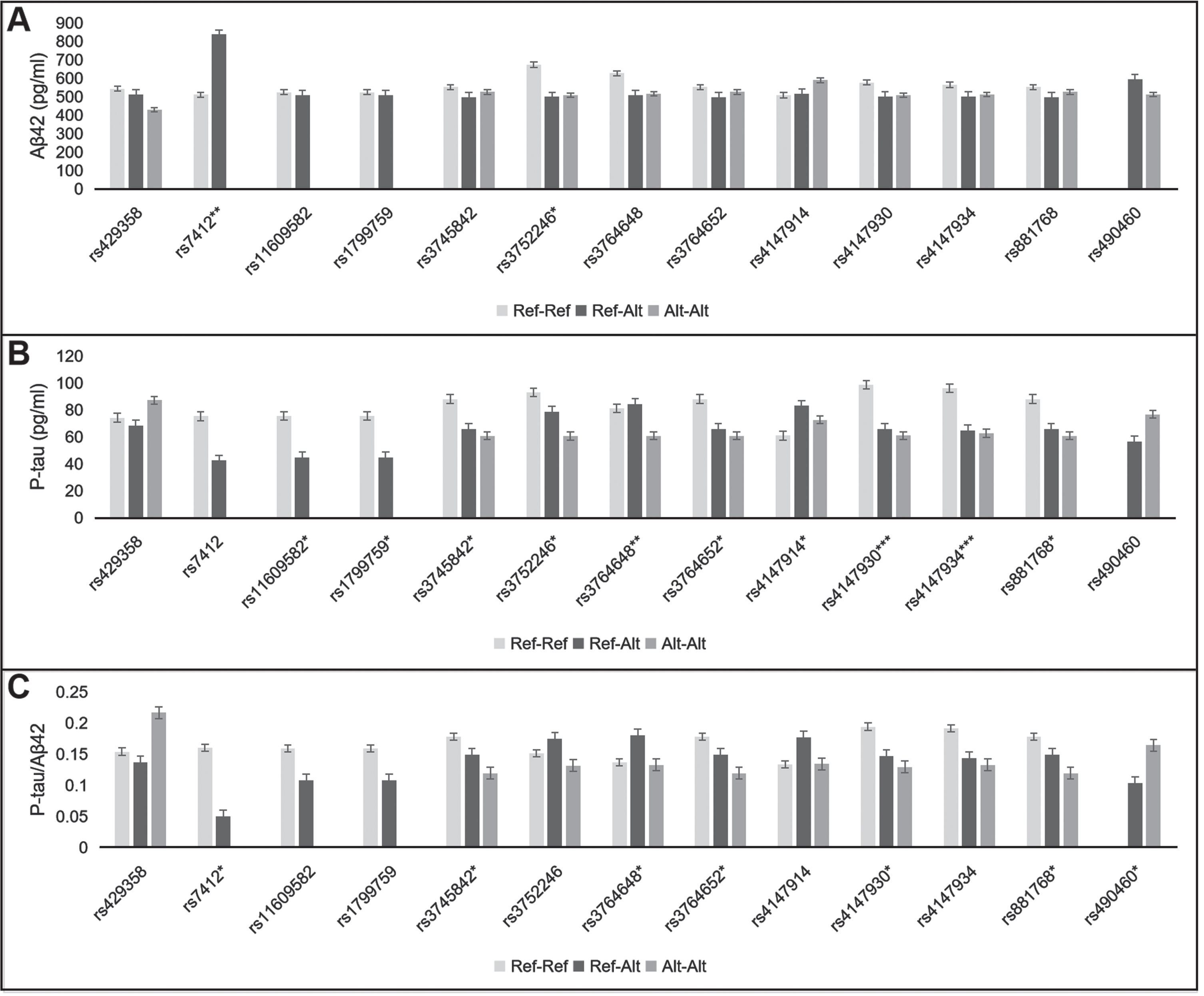
Effect of 13 SNPs on CSF Aβ42 (A), p-tau (B), and p-tau/Aβ42 ratio (C). Ref-Ref, reference allele homozygote; Ref-Alt, heterozygote; Alt-Alt, alternate allele homozygote. *
RESULTS
Demographic characteristics
As shown in Table 1, there were 75 subjects: 35 male and 40 female. They had an average age of 61.7±7.7 years old. The average disease course was 3.5±2.8 years. None of them had positive family history of dementia. None of them harbored potential pathogenic mutations implicated in dementia.
General characteristics of 75 subjects
SNPs frequency
As shown in Supplementary Table 1, there were 69 SNPs in 13 risk genes. Of them, 58 were in the exon (24 missense and 34 synonymous), whereas 11 were within 10 bp of splicing junction in the intron.
Compared with the allele frequency in east Asian populations from 1000 Genomes database, the cohort had higher rs429358-C (
Association between SNPs and CSF biomarkers
As illustrated in Fig. 1 and Supplementary Tables 2 and 3, 13 SNPs were relevant to CSF biomarkers, including two
After adjustment by the rs429358 (
The rs3752246-C (
The rs11609582-TT (
The rs490460-TT (
Sex*SNP interaction
As shown in Table 2, two SNPs showed sex*SNPs interactions on CSF Aβ42 level, including the rs3752240 (
Sex*SNP interactions on CSF biomarkers [F value (
Among the subjects with the rs3752240-AA and the rs7232-AT genotypes, males had lower Aβ42 level than females. Among the participants with the rs2598044-CT and the rs2722372-GA genotypes, the p-tau/Aβ42 ratio was higher in males relative to females. However, among those with rs3752240-AG, rs7232-AA, rs2598044-CC, and rs2722372-GG genotypes, Aβ42 or p-tau/Aβ42 ratio did not differ between males and females.
DISCUSSION
APOE, ABCA7, A2M, BACE1 SNPs and CSF Aβ42, p-tau
The
Among the 13 potential functional SNPs, 61.5% (8/13) SNPs are from the
Two novel
One novel
Allele frequency variation of CR1, SORL1
The alternate allele frequencies of rs2274567 (
Sex*SNP (ABCA7, MS4A6A, NME8) interaction
This is the first time that we demonstrate the interactions between sex and
P-tau biased association
In this paper, ten SNPs show effects on CSF p-tau. Comparatively, there is less association between these SNPs and CSF Aβ42. This might be partly due to the dynamic evolution of CSF biomarkers at different stages of cognitive impairment. The CSF Aβ42 level decreases mainly in the preclinical stage and reaches a plateau in the dementia stage, whereas the CSF tau level increases gradually in the mild dementia stage [34–36]. T-tau is a non-specific biomarker for AD which reflects the intensity of neuronal injury, whereas p-tau is relatively specific for AD which indicates a pathological state of paired helical filament tau deposits [5]. This might explain why these SNPs are associated with p-tau rather than t-tau.
Conclusion and limitation
This study involves 69 common SNPs within or near coding regions of 13 AD risk genes.
The main limitation of this study is the small sample size and absence of pathological evidence. All the subjects in this study are clinically diagnosed with probable AD. None of them have pathological evidence. According to the CSF biomarker cut-offs for AD in our laboratory, only 50.7% (38/75) subjects reach the cutoff values of all the five variables. 29.3% (22/75) subjects have CSF Aβ42 > 570.9 pg/ml, and 25.3% (19/75) subjects have CSF p-tau <56.49 pg/ml. The atypical CSF profile might be attributed to individual differences in CSF dynamics. Grothe studied the CSF biomarkers in pathology-confirmed AD patients from ADNI database. They reported a considerably higher CSF Aβ42 cutoff (1,097 pg/ml) for differentiating high and low Thal phases, as well as a lower p-tau cutoff (19 pg/ml) for discriminating high and low neuritic plaque scores [37]. In this case, pathological evidence is highly expected in order to reduce the bias of the study. Next, we expect to expand the sample size and strive for pathological evidence. We will explore the synergistic effect of multiple SNPs on CSF biomarkers. As for the potential susceptibility loci, functional analysis should be considered.
Footnotes
ACKNOWLEDGMENTS
Dr. Jing Gao was supported by grants from National Key Research and Development Program of China (2020YFA0804500, 2016YFC1306300), CAMS Innovation fund for Medical Sciences (2016-I2M-1-004), National Natural Science Foundation of China (81550021, 30470618), and strategic priority research program (pilot study) “Biological basis of aging and therapeutic strategies” of Chinese Academy of Sciences (XDPB10).
